AGENDA for Personalized Immunotherapy – Personalized Oncology in the Genomic Era: Expanding the Druggable Space CHI’S 4TH ANNUAL IMMUNO-ONCOLOGY SUMMIT – AUGUST 30-31, 2016 | Marriott Long Wharf Hotel – Boston, MA
Reporter: Aviva Lev-Ari, PhD, RN
Personalized Immunotherapy Personalized Oncology in the Genomic Era: Expanding the Druggable Space
http://www.immuno-oncologysummit.com/uploadedFiles/Immuno_Oncology_Summit/Agenda/16/2016-The-Immuno-Oncology-Summit-Brochure.pdf
TUESDAY, AUGUST 30 & WED, AUGUST 31
TUESDAY, AUGUST 30
12:00 pm Registration
TUMOR NEOANTIGENS FOR PERSONALIZED IMMUNOTHERAPY
1:15 Chairperson’s Opening Remarks
Pramod K. Srivastava, M.D., Ph.D., Professor, Immunology and Medicine, Director, Carole and Ray Neag Comprehensive Cancer Center, University of Connecticut School of Medicine
1:20 Basics of Personalized Immunotherapy: What Is a Good Antigen? Pramod K. Srivastava, M.D., Ph.D., Professor, Immunology and Medicine, Director, Carole and Ray Neag Comprehensive Cancer Center, University of Connecticut School of Medicine
The definition of host-protective immunogenic antigen(s) of any human cancer of non-viral origin is still an enigma. New approaches in cancer genomics and bioinformatics are now offering a plethora of candidate antigens, whose role in cancer immunity, and specifically in host-protective cancer immunity, is under extensive testing. Outlines of some broad rules are emerging and some of these shall be discussed.
1:50 Novel Antibodies against Immunogenic Neoantigens
Philip M. Arlen, M.D., President & CEO, Precision Biologics, Inc.
Two novel antibodies, NEO-102 (ensituximab) and NEO- 201, were developed from an allogeneic colorectal cancer vaccine that had previously shown activity in patients with metastatic colorectal cancer This vaccine was derived from an immunogenic component of the cell membrane from pooled surgical specimens from both primary and metastatic colon cancer. Patients who benefited from the vaccine in the prior clinical trial produced and sustained high levels of serum IgG against the vaccine. Several thousand candidate antibodies were screened against this vaccine and NEO- 102 and NEO-201 were candidates that demonstrated the ability to bind to colon cancer vs. normal tissue.
2:20 PD-1 Blockade in Tumors with Mismatch-Repair Deficiency
Luis Alberto Diaz, M.D., Associate Professor, Oncology, Johns Hopkins Sidney Kimmel Comprehensive Cancer Center
Somatic mutations have the potential to encode “non-self” immunogenic antigens. Tumors with a large number of somatic mutations due to mismatch-repair defects appear to be highly susceptible to immune checkpoint blockade. This presentation will summarize the clinical and genomic data of using mutations as neoantigens.
2:50 Sponsored Presentations (Opportunities Available)
3:20 Refreshment Break in the Exhibit Hall with Poster Viewing
4:00 PLENARY KEYNOTE SESSION See Keynotes for details.
4:00 A New Era of Personalized Therapy: Using Tumor Neoantigens to Unlock the Immune System Matthew J. Goldstein, M.D., Ph.D., Director, Translational Medicine, Neon Therapeutics, Inc.
Neon Therapeutics, Inc. launched in 2015 to focus on advancing neoantigen biology to improve cancer patient care. A neoantigen-based product engine will allow Neon to develop further treatment modalities including next-generation vaccines and T cell therapies targeting both personalized as well as shared neoantigens. The company’s first trial will launch later this year investigating the combination of a personalized, vaccine with nivolumab in advanced Melanoma, NSCLC, and Bladder Cancer.
4:30 Emerging Innate Immune Targets for Enhancing Adaptive Anti-Tumor Responses
Michael Rosenzweig, Ph.D., Executive Director, Biology-Discovery, IMR Early Discovery, Merck Research Laboratories
Novel cancer immunotherapies targeting T cell checkpoint proteins have emerged as powerful tools to induce profound, durable regression and remission of many types of cancer. Despite these advances, multiple studies have demonstrated that not all patients respond to these therapies, and the ability to predict which patients may respond is limited. Harnessing the innate immune system to augment the adaptive anti-tumor response represents an attractive target for therapy, which has the potential to enhance both the percentage and rate of response to checkpoint blockade.
5:00 Reading Tea Leaves: The Dilemma of Prediction and Prognosis in Immunotherapy
Morganna Freeman, D.O., Associate Director, Melanoma & Cutaneous Oncology Program, The Angeles Clinic and Research Institute
With the rapid expansion of immunotherapeutics in oncology, scientifically significant advances have been made with both the depth and duration of antitumor responses. However, not all patients benefit, or quickly relapse, thus much scientific inquiry has been devoted to appropriate patient selection and how such obstacles might be overcome. While more is known about potential biomarkers, accurate prognostication persists as a knowledge gap, and efforts to bridge it will be discussed here.
5:30 Welcome Reception in the Exhibit Hall with Poster Viewing
WEDNESDAY, AUGUST 31
8:00 am Morning Coffee
PERSONALIZED IMMUNOTHERAPY WITH CANCER VACCINES
8:25 Chairperson’s Remarks
Ravi Madan, M.D., Clinical Director, Genitourinary Malignancies Branch, National Cancer Institute, National Institutes of Health
8:30 Cancer Vaccines in the Era of Checkpoint Inhibitors
Keith L. Knutson, Ph.D., Professor, Immunology, Mayo Clinic
Vaccination has been one of the most successful approaches to reduce incidence and mortality rates of infectious diseases and more recently cancer, including cervical cancer. Our goal is to develop vaccine strategies that can be delivered to breast cancer patients to boost host immune defenses following conventional treatments (e.g., surgery, chemotherapy, and radiation), in order to prevent recurrence of treatment resistant tumors. We believe that one of the better approaches is to vaccinate against abnormally expressed ‘self’ (non-mutated) antigens that contribute to the cancer initiation and progression.
9:00 Developing Therapeutic Cancer Vaccine Strategies for Prostate Cancer
Ravi Madan, M.D., Clinical Director, Genitourinary Malignancies Branch, National Cancer Institute, National Institutes of Health
The development of immunotherapy strategies has become the primary focus in oncology. This lecture will provide prostate cancer as a template to demonstrate synergies between immune-based therapies and chemotherapy, radiopharmaceuticals and hormonal therapies.
9:30 Getting Very Personal: Fully Individualized Tumor Neoantigen-Based Vaccine Approaches to Cancer Therapy
Karin Jooss, Ph.D., CSO, Gritstone Oncology
Genetic instability in tumors generates tumor-specific neoantigens which have been identified as the targets of new T cells in patients responding to checkpoint inhibitor therapy. Predicting neoantigens by sequencing routine clinical biopsy material, and then incorporating them into therapeutic cancer vaccines is an attractive concept being developed by Gritstone Oncology. The complexities of neoantigen prediction will be discussed, together with insights into how vaccine vectors are selected and designed.
10:00 Approaches to Assess Tumor Mutation Load for Selecting Patients for Cancer Immunotherapy
John Simmons, Ph.D., Manager, Research Services, Personal Genome
Diagnostics Tests to identify patients who are most likely to benefit from cancer immunotherapies are urgently needed. Here we discuss PGDx approaches to assess tumor mutation load as a potential predictor of clinical benefit for checkpoint inhibitors in multiple cancer types.
10:15 Sponsored Presentation (Opportunity Available)
10:30 Coffee Break in the Exhibit Hall with Poster Viewing
11:15 In situ Vaccination for Lymphoma
Joshua Brody, M.D., Director, Lymphoma Immunotherapy Program, Icahn School of Medicine at Mount Sinai
Prior ex vivo combinations of dendritic cells (DC) with tumor antigens have yielded immunologic and clinical responses. Intratumoral immunomodulation may bypass the need for ex vivo production of vaccine. In situ vaccination combines: intratumoral Flt3L to recruit DC, low dose radiotherapy to load DC with tumor antigens, and intratumoral TLR agonist to activate tumor-antigen-loaded DC. Preliminary results demonstrate DC recruitment and activation, systemic tumor regressions, and induction of neoantigen specific CD8 T cell responses after vaccination.
11:45 Immunotherapy Using Ad5 [E1-, E2b-] Vector Vaccines in the Cancer MoonShot 2020 Program
Frank R. Jones, Ph.D., Chairman & CEO, Etubics Corporation
The Cancer MoonShot 2020 project intends to design, initiate and complete randomized clinical trials at all stages of cancer in up to 20 tumor types in as many as 20,000 patients by the year 2020. Etubics is participating in the Cancer MoonShot 2020 program by providing its proprietary viral platform, known as Ad5 [E1-, E2b-] as a treatment agent in several of the program’s immunotherapeutic vaccination initiatives and trials.
12:15 pm Sponsored Presentations (Opportunities Available)
12:45 Luncheon Presentation to be Announced
Robert G. Petit, Ph.D., Executive Vice President & CSO, Advaxis Immunotherapies
1:15 Session Break
PERSONALIZED CELL THERAPY
1:55 Chairperson’s Remarks
Andrew M. Evens, D.O., Professor and Chief, Hematology/Oncology, Tufts University School of Medicine; Director, Tufts Cancer Center
2:00 Integration of Natural Killer-Based Therapy into the Treatment of Lymphoma Andrew M. Evens, D.O., Professor and Chief, Hematology/Oncology, Tufts University School of Medicine; Director, Tufts Cancer Center
Targeting signaling pathways or epitopes with small molecules and antibody-based immunotherapeutic agents is a leading strategy for cancer therapy. Promising immunotherapy agents being examined for the treatment of lymphoma include monoclonal antibodies, immunomodulatory agents, PD-1 inhibitors, chimeric antigen receptor (CAR) T-cells, and NK-based therapies. The optimum combinations or sequences of these therapeutics continue to be defined. Additionally, understanding tumor and patient/host heterogeneity is desired in order to optimize personalized medicine.
2:30 Dendritic Cells: Personalized Cancer Vaccines and Inducers of Multi-Epitope Specific T Cells for Adoptive Cell Therapy
Pawel Kalinski, M.D., Ph.D., Professor, Surgery, Immunology, and Bioengineering, University of Pittsburgh School of Medicine, University of Pittsburgh Cancer Institute
Conditions of dendritic cell (DC) maturation affect their ability to cross-present cancer cell-derived antigens and induce clonal expansion and effector functions of responding T cells. We will discuss the pathways of DC maturation which promote their preferential interaction with naïve, memory and effector T cells, cross-presentation of antigens from dead cancer cells, and induction of large numbers of type-1 CD4+ and CD8+ T cells specific for multiple tumor-associated antigens ex vivo and in vivo.
3:00 Mesothelin-Targeted CAR T-Cell Therapy for Solid Tumors
Prasad S. Adusumilli, M.D., FACS, Deputy Chief of Translational & Clinical Research, Thoracic Surgery, Memorial Sloan-Kettering Cancer Center
Mesothelin, a cell-surface antigen, provides an exciting prospect based on its higher expression in a majority of solid tumors (estimated annual incidence of 340,000 and prevalence of 2 million patients in the U.S.), limited expression in normal tissues and its association with tumor aggressiveness. CAR T-cell therapy with second generation mesothelin-targeted CARs has been translated to clinical trials targeting mesothelioma, non-small cell lung cancer, triple-negative breast cancer, and other solid tumors.
3:30 Refreshment Break with Exhibit and Poster Viewing
4:15 Synthetic Regulation of T Cell Therapies Adds Safety and Enhanced Efficacy to Previously Unpredicted Therapies
David M. Spencer, Ph.D., CSO, Bellicum Pharmaceuticals
CAR- and TCR-based T cell therapies have had some spectacular successes in a handful of malignancies, but safety and efficacy concerns still impede broader adoption of these new technologies. Bellicum Pharmaceuticals has developed a suite of synthetic ligand-inducible switches to rapidly and rigorously regulate T cell therapies. These potent switches address both safety and anti-tumor efficacy and promise to further expand the reach of immunotherapy.
4:45 Long-Term Relapse-Free Survival of Patients with Acute Myeloid Leukemia (AML) Receiving a Telomerase- Engineered Dendritic Cell Immunotherapy
Jane Lebkowski, Ph.D., President & CSO, Research and Development, Asterias Biotherapeutics
There are few treatment options for patients with intermediate and high risk AML, and remission and relapse rates are dismal, especially in patients ≥ 60 years old. A Phase II clinical trial was conducted in subjects with AML to assess a dendritic cell immunotherapy (ASTVAC1) engineered to express a modified form of telomerase that is processed through both the MHC Class I and II antigen presentation pathways. The results suggest that immunotherapy with AST-VAC1 is safe, can stimulate immune responses to telomerase, and may extend relapse-free survival even in patients with high risk AML.
5:15 Activated and Exhausted Tumor Infiltrating B Cells in Non-Small Cell Lung Cancer Patients Present Antigen and Influence the Phenotype of CD4 Tumor Infiltrating T Cells
Tullia Bruno, Ph.D., Research Assistant Professor, Immunology, University of Pittsburgh
The focus of immunotherapy has been on subsets of CD8 and CD4 tumor infiltrating lymphocytes (TILs), however, tumor infiltrating B cells (TIL-Bs) have been reported in tertiary lymphoid structures (TLS) with CD4 TILs, and both TIL-Bs and TLS correlate with NSCLC patient survival. While TIL-Bs have been identified in NSCLC patients, their function in the tumor microenvironment has been understudied with no focus on their role as antigen presenting cells (APCs) and their influence on CD8 and CD4 TILs. Here, we demonstrate that TIL-Bs can efficiently present antigen to CD4 TILs and influence CD4 TIL phenotype depending on their exhaustion profile.
5:30 Dinner Short Course Registration
5:45 Close of Personalized Immunotherapy Conference
Read Full Post »










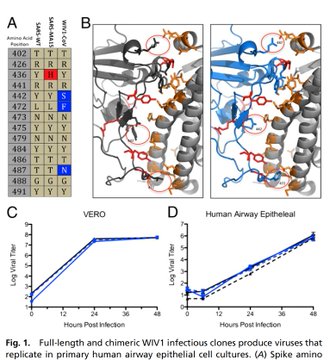






Paper in collection COVID-19 SARS-CoV-2 preprints from medRxiv and bioRxiv