Genomics & Genetics of Cardiovascular Disease Diagnoses: A Literature Survey of AHA’s Circulation Cardiovascular Genetics, 3/2010 – 3/2013
Curators: Aviva Lev-Ari, PhD, RN and Larry H. Bernstein, MD, FCAP
348 articles that appeared in AHA’s Circulation Cardiovascular Genetics, 3/2010 – 3/2013 were classified by the curators of this article into the following TEN categories. The first 9, represent DIAGNOSES of cardiovascular diseases, the last, deals with Pharmacogenomics.
The Cardiovascular Diagnoses that were covered in the period of 3/2010 – 3/2013, include the following:
- MicroRNA in Serum as Bimarker for Cardiovascular Pathologies: acute myocardial infarction, viral myocarditis, diastolic dysfunction, and acute heart failure
- Genetic Determinants of Potassium Sensitivity and Hypertension
- Heart and Aging Research in Genomic Epidemiology: 1700 MIs and 2300 coronary heart disease events among about 29 000 eligible patients
- Genetics of CVD and Hyperlipidemia, Hyper Cholesterolemia, Metabolic Syndrome
- Genomics and Valvular Disease
Introductions
Larry H. Bernstein, MD, FCAP
The curation of this large amount of material in 10 categories begins with a first chapter on preventative cardiology, which has had much public attention for the last decade. Much of the concern with preventive cardiology has emphasized diet and exercise. There is much to be said about this in articles not yet written. However, there are several decades of research on the amino acid composition of foods, and the essential fatty acids, that indicates an essential balance between proinflammatory and antiinflammatory fatty acids in polyunsaturated fatty acids, and of the harmful effects of saturated fats. There is also much to be said of essential amino acids, and in particular, those essential for methylation processes, and sulfur metabolism.
The next eight chapters are all concerned with genomics in cardiovascular disease. This is in no small part a follow up on the completion of the genetic code in 2003, a seminal event. Let us look at these in clusters.
[1] microRNA in serum is now considered for a biomarker for cardiovascular disease. It can be measured at very low levels, but we don’t yet know where it fits. It might be more revealing once we understand the adaptive mechanism in development of congestive heart failure, renal hypertension, and post-genomic events.
[2] It appears to me that potassium sensitivity and hypertension approached from the genomic side is more complicate. Why is that? The kidney excretes a sodium load and in metabolic acidosis, the serum potassium rises with a metabolic acidemia that can’t be compensated by the respiratory loss of CO2 through the carbonic anhydrase mechanism.
[3] Heart and aging research is a rich area for work on the long term post-genomic changes, and it involves a large population base.
[4][5] The genomics of cardiac dysrrhytmias and cardiomyopathies will open new doors into our understanding of the mechanisms of these diseases, and perhaps find therapeutic targets. There has been a large volume of work on lipid synthesis, the role of the liver in generating apolipoproteins, and this has new answers on the way. The most important feature, not readily accepted is the measurement of particles, which has now been done by a monoclonal antibody. Metabolic syndrome brings together adipose tissue metabolism, endocrine and changes in CRP and IL-1.
[6] Vascular pathologies and coagulation, hyperviscosity has had an enormous increase in intensity of research. The concept of plaque rupture to account for all AMIs is being modified, and the high sensitivity cardio-specific troponins have become the most widely use test.
[7] The genomics of valvular disease fits with the increased surgical procedures for valvular disease related to atheroschlerosis and advent of minimally invasive surgical procedures for the reapir and replacement of valves, procedure called TAVR vs. Openhealrt surgery for valve replacement.
[8] Inherited cardiovascular disease is an older family of disorders, going back to Victor McKusik, and also the “Blue Baby” operation, both at Johns Hopkins.
[9] Pharmacogenomics is a vary active field of investigation and has uncovered inter-individual differences in handling Warfarin as a starter.
Preventative Cardiology
Methods in Genetics and Clinical Interpretation Randomized Trial of Personal Genomics for Preventive Cardiology Design and Challenges
Joshua W. Knowles, MD, PhD, Themistocles L. Assimes, MD, PhD, Michaela Kiernan, PhD, Aleksandra Pavlovic, BS, Benjamin A. Goldstein, PhD, Veronica Yank, MD, Michael V. McConnell, MD, Devin Absher, PhD, Carlos Bustamante, PhD, Euan A. Ashley, MD, DPhil and John P.A. Ioannidis, MD, DSc
Author Affiliations
From the Division of Cardiovascular Medicine (J.W.K., T.L.A., A.P., M.V.M., E.A.A.), Stanford Prevention Research Center (M.K., V.Y., J.P.A.I.), Division of General Medical Disciplines (V.Y.), Department of Genetics (C.B.), Department of Health Research and Policy (J.P.A.I.), Stanford University School of Medicine, Stanford, CA; Quantitative Sciences Unit, Stanford University School of Medicine, Palo Alto, CA (B.A.G.); HudsonAlpha Institute for Biotechnology, Huntsville, AL (D.A.); Department of Statistics, Stanford University School of Humanities and Sciences, Stanford, CA (J.P.A.I.).
Correspondence to Joshua W. Knowles, MD, PhD, Stanford University School of Medicine, Division of Cardiovascular Medicine, Falk CVRC, 300 Pasteur Dr, Stanford, CA 94305. E-mail knowlej@stanford.edu
Background
Genome-wide association studies (GWAS) have identified more than 1500 disease-associated single nucleotide polymorphisms (SNPs), including many related to atherosclerotic cardiovascular disease (CVD). Associations have been found for most traditional risk factors (TRFs), including lipids,1,2 blood pressure/hypertension,3,4 weight/body mass index,5,6 smoking behavior,7 and diabetes.8–13 GWAS have also identified susceptibility variants for coronary heart disease (CHD). The first and, so far, strongest of these signals was found in the 9p21.3 locus, where common variants in this region increase the relative risk of CVD by 15% to 30% per risk allele in most race/ethnic groups.13–20 Subsequent large-scale GWAS meta-analyses and replication studies in largely white/European populations have led to the reliable identification of an additional 26 loci conferring susceptibility to CHD,2,20–23 all with substantially lower effects sizes compared with the 9p21 locus. Many of these CVD susceptibility loci appear to be conferring risk independent of TRFs and thus cannot currently be assessed by surrogate clinical measures (Table 1). Among the 27 independent loci identified in the most recent large meta-analyses of CVD, 21 were reported not to be associated with any of the TRFs.20,21
SOURCE
Circulation: Cardiovascular Genetics 2012; 5: 368-376
doi: 10.1161/ CIRCGENETICS.112.962746
MicroRNA in Serum as Bimarker for Cardiovascular Pathologies: acute myocardial infarction, viral myocarditis, diastolic dysfunction, and acute heart failure
Increased MicroRNA-1 and MicroRNA-133a Levels in Serum of Patients With Cardiovascular Disease Indicate Myocardial Damage
Yasuhide Kuwabara, MD, Koh Ono, MD, PhD, Takahiro Horie, MD, PhD, Hitoo Nishi, MD, PhD, Kazuya Nagao, MD, PhD, Minako Kinoshita, MD, PhD, Shin Watanabe, MD, PhD, Osamu Baba, MD, Yoji Kojima, MD, PhD, Satoshi Shizuta, MD, Masao Imai, MD, Toshihiro Tamura, MD, Toru Kita, MD, PhD and Takeshi Kimura, MD, PhD
Author Affiliations
From the Department of Cardiovascular Medicine, Graduate School of Medicine, Kyoto University, Kyoto, Japan (Y. Kuwabara, K.O., T.H., H.N., K.N., M.K., S.W., O.B., Y. Kojima, S.S., M.I., T.T., T. Kimura); and Kobe City Medical Center General Hospital, Kobe, Japan (T. Kita).
Correspondence to Koh Ono, MD, PhD, Department of Cardiovascular Medicine, Graduate School of Medicine, Kyoto University, 54 Shogoin-kawahara-cho, Sakyo-ku, Kyoto, Japan 606-8507. E-mail kohono@kuhp.kyoto-u.ac.jp
Abstract
Background—Recently, elevation of circulating muscle-specific microRNA (miRNA) levels has been reported in patients with acute myocardial infarction. However, it is still unclear from which part of the myocardium or under what conditions miRNAs are released into circulating blood. The purpose of this study was to identify the source of elevated levels of circulating miRNAs and their function in cardiovascular diseases.
Conclusions—These results suggest that elevated levels of circulating miR-133a in patients with cardiovascular diseases originate mainly from the injured myocardium. Circulating miR-133a can be used as a marker for cardiomyocyte death, and it may have functions in cardiovascular diseases.
SOURCE:
Circulation: Cardiovascular Genetics. 2011; 4: 446-454
Published online before print June 2, 2011,
doi: 10.1161/ CIRCGENETICS.110.958975
Selective Articles citing this article
Plasma microRNAs serve as biomarkers of therapeutic efficacy and disease progression in hypertension-induced heart failure
Eur J Heart Fail. 2013;0:hft018v1-hft018,
AbstractFull TextPDF
Intercellular Transport of MicroRNAs
Arterioscler. Thromb. Vasc. Bio.. 2013;33:186-192,
AbstractFull TextPDF
MicroRNAs Within the Continuum of Postgenomics Biomarker Discovery
Arterioscler. Thromb. Vasc. Bio.. 2013;33:206-214,
AbstractFull TextPDF
Reply to “Need for Rigor in Design, Reporting, and Interpretation of Transcriptomic Biomarker Studies”
J. Clin. Microbiol.. 2012;50:4192-4193,
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3502969/
Full TextPDF
Circulating microRNAs as diagnostic biomarkers for cardiovascular diseases
Am. J. Physiol. Heart Circ. Physiol.. 2012;303:H1085-H1095,
AbstractFull TextPDF
Sonic Hedgehog-Modified Human CD34+ Cells Preserve Cardiac Function After Acute Myocardial Infarction
Circ. Res.. 2012;111:312-321,
AbstractFull TextPDF
Metabolomics, Proteomics, Genomics, and Transcriptomics in Circulation and the Circulation Subspecialty Journals
Circulation. 2012;125:e507-e519,
AbstractFull TextPDF
Circulating MicroRNAs: Novel Biomarkers and Extracellular Communicators in Cardiovascular Disease?
Circ. Res.. 2012;110:483-495,
AbstractFull TextPDF
Circulating MicroRNAs in Cardiovascular Disease
Circulation. 2011;124:1908-1910,
Full TextPDF
Circulating MicroRNAs: Biomarkers or Mediators of Cardiovascular Diseases?
Arterioscler. Thromb. Vasc. Bio.. 2011;31:2383-2390,
AbstractFull TextPDF
Circulating MicroRNA-208b and MicroRNA-499 Reflect Myocardial Damage in Cardiovascular Disease
Maarten F. Corsten, MD, Robert Dennert, MD, Sylvia Jochems, BSc, Tatiana Kuznetsova, MD, PhD, Yvan Devaux, PhD, Leon Hofstra, MD, PhD, Daniel R. Wagner, MD, PhD, Jan A. Staessen, MD, PhD, Stephane Heymans, MD, PhD and Blanche Schroen, PhD
Author Affiliations
From the Center for Heart Failure Research (M.F.C., R.D., S.J., S.H., B.S.), Cardiovascular Research Institute, Maastricht, The Netherlands; the Division of Hypertension and Cardiovascular Rehabilitation (T.K., J.A.S.), Department of Cardiovascular Diseases, University of Leuven, Leuven, Belgium and Department of Epidemiology, Maastricht University Medical Center, Maastricht, The Netherlands; Centre de Recherche Public–Santé, Luxembourg (Y.D., D.R.W.), Luxembourg; Maastricht University Medical Center (L.H.), Maastricht, The Netherlands; and Centre Hospitalier Luxembourg (D.R.W.), Luxembourg.
Correspondence to Blanche Schroen, PhD, Center for Heart Failure Research, Cardiovascular Research Institute Maastricht, Universiteitssingel 50, 6229 ER Maastricht, The Netherlands. E-mail b.schroen@cardio.unimaas.nl
Drs Heymans and Schroen contributed equally to this work.
Abstract
Background— Small RNA molecules, called microRNAs, freely circulate in human plasma and correlate with varying pathologies. In this study, we explored their diagnostic potential in a selection of prevalent cardiovascular disorders.
Methods and Results— MicroRNAs were isolated from plasmas from well-characterized patients with varying degrees of cardiac damage:
(1) acute myocardial infarction,
(2) viral myocarditis,
(3) diastolic dysfunction, and
(4) acute heart failure.
Plasma levels of selected microRNAs, including heart-associated (miR-1, -133a, -208b, and -499), fibrosis-associated (miR-21 and miR-29b), and leukocyte-associated (miR-146, -155, and -223) candidates, were subsequently assessed using real-time polymerase chain reaction. Strikingly, in plasma from acute myocardial infarction patients, cardiac myocyte–associated miR-208b and -499 were highly elevated, 1600-fold (P<0.005) and 100-fold (P<0.0005), respectively, as compared with control subjects. Receiver operating characteristic curve analysis revealed an area under the curve of 0.94 (P<10−10) for miR-208b and 0.92 (P<10−9) for miR-499. Both microRNAs correlated with plasma troponin T, indicating release of microRNAs from injured cardiomyocytes. In viral myocarditis, we observed a milder but significant elevation of these microRNAs, 30-fold and 6-fold, respectively. Plasma levels of leukocyte-expressed microRNAs were not significantly increased in acute myocardial infarction or viral myocarditis patients, despite elevated white blood cell counts. In patients with acute heart failure, only miR-499 was significantly elevated (2-fold), whereas no significant changes in microRNAs studied could be observed in diastolic dysfunction. Remarkably, plasma microRNA levels were not affected by a wide range of clinical confounders, including age, sex, body mass index, kidney function, systolic blood pressure, and white blood cell count.
Conclusions— Cardiac damage initiates the detectable release of cardiomyocyte-specific microRNAs-208b and -499 into the circulation.
SOURCE:
Circulation: Cardiovascular Genetics. 2010; 3: 499-506
Published online before print October 4, 2010,
doi: 10.1161/ CIRCGENETICS.110.957415
Selective Articles citing this article
Plasma microRNAs serve as biomarkers of therapeutic efficacy and disease progression in hypertension-induced heart failure
Eur J Heart Fail. 2013;0:hft018v1-hft018,
AbstractFull TextPDF
Aspirin treatment hampers the use of plasma microRNA-126 as a biomarker for the progression of vascular disease
Eur Heart J. 2013;0:eht007v1-eht007,
AbstractFull TextPDF
MicroRNAs in Myocardial Infarction
Arterioscler. Thromb. Vasc. Bio.. 2013;33:201-205,
AbstractFull TextPDF
MicroRNAs Within the Continuum of Postgenomics Biomarker Discovery
Arterioscler. Thromb. Vasc. Bio.. 2013;33:206-214,
AbstractFull TextPDF
MicroRNAs Are Involved in End-Organ Damage During Hypertension
Hypertension. 2012;60:1088-1093,
AbstractFull TextPDF
Circulating microRNAs as diagnostic biomarkers for cardiovascular diseases
Am. J. Physiol. Heart Circ. Physiol.. 2012;303:H1085-H1095,
AbstractFull TextPDF
Circulation Editors’ Picks: Most Read Articles in Cardiovascular Genetics
Circulation. 2012;126:e163-e169,
AbstractFull TextPDF
MicroRNAs in Patients on Chronic Hemodialysis (MINOS Study)
CJASN. 2012;7:619-623,
AbstractFull TextPDF
Novel techniques and targets in cardiovascular microRNA research
Cardiovasc Res. 2012;93:545-554,
AbstractFull TextPDF
Microparticles: major transport vehicles for distinct microRNAs in circulation
Cardiovasc Res. 2012;93:633-644,
AbstractFull TextPDF
Profiling of circulating microRNAs: from single biomarkers to re-wired networks
Cardiovasc Res. 2012;93:555-562,
AbstractFull TextPDF
Small but smart–microRNAs in the centre of inflammatory processes during cardiovascular diseases, the metabolic syndrome, and ageing
Cardiovasc Res. 2012;93:605-613,
AbstractFull TextPDF
Circulation: Heart Failure Editors’ Picks: Most Important Papers in Pathophysiology and Genetics
Circ Heart Fail. 2012;5:e32-e49,
AbstractFull TextPDF
Use of Circulating MicroRNAs to Diagnose Acute Myocardial Infarction
Clin. Chem.. 2012;58:559-567,
AbstractFull TextPDF
Circulating MicroRNAs: Novel Biomarkers and Extracellular Communicators in Cardiovascular Disease?
Circ. Res.. 2012;110:483-495,
AbstractFull TextPDF
Serum levels of microRNAs in patients with heart failure
Eur J Heart Fail. 2012;14:147-154,
AbstractFull TextPDF
Circulating microRNAs to identify human heart failure
Eur J Heart Fail. 2012;14:118-119,
Full TextPDF
Next Steps in Cardiovascular Disease Genomic Research–Sequencing, Epigenetics, and Transcriptomics
Clin. Chem.. 2012;58:113-126,
AbstractFull TextPDF
Most Read in Cardiovascular Genetics on Biomarkers, Inherited Cardiomyopathies and Arrhythmias, Metabolomics, and Genomics
Circ Cardiovasc Genet. 2011;4:e24-e30,
AbstractFull TextPDF
MicroRNA-126 modulates endothelial SDF-1 expression and mobilization of Sca-1+/Lin- progenitor cells in ischaemia
Cardiovasc Res. 2011;92:449-455,
AbstractFull TextPDF
Circulating MicroRNAs: Biomarkers or Mediators of Cardiovascular Diseases?
Arterioscler. Thromb. Vasc. Bio.. 2011;31:2383-2390,
AbstractFull TextPDF
Transcoronary Concentration Gradients of Circulating MicroRNAs
Circulation. 2011;124:1936-1944,
AbstractFull TextPDF
MicroRNAs: New Players in Cardiac Injury and Protection
Mol. Pharmacol.. 2011;80:558-564,
AbstractFull TextPDF
Circulation Editors’ Picks: Most Read Articles in Cardiovascular Genetics, Part I
Circulation. 2011;124:e345-e357,
AbstractFull TextPDF
MicroRNAs regulating oxidative stress and inflammation in relation to obesity and atherosclerosis
FASEB J.. 2011;25:2515-2527,
AbstractFull TextPDF
miR-143 and miR-145: Molecular Keys to Switch the Phenotype of Vascular Smooth Muscle Cells
Circ Cardiovasc Genet. 2011;4:197-205,
Full TextPDF
Response to Letter Regarding Article, “Circulating MicroRNA-208b and MicroRNA-499 Reflect Myocardial Damage in Cardiovascular Disease”
Circ Cardiovasc Genet. 2011;4:e8,
Full TextPDF
Genetic Determinants of Potassium Sensitivity and Hypertension
Integrated Computational and Experimental Analysis of the Neuroendocrine Transcriptome in Genetic Hypertension Identifies Novel Control Points for the Cardiometabolic Syndrome
Ryan S. Friese, PhD, Chun Ye, PhD, Caroline M. Nievergelt, PhD, Andrew J. Schork, BS, Nitish R. Mahapatra, PhD, Fangwen Rao, MD, Philip S. Napolitan, BS, Jill Waalen, MD, MPH, Georg B. Ehret, MD, Patricia B. Munroe, PhD, Geert W. Schmid-Schönbein, PhD, Eleazar Eskin, PhD and Daniel T. O’Connor, MD
Author Affiliations
From the Departments of Bioengineering (R.S.F., G.W.S.-S.), Medicine (R.S.F., A.J.S., F.R., P.S.N., D.T.O.), Pharmacology (D.T.O.), and Psychiatry (C.M.N.), the Bioinformatics Program (C.Y.), and the Institute for Genomic Medicine (D.T.O.), University of California at San Diego; the VA San Diego Healthcare System, San Diego, CA (D.T.O.); the Departments of Computer Science & Human Genetics, University of California at Los Angeles (E.E.); the Department of Biotechnology, Indian Institute of Technology Madras, Chennai, India (N.R.M.); Clinical Pharmacology and The Genome Centre, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, London, United Kingdom (P.B.M.); Center for Complex Disease Genomics, McKusick-Nathans Institute of Genetic Medicine, Johns Hopkins University School of Medicine, Baltimore, MD (G.B.E.); and Scripps Research Institute, La Jolla, CA (J.W.).
Correspondence to Daniel T. O’Connor, MD, Department of Medicine, University of California at San Diego School of Medicine, VASDHS (0838), Skaggs (SSPPS) Room 4256, 9500 Gilman Drive, La Jolla, CA 92093-0838. E-mail doconnor@ucsd.edu
Abstract
Background—Essential hypertension, a common complex disease, displays substantial genetic influence. Contemporary methods to dissect the genetic basis of complex diseases such as the genomewide association study are powerful, yet a large gap exists betweens the fraction of population trait variance explained by such associations and total disease heritability.
Methods and Results—We developed a novel, integrative method (combining animal models, transcriptomics, bioinformatics, molecular biology, and trait-extreme phenotypes) to identify candidate genes for essential hypertension and the metabolic syndrome. We first undertook transcriptome profiling on adrenal glands from blood pressure extreme mouse strains: the hypertensive BPH (blood pressure high) and hypotensive BPL (blood pressure low). Microarray data clustering revealed a striking pattern of global underexpression of intermediary metabolism transcripts in BPH. The MITRA algorithm identified a conserved motif in the transcriptional regulatory regions of the underexpressed metabolic genes, and we then hypothesized that regulation through this motif contributed to the global underexpression. Luciferase reporter assays demonstrated transcriptional activity of the motif through transcription factors HOXA3, SRY, and YY1. We finally hypothesized that genetic variation at HOXA3, SRY, and YY1 might predict blood pressure and other metabolic syndrome traits in humans. Tagging variants for each locus were associated with blood pressure in a human population blood pressure extreme sample with the most extensive associations for YY1 tagging single nucleotide polymorphism rs11625658 on systolic blood pressure, diastolic blood pressure, body mass index, and fasting glucose. Meta-analysis extended the YY1 results into 2 additional large population samples with significant effects preserved on diastolic blood pressure, body mass index, and fasting glucose.
Conclusions—The results outline an innovative, systematic approach to the genetic pathogenesis of complex cardiovascular disease traits and point to transcription factor YY1 as a potential candidate gene involved in essential hypertension and the cardiometabolic syndrome.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 430-440
Published online before print June 5, 2012,
doi: 10.1161/ CIRCGENETICS.111.962415
Genome-Wide Linkage and Positional Candidate Gene Study of Blood Pressure Response to Dietary Potassium Intervention
The Genetic Epidemiology Network of Salt Sensitivity Study
Tanika N. Kelly, PhD, James E. Hixson, PhD, Dabeeru C. Rao, PhD, Hao Mei, MD, PhD, Treva K. Rice, PhD, Cashell E. Jaquish, PhD, Lawrence C. Shimmin, PhD, Karen Schwander, MS, Chung-Shuian Chen, MS, Depei Liu, PhD, Jichun Chen, MD, Concetta Bormans, PhD, Pramila Shukla, MS, Naveed Farhana, MS, Colin Stuart, BS, Paul K. Whelton, MD, MSc, Jiang He, MD, PhD and Dongfeng Gu, MD, PhD
Author Affiliations
From the Department of Epidemiology (T.N.K., H.M., C.-S.C., J.H.), Tulane University School of Public Health and Tropical Medicine, and Department of Medicine (J.H.), Tulane University School of Medicine, New Orleans, La; Department of Epidemiology (J.E.H., L.C.S., C.B., P.S., N.F., C.S.), University of Texas School of Public Health, Houston, Tex; Division of Biostatistics (D.C.R., T.K.R., K.S.), Washington University School of Medicine, St Louis, Mo; Division of Prevention and Population Sciences (C.E.J.), National Heart, Lung, Blood Institute, Bethesda, Md; National Laboratory of Medical Molecular Biology (D.L.), Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China; Cardiovascular Institute and Fuwai Hospital (J.C., D.G.), Chinese Academy of Medical Sciences and Peking Union Medical College and Chinese National Center for Cardiovascular Disease Control and Research, Beijing, China; and Office of the President (P.K.W.), Loyola University Health System and Medical Center, Maywood, Ill.
Correspondence to Dongfeng Gu, MD, PhD, Division of Population Genetics and Prevention, Cardiovascular Institute and Fuwai Hospital, 167 Beilishi Rd, Beijing 100037, China. E-mail gudongfeng@vip.sina.com
Abstract
Background— Genetic determinants of blood pressure (BP) response to potassium, or potassium sensitivity, are largely unknown. We conducted a genome-wide linkage scan and positional candidate gene analysis to identify genetic determinants of potassium sensitivity.
Conclusions— Genetic regions on chromosomes 3 and 11 may harbor important susceptibility loci for potassium sensitivity. Furthermore, the AGTR1 gene was a significant predictor of BP responses to potassium intake.
SOURCE:
Circulation: Cardiovascular Genetics. 2010; 3: 539-547
Published online before print September 22, 2010,
doi: 10.1161/ CIRCGENETICS.110.940635
Genome-Wide Association Study of Cardiac Structure and Systolic Function in African Americans
The Candidate Gene Association Resource (CARe) Study
Ervin R. Fox, MD*, Solomon K. Musani, PhD*, Maja Barbalic, PhD*, Honghuang Lin, PhD, Bing Yu, MS, Kofo O. Ogunyankin, MD, Nicholas L. Smith, PhD, Abdullah Kutlar, MD, Nicole L. Glazer, MD, Wendy S. Post, MD, MS, Dina N. Paltoo, PhD, MPH, Daniel L. Dries, MD, MPH, Deborah N. Farlow, PhD, Christine W. Duarte, PhD, Sharon L. Kardia, PhD, Kristin J. Meyers, PhD, Yan V. Sun, PhD, Donna K. Arnett, PhD, Amit A. Patki, MS, Jin Sha, MS, Xiangqui Cui, PhD, Tandaw E. Samdarshi, MD, MPH, Alan D. Penman, PhD, Kirsten Bibbins-Domingo, MD, PhD, Petra Bůžková, PhD, Emelia J. Benjamin, MD, David A. Bluemke, MD, PhD, Alanna C. Morrison, PhD, Gerardo Heiss, MD, J. Jeffrey Carr, MD, MSc, Russell P. Tracy, PhD, Thomas H. Mosley, PhD, Herman A. Taylor, MD, Bruce M. Psaty, MD, PhD, Susan R. Heckbert, MD, PhD, Thomas P. Cappola, MD, ScM and Ramachandran S. Vasan, MD
Author Affiliations
Guest Editor for this article was Barry London, MD, PhD.
Correspondence to Ervin Fox, MD MPH, FAHA, FACC, Professor of Medicine, Department of Medicine, University of Mississippi Medical Center, 2500 North State St, Jackson, MS 39216. E-mail efox@medicine.umsmed.edu
↵* These authors contributed equally as joint first authors.
Abstract
Background—Using data from 4 community-based cohorts of African Americans, we tested the association between genome-wide markers (single-nucleotide polymorphisms) and cardiac phenotypes in the Candidate-gene Association Resource study.
Methods and Results—Among 6765 African Americans, we related age, sex, height, and weight-adjusted residuals for 9 cardiac phenotypes (assessed by echocardiogram or magnetic resonance imaging) to 2.5 million single-nucleotide polymorphisms genotyped using Genome-wide Affymetrix Human SNP Array 6.0 (Affy6.0) and the remainder imputed. Within the cohort, genome-wide association analysis was conducted, followed by meta-analysis across cohorts using inverse variance weights (genome-wide significance threshold=4.0 ×10−7). Supplementary pathway analysis was performed. We attempted replication in 3 smaller cohorts of African ancestry and tested lookups in 1 consortium of European ancestry (EchoGEN). Across the 9 phenotypes, variants in 4 genetic loci reached genome-wide significance: rs4552931 in UBE2V2 (P=1.43×10−7) for left ventricular mass, rs7213314 in WIPI1 (P=1.68×10−7) for left ventricular internal diastolic diameter, rs1571099 in PPAPDC1A (P=2.57×10−8) for interventricular septal wall thickness, and rs9530176 in KLF5 (P=4.02×10−7) for ejection fraction. Associated variants were enriched in 3 signaling pathways involved in cardiac remodeling. None of the 4 loci replicated in cohorts of African ancestry was confirmed in lookups in EchoGEN.
Conclusions—In the largest genome-wide association study of cardiac structure and function to date in African Americans, we identified 4 genetic loci related to left ventricular mass, interventricular septal wall thickness, left ventricular internal diastolic diameter, and ejection fraction, which reached genome-wide significance. Replication results suggest that these loci may be unique to individuals of African ancestry. Additional large-scale studies are warranted for these complex phenotypes.
SOURCE:
Circulation: Cardiovascular Genetics. 2013; 6: 37-46
Published online before print December 28, 2012,
doi: 10.1161/ CIRCGENETICS.111.962365
Heart and Aging Research in Genomic Epidemiology: 1700 MIs and 2300 coronary heart disease events among about 29 000 eligible patients
Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium
Design of Prospective Meta-Analyses of Genome-Wide Association Studies From 5 Cohorts
Bruce M. Psaty, MD, PhD, Christopher J. O’Donnell, MD, MPH, Vilmundur Gudnason, MD, PhD, Kathryn L. Lunetta, PhD, Aaron R. Folsom, MD, Jerome I. Rotter, MD, André G. Uitterlinden, PhD, Tamara B. Harris, MD, Jacqueline C.M. Witteman, PhD, Eric Boerwinkle, PhD and on Behalf of the CHARGE Consortium
Author Affiliations
From the Cardiovascular Health Research Unit, Departments of Medicine, Epidemiology, and Health Services (B.M.P.), University of Wash; Center for Health Studies, Group Health (B.M.P.), Seattle, Wash; the National Heart, Lung and Blood Institute and the Framingham Heart Study (C.J.O.D.), Framingham, Mass; Icelandic Heart Association and the Department of Cardiovascular Genetics (Y.G.), University of Iceland, Reykjavik, Iceland; Department of Biostatistics (K.L.), Boston University School of Public Health, Mass; Division of Epidemiology and Community Health (A.R.F.), University of Minnesota, Minneapolis; Medical Genetics Institute (J.I.R.), Cedars-Sinai Medical Center, Los Angeles, Calif; Departments of Internal Medicine (A.G.U.) and Epidemiology (A.G.U., J.C.M.W.), Erasmus Medical Center, Rotterdam, The Netherlands; Laboratory of Epidemiology, Demography, and Biometry (T.B.H.), Intramural Research Program, National Institute on Aging, Bethesda, Md; and Human Genetics Center and Division of Epidemiology (E.B.), University of Texas, Houston.
Guest editor for this article was Elizabeth R. Hauser, PhD.
Abstract
Background— The primary aim of genome-wide association studies is to identify novel genetic loci associated with interindividual variation in the levels of risk factors, the degree of subclinical disease, or the risk of clinical disease. The requirement for large sample sizes and the importance of replication have served as powerful incentives for scientific collaboration.
Methods— The Cohorts for Heart and Aging Research in Genomic Epidemiology Consortium was formed to facilitate genome-wide association studies meta-analyses and replication opportunities among multiple large population-based cohort studies, which collect data in a standardized fashion and represent the preferred method for estimating disease incidence. The design of the Cohorts for Heart and Aging Research in Genomic Epidemiology Consortium includes 5 prospective cohort studies from the United States and Europe: the Age, Gene/Environment Susceptibility—Reykjavik Study, the Atherosclerosis Risk in Communities Study, the Cardiovascular Health Study, the Framingham Heart Study, and the Rotterdam Study. With genome-wide data on a total of about 38 000 individuals, these cohort studies have a large number of health-related phenotypes measured in similar ways. For each harmonized trait, within-cohort genome-wide association study analyses are combined by meta-analysis. A prospective meta-analysis of data from all 5 cohorts, with a properly selected level of genome-wide statistical significance, is a powerful approach to finding genuine phenotypic associations with novel genetic loci.
Conclusions— The Cohorts for Heart and Aging Research in Genomic Epidemiology Consortium and collaborating non-member studies or consortia provide an excellent framework for the identification of the genetic determinants of risk factors, subclinical-disease measures, and clinical events.
Example of Coronary Heart Disease
The cohort-study methods papers provide detail about many of the phenotypes listed in Table 2. For coronary heart disease, investigators knowledgeable about the phenotype in each study decided to focus on fatal and nonfatal myocardial infarction (MI) as the primary outcome because the MI criteria differed in only trivial ways among the studies. There were some minor differences in the definition of the composite outcome of MI, fatal coronary heart disease, and sudden death, which became the secondary outcome. Only subjects at risk for an incident event were included in the analysis. MI survivors whose DNA was drawn after the event were not eligible. The primary analysis was restricted to Europeans or European Americans. Patients entered the analysis at the time of the DNA blood draw, and were followed until an event, death, loss to follow up, or the last visit. The main recommendations of the Analysis Committee were adopted, and a threshold of 5×10−8 was selected for genome-wide statistical significance. Analyses in progress include about 1700 MIs and 2300 coronary heart disease events among about 29 000 eligible patients. Each cohort conducted its own analysis, and results were uploaded to a secure share site for the fixed-effects meta-analysis. Even with this number of events (Supplemental Figure 2), power is good for only for relatively high minor allele frequencies (>0.25) and large relative risks (>1.3).
The authors had full access to and take full responsibility for the integrity of the data. All authors have read and agree to the manuscript as written.
Discussion
In thousands of published papers, the 5 CHARGE cohort studies and many of the collaborating studies have already characterized the risk factors for and the incidence and prognosis of a variety of aging-related and cardiovascular conditions. The analysis of the incident MI, for instance, is free from the survival bias typically associated with cross-sectional or case-control studies. The methodologic advantages of the prospective population-based cohort design, the similarity of phenotypes across 5 studies, the availability of genome-wide genotyping data in each cohort, and the need for large sample sizes to provide reliable estimates of genotype-phenotype associations have served as the primary incentives for the formation of the CHARGE consortium, which includes GWAS data on about 38 000 individuals. The consortium effort relies on collaborative methods that are similar to those used by the individual contributing cohorts.
Phenotype experts who know the studies and the data well are responsible for phenotype-standardization across cohorts. The coordinated prospectively planned meta-analyses of CHARGE provide results that are virtually identical to a cohort-adjusted pooled analysis of individual level data. This approach–the within-study analysis followed by a between-study meta-analysis–avoids the human subjects issues associated with individual-level data sharing.
Editors, reviewers, and readers expect replication as the standard in science.6 The finding of a genetic association in one population with evidence for replication in multiple independent populations provides moderate assurance against false-positive reports and helps to establish the validity of the original finding. In a single experiment, the discovery-replication structure is traditionally embodied in a 2-stage design. The CHARGE consortium includes up to 5 independent replicate samples as well as additional collaborating studies for some phenotype working groups, so that it would have been possible to set up analysis plans within CHARGE to mimic the traditional 2-stage design for replication. For instance, the 2 largest cohorts could have served as the discovery set and the others as the replication set. However, attaining the extremely small probability values expected in GWAS requires large sample sizes. For any phenotype, a prospective meta-analysis of all participating cohorts, with a properly selected level of genome-wide statistical significance to minimize the chance of false-positives, is the most powerful approach to finding new genuine associations for genetic loci.25 When findings narrowly miss the prespecified significance threshold, genotyping individuals in other independent populations provides additional evidence about the association. For findings that substantially exceed pre-established significance thresholds, the results of a CHARGE meta-analysis effectively provide evidence of a multistudy replication.
The effort to assemble and manage the CHARGE consortium has provided some interesting and unanticipated challenges. Participating cohorts often had relationships with outside study groups that predated the formation of CHARGE. Timelines for genotyping and imputation have shifted. Purchases of new computer systems for the volume of work were sometimes necessary. Each cohort came to the consortium with their own traditions for methods of analysis, organization, and authorship policies that, while appropriate for their own work, were not always optimal for collaboration with multiple external groups. Within each cohort, the investigators had often formed working groups that divided up the large number of available phenotypes in ways that made sense locally but did not necessarily match the configuration that had been adopted by other cohorts. The Research Steering Committee has attempted to create a set of CHARGE working groups that accommodate the needs and the conventions of the various cohorts. Transparency, disclosure, and professional collaborative behavior by all participating investigators have been essential to the process.
Resource limitations are another challenge. Grant applications that funded the original single-study genome-wide genotyping effort typically imagined a much simpler design. The CHS whole-genome study had as its primary aim, for instance, the analysis of data on 3 endpoints, coronary disease, stroke and heart failure. With a score of active phenotype working groups, the CHARGE collaboration broadened the scope of the short-term work well beyond initial expectations for all the participating cohorts.
One of the premier challenges has been communications among scores of investigators at a dozen sites. CHS and ARIC are themselves multi-site studies. To be successful, the CHARGE collaboration has required effective communications: (1) within each cohort; (2) between cohorts; (3) within the CHARGE working groups; and (4) among the major CHARGE committees. In addition to the traditional methods of conference calls and email, the CHARGE “wiki,” set up by Dr J. Bis (Seattle, Wash), has provided a crucial and highly functional user-driven website for calendars, minutes, guidelines, working group analysis plans, manuscript proposals, and other documents. In the end, there is no substitute for face-to-face meetings, especially at the beginning of the collaboration, and this complex meta-organization has benefited from several CHARGE-wide meetings.
The major emerging opportunity is the collaboration with other studies and consortia. Many working groups have already incorporated nonmember studies into their efforts. Several working groups have coordinated submissions of initial manuscripts with the parallel submission of manuscripts from other studies or consortia. Several working groups have embarked on plans for joint meta-analyses between CHARGE and other consortia. CHARGE has tried to acknowledge and reward the efforts of champions, who assume leadership responsibility for moving these large complex projects forward and who are often hard-working young investigators, the key to the future success of population science.
The CHARGE Consortium represents an innovative model of collaborative research conducted by research teams that know well the strengths, the limitations, and the data from 5 prospective population-based cohort studies. By leveraging the dense genotyping, deep phenotyping and the diverse expertise, prospective meta-analyses are underway to identify and replicate the major common genetic determinants of risk factors, measures of subclinical disease, and clinical events for cardiovascular disease and aging.
SOURCE:
Circulation: Cardiovascular Genetics.2009; 2: 73-80
doi: 10.1161/ CIRCGENETICS.108.829747
Genomics of Ventricular arrhythmias, A-Fib, Right Ventricular Dysplasia, Cardiomyopathy
Comprehensive Desmosome Mutation Analysis in North Americans With Arrhythmogenic Right Ventricular Dysplasia/Cardiomyopathy
A. Dénise den Haan, MD, Boon Yew Tan, MBChB, Michelle N. Zikusoka, MD, Laura Ibañez Lladó, MS, Rahul Jain, MD, Amy Daly, MS, Crystal Tichnell, MGC, Cynthia James, PhD, Nuria Amat-Alarcon, MS, Theodore Abraham, MD, Stuart D. Russell, MD, David A. Bluemke, MD, PhD, Hugh Calkins, MD, Darshan Dalal, MD, PhD and Daniel P. Judge, MD
Author Affiliations
From the Department of Medicine/Cardiology (A.D.d.H., B.Y.T., M.N.Z., L.I.L., R.J., A.D., C.T., C.J., N.A.-A., T.A., S.D.R., H.C., D.D., D.P.J.), Johns Hopkins University School of Medicine, Baltimore, Md; Department of Cardiology, Division of Heart and Lungs (A.D.d.H.), University Medical Center Utrecht, Utrecht, The Netherlands; and National Institutes of Health, Radiology and Imaging Sciences (D.A.B.), Bethesda, Md.
Correspondence to Daniel P. Judge, MD, Johns Hopkins University, Division of Cardiology, Ross 1049; 720 Rutland Avenue, Baltimore, MD 21205. E-mail djudge@jhmi.edu
Abstract
Background— Arrhythmogenic right ventricular dysplasia/cardiomyopathy (ARVD/C) is an inherited disorder typically caused by mutations in components of the cardiac desmosome. The prevalence and significance of desmosome mutations among patients with ARVD/C in North America have not been described previously. We report comprehensive desmosome genetic analysis for 100 North Americans with clinically confirmed or suspected ARVD/C.
Methods and Results— In 82 individuals with ARVD/C and 18 people with suspected ARVD/C, DNA sequence analysis was performed on PKP2, DSG2, DSP, DSC2, and JUP. In those with ARVD/C, 52% harbored a desmosome mutation. A majority of these mutations occurred in PKP2. Notably, 3 of the individuals studied have a mutation in more than 1 gene. Patients with a desmosome mutation were more likely to have experienced ventricular tachycardia (73% versus 44%), and they presented at a younger age (33 versus 41 years) compared with those without a desmosome mutation. Men with ARVD/C were more likely than women to carry a desmosome mutation (63% versus 38%). A mutation was identified in 5 of 18 patients (28%) with suspected ARVD. In this smaller subgroup, there were no significant phenotypic differences identified between individuals with a desmosome mutation compared with those without a mutation.
Conclusions— Our study shows that in 52% of North Americans with ARVD/C a mutation in one of the cardiac desmosome genes can be identified. Compared with those without a desmosome gene mutation, individuals with a desmosome gene mutation had earlier-onset ARVD/C and were more likely to have ventricular tachycardia.
SOURCE:
Circulation: Cardiovascular Genetics.2009; 2: 428-435
Published online before print June 3, 2009,
doi: 10.1161/ CIRCGENETICS.109.858217
Selective Articles citing this article
Genetics of arrhythmogenic right ventricular cardiomyopathy
J. Med. Genet.. 2013;0:jmedgenet-2013-101523v1-jmedgenet-2013-101523,
AbstractFull Text
Genetic complexity in hypertrophic cardiomyopathy revealed by high-throughput sequencing
J. Med. Genet.. 2013;0:jmedgenet-2012-101270v1-jmedgenet-2012-101270,
AbstractFull Text
An Update on Channelopathies: From Mechanisms to Management
Circulation. 2013;127:126-140,
Full TextPDF
Arrhythmogenic Right Ventricular Cardiomyopathy
Circ Arrhythm Electrophysiol. 2012;5:1233-1246,
Full TextPDF
Ventricular arrhythmias associated with long-term endurance sports: what is the evidence?
Br. J. Sports. Med.. 2012;46:i44-i50,
AbstractFull TextPDF
Phospholamban R14del mutation in patients diagnosed with dilated cardiomyopathy or arrhythmogenic right ventricular cardiomyopathy: evidence supporting the concept of arrhythmogenic cardiomyopathy
Eur J Heart Fail. 2012;14:1199-1207,
AbstractFull TextPDF
Assessing the Significance of Pathogenic Mutations and Autopsy Findings in the Light of 2010 Arrhythmogenic Right Ventricular Cardiomyopathy Diagnostic Criteria: A Clinical Challenge
Circ Cardiovasc Genet. 2012;5:384-386,
Full TextPDF
Loss of Cadherin-Binding Proteins {beta}-Catenin and Plakoglobin in the Heart Leads to Gap Junction Remodeling and Arrhythmogenesis
Mol. Cell. Biol.. 2012;32:1056-1067,
AbstractFull TextPDF
Loss of {alpha}T-catenin alters the hybrid adhering junctions in the heart and leads to dilated cardiomyopathy and ventricular arrhythmia following acute ischemia
J. Cell Sci.. 2012;125:1058-1067,
AbstractFull TextPDF
General and Disease-Specific Psychosocial Adjustment in Patients With Arrhythmogenic Right Ventricular Dysplasia/Cardiomyopathy With Implantable Cardioverter Defibrillators: A Large Cohort Study
Circ Cardiovasc Genet. 2012;5:18-24,
AbstractFull TextPDF
Restrictive loss of plakoglobin in cardiomyocytes leads to arrhythmogenic cardiomyopathy
Hum Mol Genet. 2011;20:4582-4596,
AbstractFull TextPDF
Circulation Editors’ Picks: Most Read Articles in Cardiovascular Genetics, Part I
Circulation. 2011;124:e345-e357,
AbstractFull TextPDF
Genetic Variation in Titin in Arrhythmogenic Right Ventricular Cardiomyopathy-Overlap Syndromes
Circulation. 2011;124:876-885,
AbstractFull TextPDF
Circulation: Cardiovascular Genetics Editors’ Picks: A Selection of the Most Important Articles Published
Circulation. 2011;124:e179-e187,
AbstractFull TextPDF
Sudden Cardiac Arrest Without Overt Heart Disease
Circulation. 2011;123:2994-3008,
Full TextPDF
Familial Evaluation in Arrhythmogenic Right Ventricular Cardiomyopathy: Impact of Genetics and Revised Task Force Criteria
Circulation. 2011;123:2701-2709,
AbstractFull TextPDF
Arrhythmogenic Right Ventricular Dysplasia/Cardiomyopathy: Pathogenic Desmosome Mutations in Index-Patients Predict Outcome of Family Screening: Dutch Arrhythmogenic Right Ventricular Dysplasia/Cardiomyopathy Genotype-Phenotype Follow-Up Study
Circulation. 2011;123:2690-2700,
AbstractFull TextPDF
Arrhythmogenic Right Ventricular Dysplasia/Cardiomyopathy: A Family Affair
Circulation. 2011;123:2661-2663,
Full TextPDF
Lack of plakoglobin leads to lethal congenital epidermolysis bullosa: a novel clinico-genetic entity
Hum Mol Genet. 2011;20:1811-1819,
AbstractFull TextPDF
Mechanistic insights into arrhythmogenic right ventricular cardiomyopathy caused by desmocollin-2 mutations
Cardiovasc Res. 2011;90:77-87,
AbstractFull TextPDF
Cardiac Tissue-Restricted Deletion of Plakoglobin Results in Progressive Cardiomyopathy and Activation of {beta}-Catenin Signaling
Mol. Cell. Biol.. 2011;31:1134-1144,
AbstractFull TextPDF
Psychological Issues in Genetic Testing for Inherited Cardiovascular Diseases
Circ Cardiovasc Genet. 2011;4:81-90,
Full TextPDF
De novo desmin-mutation N116S is associated with arrhythmogenic right ventricular cardiomyopathy
Hum Mol Genet. 2010;19:4595-4607,
AbstractFull TextPDF
Wide spectrum of desmosomal mutations in Danish patients with arrhythmogenic right ventricular cardiomyopathy
J. Med. Genet.. 2010;47:736-744,
AbstractFull TextPDF
Dilated Cardiomyopathy With Conduction Disease and Arrhythmia
Circulation. 2010;122:527-534,
Full TextPDF
Lower than expected desmosomal gene mutation prevalence in endurance athletes with complex ventricular arrhythmias of right ventricular origin
Heart. 2010;96:1268-1274, AbstractFull TextPDF
Desmosomal gene analysis in arrhythmogenic right ventricular dysplasia/cardiomyopathy: spectrum of mutations and clinical impact in practice
Europace. 2010;12:861-868,
AbstractFull TextPDF
Large-Scale Candidate Gene Analysis in Whites and African Americans Identifies IL6R Polymorphism in Relation to Atrial Fibrillation
The National Heart, Lung, and Blood Institute’s Candidate Gene Association Resource (CARe) Project
Renate B. Schnabel, MD, MSc*, Kathleen F. Kerr, PhD*, Steven A. Lubitz, MD*, Ermeg L. Alkylbekova, MD*, Gregory M. Marcus, MD, MAS, Moritz F. Sinner, MD, Jared W. Magnani, MD, Philip A. Wolf, MD, Rajat Deo, MD, Donald M. Lloyd-Jones, MD, ScM, Kathryn L. Lunetta, PhD, Reena Mehra, MD, MS, Daniel Levy, MD, Ervin R. Fox, MD, MPH, Dan E. Arking, PhD, Thomas H. Mosley, PhD, Martina Müller-Nurasyid, MSc, PhD, Taylor R. Young, MA, H.-Erich Wichmann, MD, PhD, Sudha Seshadri, MD, Deborah N. Farlow, PhD, Jerome I. Rotter, MD, Elsayed Z. Soliman, MD, MSc, MS, Nicole L. Glazer, PhD, James G. Wilson, MD, Monique M.B. Breteler, MD, Nona Sotoodehnia, MD, MPH, Christopher Newton-Cheh, MD, MPH, Stefan Kääb, MD, PhD, Patrick T. Ellinor, MD, PhD*, Alvaro Alonso, MD*, Emelia J. Benjamin, MD, ScM*, Susan R. Heckbert, MD, PhD* and for the Candidate Gene Association Resource (CARe) Atrial Fibrillation/Electrocardiography Working Group
Correspondence to Susan R. Heckbert, MD, PhD, Cardiovascular Health Research Unit, University of Washington, 1730 Minor Ave, Suite 1360, Seattle, WA 98101. E-mail heckbert@u.washington.edu; Emelia J. Benjamin, MD, ScM, Medicine and Epidemiology, Boston University Schools of Medicine and Public Health, The Framingham Heart Study, 73 Mount Wayte Ave, Framingham, MA 01702–5827. E-mail emelia@bu.edu; Renate B. Schnabel, MD, MSc, Department of Medicine 2, Cardiology, Johannes Gutenberg University, Langenbeckstr 1, 55131 Mainz, Germany. E-mail schnabelr@gmx.de
↵* These authors contributed equally to the manuscript.
Abstract
Background—The genetic background of atrial fibrillation (AF) in whites and African Americans is largely unknown. Genes in cardiovascular pathways have not been systematically investigated.
Methods and Results—We examined a panel of approximately 50 000 common single-nucleotide polymorphisms (SNPs) in 2095 cardiovascular candidate genes and AF in 3 cohorts with participants of European (n=18 524; 2260 cases) or African American descent (n=3662; 263 cases) in the National Heart, Lung, and Blood Institute’s Candidate Gene Association Resource. Results in whites were followed up in the German Competence Network for AF (n=906, 468 cases). The top result was assessed in relation to incident ischemic stroke in the Cohorts for Heart and Aging Research in Genomic Epidemiology Stroke Consortium (n=19 602 whites, 1544 incident strokes). SNP rs4845625 in the IL6R gene was associated with AF (relative risk [RR] C allele, 0.90; 95% confidence interval [CI], 0.85–0.95; P=0.0005) in whites but did not reach statistical significance in African Americans (RR, 0.86; 95% CI, 0.72–1.03; P=0.09). The results were comparable in the German AF Network replication, (RR, 0.71; 95% CI, 0.57–0.89; P=0.003). No association between rs4845625 and stroke was observed in whites. The known chromosome 4 locus near PITX2 in whites also was associated with AF in African Americans (rs4611994; hazard ratio, 1.40; 95% CI, 1.16–1.69; P=0.0005).
Conclusions—In a community-based cohort meta-analysis, we identified genetic association in IL6R with AF in whites. Additionally, we demonstrated that the chromosome 4 locus known from recent genome-wide association studies in whites is associated with AF in African Americans.
SOURCE:
Circulation: Cardiovascular Genetics.2011; 4: 557-564
Published online before print August 16, 2011,
doi: 10.1161/ CIRCGENETICS.110.959197
PITX2c Is Expressed in the Adult Left Atrium, and Reducing Pitx2c Expression Promotes Atrial Fibrillation Inducibility and Complex Changes in Gene Expression
Paulus Kirchhof, MD*, Peter C. Kahr*, Sven Kaese, Ilaria Piccini, PhD, Ismail Vokshi, BSc, Hans-Heinrich Scheld, MD, Heinrich Rotering, MD, Lisa Fortmueller, MD (vet), Sandra Laakmann, MD (vet), Sander Verheule, PhD, Ulrich Schotten, MD, PhD, Larissa Fabritz, MD and Nigel A. Brown, PhD
Author Affiliations
From the Department of Cardiology and Angiology (P.K., P.C.K., S.K., I.P., L.F., S.L., L.F.) and the Department of Thoracic and Cardiovascular Surgery (H.-H.S., H.R.), University Hospital Muenster, Germany; Division of Biomedical Sciences (P.C.K., I.V., N.A.B.), St. George’s, University of London, United Kingdom; and the Department of Physiology (S.V., U.S.), Maastricht University, The Netherlands.
Correspondence to Nigel A. Brown, PhD, Division of Biomedical Sciences, St George’s, University of London, Cranmer Terrace, London, SW17 0RE, UK. E-mail nbrown@sgul.ac.uk
↵* Drs Kirchhof and Kahr contributed equally to this work.
Abstract
Background— Intergenic variations on chromosome 4q25, close to the PITX2 transcription factor gene, are associated with atrial fibrillation (AF). We therefore tested whether adult hearts express PITX2 and whether variation in expression affects cardiac function.
Methods and Results— mRNA for PITX2 isoform c was expressed in left atria of human and mouse, with levels in right atrium and left and right ventricles being 100-fold lower. In mice heterozygous for Pitx2c (Pitx2c+/−), left atrial Pitx2c expression was 60% of wild-type and cardiac morphology and function were not altered, except for slightly elevated pulmonary flow velocity. Isolated Pitx2c+/− hearts were susceptible to AF during programmed stimulation. At short paced cycle lengths, atrial action potential durations were shorter in Pitx2c+/− than in wild-type. Perfusion with the β-receptor agonist orciprenaline abolished inducibility of AF and reduced the effect on action potential duration. Spontaneous heart rates, atrial conduction velocities, and activation patterns were not affected in Pitx2c+/− hearts, suggesting that action potential duration shortening caused wave length reduction and inducibility of AF. Expression array analyses comparing Pitx2c+/− with wild-type, for left atrial and right atrial tissue separately, identified genes related to calcium ion binding, gap and tight junctions, ion channels, and melanogenesis as being affected by the reduced expression of Pitx2c.
Conclusions— These findings demonstrate a physiological role for PITX2 in the adult heart and support the hypothesis that dysregulation of PITX2 expression can be responsible for susceptibility to AF.
SOURCE:
Circulation: Cardiovascular Genetics.2011; 4: 123-133
Published online before print January 31, 2011,
doi: 10.1161/ CIRCGENETICS.110.958058
Genetics of CVD and Hyperlipidemia, Hyper Cholesterolemia, Metabolic Syndrome
Genetic Loci Associated With Plasma Concentration of Low-Density Lipoprotein Cholesterol, High-Density Lipoprotein Cholesterol, Triglycerides, Apolipoprotein A1, and Apolipoprotein B Among 6382 White Women in Genome-Wide Analysis With Replication
Daniel I. Chasman, PhD*, Guillaume Paré, MD, MS*, Robert Y.L. Zee, PhD, MPH, Alex N. Parker, PhD, Nancy R. Cook, ScD, Julie E. Buring, ScD, David J. Kwiatkowski, MD, PhD, Lynda M. Rose, MS, Joshua D. Smith, BS, Paul T. Williams, PhD, Mark J. Rieder, PhD, Jerome I. Rotter, MD, Deborah A. Nickerson, PhD, Ronald M. Krauss, MD, Joseph P. Miletich, MD and Paul M Ridker, MD, MPH
Author Affiliations
From the Center for Cardiovascular Disease Prevention (D.I.C., G.P., R.Y.L.Z., N.R.C., J.E.B., L.M.R., P.M.R.) and Donald W. Reynolds Center for Cardiovascular Research (D.I.C., G.P., R.Y.L.Z., N.R.C., D.J.K., P.M.R.), Brigham and Women’s Hospital, Harvard Medical School, Boston, Mass; Amgen, Inc, Cambridge, Mass (A.N.P., J.M.P.); Department of Genome Sciences, University of Washington, Seattle, Wash (J.D.S., M.J.R., D.A.N.); Life Science Division, Lawrence Berkeley National Laboratory, Berkeley, Calif (P.T.W., R.M.K.); Medical Genetics Institute, Cedars-Sinai Medical Center, Los Angeles, Calif (J.I.R.); and Children’s Hospital Oakland Research Institute, Oakland, Calif (R.M.K.).
Correspondence to Daniel I. Chasman, Center for Cardiovascular Disease Prevention, Brigham and Women’s Hospital, 900 Commonwealth Ave E, Boston, MA 02215. E-mail dchasman@rics.bwh.harvard.edu
Abstract
Background— Genome-wide genetic association analysis represents an opportunity for a comprehensive survey of the genes governing lipid metabolism, potentially revealing new insights or even therapeutic strategies for cardiovascular disease and related metabolic disorders.
Methods and Results— We have performed large-scale, genome-wide genetic analysis among 6382 white women with replication in 2 cohorts of 970 additional white men and women for associations between common single-nucleotide polymorphisms and low-density lipoprotein cholesterol, high-density lipoprotein cholesterol, triglycerides, apolipoprotein (Apo) A1, and ApoB. Genome-wide associations (P<5×10−8) were found at the PCSK9 gene, the APOB gene, the LPL gene, the APOA1-APOA5 locus, the LIPC gene, the CETP gene, the LDLR gene, and the APOE locus. In addition, genome-wide associations with triglycerides at the GCKR gene confirm and extend emerging links between glucose and lipid metabolism. Still other genome-wide associations at the 1p13.3 locus are consistent with emerging biological properties for a region of the genome, possibly related to the SORT1 gene. Below genome-wide significance, our study provides confirmatory evidence for associations at 5 novel loci with low-density lipoprotein cholesterol, high-density lipoprotein cholesterol, or triglycerides reported recently in separate genome-wide association studies. The total proportion of variance explained by common variation at the genome-wide candidate loci ranges from 4.3% for triglycerides to 12.6% for ApoB.
Conclusion— Genome-wide associations at the GCKR gene and near the SORT1 gene, as well as confirmatory associations at 5 additional novel loci, suggest emerging biological pathways for lipid metabolism among white women.
SOURCE:
Circulation: Cardiovascular Genetics.2008; 1: 21-30
doi: 10.1161/ CIRCGENETICS.108.773168
Integrated Computational and Experimental Analysis of the Neuroendocrine Transcriptome in Genetic Hypertension Identifies Novel Control Points for the Cardiometabolic Syndrome
Ryan S. Friese, PhD, Chun Ye, PhD, Caroline M. Nievergelt, PhD, Andrew J. Schork, BS, Nitish R. Mahapatra, PhD, Fangwen Rao, MD, Philip S. Napolitan, BS, Jill Waalen, MD, MPH, Georg B. Ehret, MD, Patricia B. Munroe, PhD, Geert W. Schmid-Schönbein, PhD, Eleazar Eskin, PhD and Daniel T. O’Connor, MD
Author Affiliations
From the Departments of Bioengineering (R.S.F., G.W.S.-S.), Medicine (R.S.F., A.J.S., F.R., P.S.N., D.T.O.), Pharmacology (D.T.O.), and Psychiatry (C.M.N.), the Bioinformatics Program (C.Y.), and the Institute for Genomic Medicine (D.T.O.), University of California at San Diego; the VA San Diego Healthcare System, San Diego, CA (D.T.O.); the Departments of Computer Science & Human Genetics, University of California at Los Angeles (E.E.); the Department of Biotechnology, Indian Institute of Technology Madras, Chennai, India (N.R.M.); Clinical Pharmacology and The Genome Centre, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, London, United Kingdom (P.B.M.); Center for Complex Disease Genomics, McKusick-Nathans Institute of Genetic Medicine, Johns Hopkins University School of Medicine, Baltimore, MD (G.B.E.); and Scripps Research Institute, La Jolla, CA (J.W.).
Correspondence to Daniel T. O’Connor, MD, Department of Medicine, University of California at San Diego School of Medicine, VASDHS (0838), Skaggs (SSPPS) Room 4256, 9500 Gilman Drive, La Jolla, CA 92093-0838. E-mail doconnor@ucsd.edu
Abstract
Background—Essential hypertension, a common complex disease, displays substantial genetic influence. Contemporary methods to dissect the genetic basis of complex diseases such as the genomewide association study are powerful, yet a large gap exists betweens the fraction of population trait variance explained by such associations and total disease heritability.
Methods and Results—We developed a novel, integrative method (combining animal models, transcriptomics, bioinformatics, molecular biology, and trait-extreme phenotypes) to identify candidate genes for essential hypertension and the metabolic syndrome. We first undertook transcriptome profiling on adrenal glands from blood pressure extreme mouse strains: the hypertensive BPH (blood pressure high) and hypotensive BPL (blood pressure low). Microarray data clustering revealed a striking pattern of global underexpression of intermediary metabolism transcripts in BPH. The MITRA algorithm identified a conserved motif in the transcriptional regulatory regions of the underexpressed metabolic genes, and we then hypothesized that regulation through this motif contributed to the global underexpression. Luciferase reporter assays demonstrated transcriptional activity of the motif through transcription factors HOXA3, SRY, and YY1. We finally hypothesized that genetic variation at HOXA3, SRY, and YY1 might predict blood pressure and other metabolic syndrome traits in humans. Tagging variants for each locus were associated with blood pressure in a human population blood pressure extreme sample with the most extensive associations for YY1 tagging single nucleotide polymorphism rs11625658 on systolic blood pressure, diastolic blood pressure, body mass index, and fasting glucose. Meta-analysis extended the YY1 results into 2 additional large population samples with significant effects preserved on diastolic blood pressure, body mass index, and fasting glucose.
Conclusions—The results outline an innovative, systematic approach to the genetic pathogenesis of complex cardiovascular disease traits and point to transcription factor YY1 as a potential candidate gene involved in essential hypertension and the cardiometabolic syndrome.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 430-440
Published online before print June 5, 2012,
doi: 10.1161/ CIRCGENETICS.111.962415
Associations Between Incident Ischemic Stroke Events and Stroke and Cardiovascular Disease-Related Genome-Wide Association Studies Single Nucleotide Polymorphisms in the Population Architecture Using Genomics and Epidemiology Study
Cara L. Carty, PhD, Petra Bůžková, PhD, Myriam Fornage, PhD, Nora Franceschini, MD, Shelley Cole, PhD, Gerardo Heiss, MD, PhD, Lucia A. Hindorff, PhD, MPH, Barbara V. Howard, PhD, Sue Mann, MPH, Lisa W. Martin, MD, Ying Zhang, PhD, Tara C. Matise, PhD, Ross Prentice, PhD, Alexander P. Reiner, MD, MS and Charles Kooperberg, PhD
Author Affiliations
From the Public Health Sciences, Fred Hutchinson Cancer Research Center (C.L.C., S.M., R.P., C.K.); Department of Biostatistics, University of Washington, Seattle, WA (P.B.); Institute of Molecular Medicine, University of Texas Health Sciences Center at Houston, Houston, TX (M.F.); Division of Epidemiology, School of Public Health, University of Texas Health Sciences Center, Houston, TX (M.F.); Department of Epidemiology, University of North Carolina, Chapel Hill, NC (N.F., G.H.); Department of Genetics, Texas Biomedical Research Institute, San Antonio, TX (S.C.); Office of Population Genomics, National Human Genome Research Institute, Bethesda, MD (L.A.H.); Medstar Health Research Institute, Washington, DC (B.V.H.); George Washington University School of Medicine, Washington, DC (B.V.H., L.W.M.); University of Oklahoma Health Sciences Center, Oklahoma City, OK (Y.Z.); Department of Genetics, Rutgers University, Piscataway, NJ (T.C.M.); Department of Epidemiology, University of Washington, Seattle, WA (A.P.R.).
Correspondence to Dr Cara L. Carty, Fred Hutchinson Cancer Research Center, 1100 Fairview Ave N./M3-A410, Seattle, WA 98109. E-mail ccarty@fhcrc.org
Abstract
Background—Genome-wide association studies (GWAS) have identified loci associated with ischemic stroke (IS) and cardiovascular disease (CVD) in European-descent individuals, but their replication in different populations has been largely unexplored.
Methods and Results—Nine single nucleotide polymorphisms (SNPs) selected from GWAS and meta-analyses of stroke, and 86 SNPs previously associated with myocardial infarction and CVD risk factors, including blood lipids (high density lipoprotein [HDL], low density lipoprotein [LDL], and triglycerides), type 2 diabetes, and body mass index (BMI), were investigated for associations with incident IS in European Americans (EA) N=26 276, African-Americans (AA) N=8970, and American Indians (AI) N=3570 from the Population Architecture using Genomics and Epidemiology Study. Ancestry-specific fixed effects meta-analysis with inverse variance weighting was used to combine study-specific log hazard ratios from Cox proportional hazards models. Two of 9 stroke SNPs (rs783396 and rs1804689) were associated with increased IS hazard in AA; none were significant in this large EA cohort. Of 73 CVD risk factor SNPs tested in EA, 2 (HDL and triglycerides SNPs) were associated with IS. In AA, SNPs associated with LDL, HDL, and BMI were significantly associated with IS (3 of 86 SNPs tested). Out of 58 SNPs tested in AI, 1 LDL SNP was significantly associated with IS.
Conclusions—Our analyses showing lack of replication in spite of reasonable power for many stroke SNPs and differing results by ancestry highlight the need to follow up on GWAS findings and conduct genetic association studies in diverse populations. We found modest IS associations with BMI and lipids SNPs, though these findings require confirmation.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 210-216
Published online before print March 8, 2012,
doi: 10.1161/ CIRCGENETICS.111.962191
Common Variation in Fatty Acid Genes and Resuscitation From Sudden Cardiac Arrest
Catherine O. Johnson, PhD, MPH, Rozenn N. Lemaitre, PhD, MPH, Carol E. Fahrenbruch, MSPH, Stephanie Hesselson, PhD, Nona Sotoodehnia, MD, MPH, Barbara McKnight, PhD, Kenneth M. Rice, PhD, Pui-Yan Kwok, MD, PhD, David S. Siscovick, MD, MPH and Thomas D. Rea, MD, MPH
Author Affiliations
From the Departments of Medicine (C.O.J., R.N.L., N.S., D.S.S., T.D.R.), Biostatistics (B.M., K.M.R.), and Epidemiology (D.S.S), University of Washington, Seattle; King County Emergency Medical Services, Seattle, WA (C.E.F.); and Institute of Human Genetics, University of California San Francisco (S.H., P.-Y.K.).
Correspondence to Catherine O. Johnson, PhD, MPH, Department of Medicine, University of Washington, CHRU 1730 Minor Ave, Suite 1360, Seattle, WA 98101. E-mail johnsoco@uw.edu
Abstract
Background—Fatty acids provide energy and structural substrates for the heart and brain and may influence resuscitation from sudden cardiac arrest (SCA). We investigated whether genetic variation in fatty acid metabolism pathways was associated with SCA survival.
Methods and Results—Subjects (mean age, 67 years; 80% male, white) were out-of-hospital SCA patients found in ventricular fibrillation in King County, WA. We compared subjects who survived to hospital admission (n=664) with those who did not (n=689), and subjects who survived to hospital discharge (n=334) with those who did not (n=1019). Associations between survival and genetic variants were assessed using logistic regression adjusting for age, sex, location, time to arrival of paramedics, whether the event was witnessed, and receipt of bystander cardiopulmonary resuscitation. Within-gene permutation tests were used to correct for multiple comparisons. Variants in 5 genes were significantly associated with SCA survival. After correction for multiple comparisons, single-nucleotide polymorphisms in ACSL1 and ACSL3 were significantly associated with survival to hospital admission. Single-nucleotide polymorphisms in ACSL3, AGPAT3, MLYCD, and SLC27A6 were significantly associated with survival to hospital discharge.
Conclusions—Our findings indicate that variants in genes important in fatty acid metabolism are associated with SCA survival in this population.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 422-429
Published online before print June 1, 2012,
doi: 10.1161/ CIRCGENETICS.111.961912
Genome-Wide Association Study Pinpoints a New Functional Apolipoprotein B Variant Influencing Oxidized Low-Density Lipoprotein Levels But Not Cardiovascular Events
AtheroRemo Consortium
Kari-Matti Mäkelä, BM, BSc, Ilkka Seppälä, MSc, Jussi A. Hernesniemi, MD, PhD, Leo-Pekka Lyytikäinen, MD, Niku Oksala, MD, PhD, DSc, Marcus E. Kleber, PhD, Hubert Scharnagl, PhD, Tanja B. Grammer, MD, Jens Baumert, PhD, Barbara Thorand, PhD, Antti Jula, MD, PhD, Nina Hutri-Kähönen, MD, PhD, Markus Juonala, MD, PhD, Tomi Laitinen, MD, PhD, Reijo Laaksonen, MD, PhD, Pekka J. Karhunen, MD, PhD, Kjell C. Nikus, MD, PhD, Tuomo Nieminen, MD, PhD, MSc, Jari Laurikka, MD, PhD, Pekka Kuukasjärvi, MD, PhD, Matti Tarkka, MD, PhD, Jari Viik, PhD, Norman Klopp, PhD, Thomas Illig, PhD, Johannes Kettunen, PhD, Markku Ahotupa, PhD, Jorma S.A. Viikari, MD, PhD, Mika Kähönen, MD, PhD, Olli T. Raitakari, MD, PhD, Mahir Karakas, MD, Wolfgang Koenig, MD, PhD, Bernhard O. Boehm, MD, Bernhard R. Winkelmann, MD, Winfried März, MD and Terho Lehtimäki, MD, PhD
Correspondence to Kari-Matti Mäkelä, Department of Clinical Chemistry, Finn-Medi 2, PO Box 2000, FI-33521 Tampere, Finland. E-mail kari-matti.makela@uta.fi
Abstract
Background—Oxidized low-density lipoprotein may be a key factor in the development of atherosclerosis. We performed a genome-wide association study on oxidized low-density lipoprotein and tested the impact of associated single-nucleotide polymorphisms (SNPs) on the risk factors of atherosclerosis and cardiovascular events.
Methods and Results—A discovery genome-wide association study was performed on a population of young healthy white individuals (N=2080), and the SNPs associated with a P<5×10–8 were replicated in 2 independent samples (A: N=2912; B: N=1326). Associations with cardiovascular endpoints were also assessed with 2 additional clinical cohorts (C: N=1118; and D: N=808). We found 328 SNPs associated with oxidized low-density lipoprotein. The genetic variant rs676210 (Pro2739Leu) in apolipoprotein B was the proxy SNP behind all associations (P=4.3×10–136, effect size=13.2 U/L per allele). This association was replicated in the 2 independent samples (A and B, P=2.5×10–47 and 1.1×10–11, effect sizes=10.3 U/L and 7.8 U/L, respectively). In the meta-analyses of cohorts A, C, and D (excluding cohort B without angiographic data), the top SNP did not associate significantly with the age of onset of angiographically verified coronary artery disease (hazard ratio=1.00 [0.94–1.06] per allele), 3-vessel coronary artery disease (hazard ratio=1.03 [0.94–1.13]), or myocardial infarction (hazard ratio=1.04 [0.96–1.12]).
Conclusions—This novel genetic marker is an important factor regulating oxidized low-density lipoprotein levels but not a major genetic factor for the studied cardiovascular endpoints.
SOURCE:
Circulation: Cardiovascular Genetics.2013; 6: 73-81
Published online before print December 17, 2012,
doi: 10.1161/ CIRCGENETICS.112.964965
Genome-Wide Screen for Metabolic Syndrome Susceptibility Loci Reveals Strong Lipid Gene Contribution But No Evidence for Common Genetic Basis for Clustering of Metabolic Syndrome Traits
Kati Kristiansson, PhD, Markus Perola, MD, PhD, Emmi Tikkanen, MSc, Johannes Kettunen, PhD, Ida Surakka, MSc, Aki S. Havulinna, DSc (Tech.), Alena Stančáková, MD, PhD, Chris Barnes, PhD, Elisabeth Widen, MD, PhD, Eero Kajantie, MD, PhD, Johan G. Eriksson, MD, DMSc, Jorma Viikari, MD, PhD, Mika Kähönen, MD, PhD, Terho Lehtimäki, MD, PhD, Olli T. Raitakari, MD, PhD, Anna-Liisa Hartikainen, MD, PhD, Aimo Ruokonen, MD, PhD, Anneli Pouta, MD, PhD, Antti Jula, MD, PhD, Antti J. Kangas, MSc, Pasi Soininen, PhD, Mika Ala-Korpela, PhD, Satu Männistö, PhD, Pekka Jousilahti, MD, PhD, Lori L. Bonnycastle, PhD, Marjo-Riitta Järvelin, MD, PhD, Johanna Kuusisto, MD, PhD, Francis S. Collins, MD, PhD, Markku Laakso, MD, PhD, Matthew E. Hurles, PhD, Aarno Palotie, MD, PhD, Leena Peltonen, MD, PhD*, Samuli Ripatti, PhD and Veikko Salomaa, MD, PhD
Correspondence to Dr Kati Kristiansson, National Institute for Health and Welfare, University of Helsinki, Biomedicum, PL 104, FI-00251 Helsinki, Finland. E-mail kati.kristiansson@thl.fi
Abstract
Background—Genome-wide association (GWA) studies have identified several susceptibility loci for metabolic syndrome (MetS) component traits, but have had variable success in identifying susceptibility loci to the syndrome as an entity. We conducted a GWA study on MetS and its component traits in 4 Finnish cohorts consisting of 2637 MetS cases and 7927 controls, both free of diabetes, and followed the top loci in an independent sample with transcriptome and nuclear magnetic resonance-based metabonomics data. Furthermore, we tested for loci associated with multiple MetS component traits using factor analysis, and built a genetic risk score for MetS.
Methods and Results—A previously known lipid locus, APOA1/C3/A4/A5 gene cluster region (SNP rs964184), was associated with MetS in all 4 study samples (P=7.23×10−9 in meta-analysis). The association was further supported by serum metabolite analysis, where rs964184 was associated with various very low density lipoprotein, triglyceride, and high-density lipoprotein metabolites (P=0.024–1.88×10−5). Twenty-two previously identified susceptibility loci for individual MetS component traits were replicated in our GWA and factor analysis. Most of these were associated with lipid phenotypes, and none with 2 or more uncorrelated MetS components. A genetic risk score, calculated as the number of risk alleles in loci associated with individual MetS traits, was strongly associated with MetS status.
Conclusions—Our findings suggest that genes from lipid metabolism pathways have the key role in the genetic background of MetS. We found little evidence for pleiotropy linking dyslipidemia and obesity to the other MetS component traits, such as hypertension and glucose intolerance.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 242-249
Published online before print March 7, 2012,
doi: 10.1161/ CIRCGENETICS.111.961482
Genetics and Vascular Pathologies and Platelet Aggregation, Cardiac Troponin T in Serum
TGFβRIIb Mutations Trigger Aortic Aneurysm Pathogenesis by Altering Transforming Growth Factor β2 Signal Transduction
Katharine J. Bee, PhD, David C. Wilkes, PhD, Richard B. Devereux, MD, Craig T. Basson, MD, PhD and Cathy J. Hatcher, PhD
Author Affiliations
From the Center for Molecular Cardiology, Greenberg Division of Cardiology, Weill Cornell Medical College, New York, NY.
Correspondence to Cathy J. Hatcher, PhD, Greenberg Division of Cardiology, Weill Cornell Medical College, 525 E. 68th St, New York, NY 10065. E-mail cjhatche@med.cornell.edu
Abstract
Background—Thoracic aortic aneurysm (TAA) is a common progressive disorder involving gradual dilation of the ascending and/or descending thoracic aorta that eventually leads to dissection or rupture. Nonsydromic TAA can occur as a genetically triggered, familial disorder that is usually transmitted in a monogenic autosomal dominant fashion and is known as familial TAA. Genetic analyses of families affected with TAA have identified several chromosomal loci, and further mapping of familial TAA genes has highlighted disease-causing mutations in at least 4 genes: myosin heavy chain 11 (MYH11), α-smooth muscle actin (ACTA2), and transforming growth factor β receptors I and II (TGFβRI and TGFβRII).
Methods and Results—We evaluated 100 probands to determine the mutation frequency in MYH11, ACTA2, TGFβRI, and TGFβRII in an unbiased population of individuals with genetically mediated TAA. In this study, 9% of patients had a mutation in one of the genes analyzed, 3% of patients had mutations in ACTA2, 3% in MYH11, 1% in TGFβRII, and no mutations were found in TGFβRI. Additionally, we identified mutations in a 75 base pair alternatively spliced TGFβRII exon, exon 1a that produces the TGFβRIIb isoform and accounted for 2% of patients with mutations. Our in vitro analyses indicate that the TGFβRIIb activating mutations alter receptor function on TGFβ2 signaling.
Conclusions—We propose that TGFβRIIb expression is a regulatory mechanism for TGFβ2 signal transduction. Dysregulation of the TGFβ2 signaling pathway, as a consequence of TGFβRIIb mutations, results in aortic aneurysm pathogenesis.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 621-629
Published online before print October 24, 2012,doi: 10.1161/CIRCGENETICS.112.964064
Matrix Metalloproteinase-9 Genotype as a Potential Genetic Marker for Abdominal Aortic Aneurysm
Tyler Duellman, BS, Christopher L. Warren, PhD, Peggy Peissig, PhD, Martha Wynn, MD and Jay Yang, MD, PhD
Author Affiliations
From the Molecular and Cellular Pharmacology Graduate Program (T.D., J.Y.) and Department of Anesthesiology (M.W., J.Y.), University of Wisconsin School of Medicine and Public Health, Madison; Illumavista Biosciences LLC, Madison, WI (C.L.W.); and Biomedical Informatics Research Center, Marshfield Clinics Research Foundation, Marshfield, WI (P.P.).
Correspondence to Jay Yang, MD, PhD, Department of Anesthesiology, University of Wisconsin SMPH, SMI 301, 1300 University Ave, Madison, WI 53706. E-mail Jyang75@wisc.edu
Abstract
Background—Degradation of extracellular matrix support in the large abdominal arteries contribute to abnormal dilation of aorta, leading to abdominal aortic aneurysms, and matrix metalloproteinase-9 (MMP-9) is the predominant enzyme targeting elastin and collagen present in the walls of the abdominal aorta. Previous studies have suggested a potential association between MMP-9 genotype and abdominal aortic aneurysm, but these studies have been limited only to the p-1562 and (CA) dinucleotide repeat microsatellite polymorphisms in the promoter region of the MMP-9 gene. We determined the functional alterations caused by 15 MMP-9 single-nucleotide polymorphisms (SNPs) reported to be relatively abundant in the human genome through Western blots, gelatinase, and promoter–reporter assays and incorporated this information to perform a logistic-regression analysis of MMP-9 SNPs in 336 human abdominal aortic aneurysm cases and controls.
Methods and Results—Significant functional alterations were observed for 6 exon SNPs and 4 promoter SNPs. Genotype analysis of frequency-matched (age, sex, history of hypertension, hypercholesterolemia, and smoking) cases and controls revealed significant genetic heterogeneity exceeding 20% observed for 6 SNPs in our population of mostly white subjects from Northern Wisconsin. A step-wise logistic-regression analysis with 6 functional SNPs, where weakly contributing confounds were eliminated using Akaike information criteria, gave a final 2 SNP (D165N and p-2502) model with an overall odds ratio of 2.45 (95% confidence interval, 1.06–5.70).
Conclusions—The combined approach of direct experimental confirmation of the functional alterations of MMP-9 SNPs and logistic-regression analysis revealed significant association between MMP-9 genotype and abdominal aortic aneurysm.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 529-537
Published online before print August 31, 2012,
doi: 10.1161/ CIRCGENETICS.112.963082
Common Genetic Variation in the 3′–BCL11B Gene Desert Is Associated With Carotid-Femoral Pulse Wave Velocity and Excess Cardiovascular Disease Risk
The AortaGen Consortium
Gary F. Mitchell, MD*, Germaine C. Verwoert, MSc*, Kirill V. Tarasov, MD, PhD*, Aaron Isaacs, PhD, Albert V. Smith, PhD, Yasmin, BSc, MA, PhD, Ernst R. Rietzschel, MD, PhD, Toshiko Tanaka, PhD, Yongmei Liu, MD, PhD, Afshin Parsa, MD, MPH, Samer S. Najjar, MD, Kevin M. O’Shaughnessy, MA, BM, DPhil, FRCP, Sigurdur Sigurdsson, MSc, Marc L. De Buyzere, MSc, Martin G. Larson, ScD, Mark P.S. Sie, MD, PhD, Jeanette S. Andrews, MS, Wendy S. Post, MD, MS, Francesco U.S. Mattace-Raso, MD, PhD, Carmel M. McEniery, BSc, PhD, Gudny Eiriksdottir, MSc, Patrick Segers, PhD, Ramachandran S. Vasan, MD, Marie Josee E. van Rijn, MD, PhD, Timothy D. Howard, PhD, Patrick F. McArdle, PhD, Abbas Dehghan, MD, PhD, Elizabeth S. Jewell, MS, Stephen J. Newhouse, MSc, PhD, Sofie Bekaert, PhD, Naomi M. Hamburg, MD, Anne B. Newman, MD, MPH, Albert Hofman, MD, PhD, Angelo Scuteri, MD, PhD, Dirk De Bacquer, PhD, Mohammad Arfan Ikram, MD, PhD†, Bruce M. Psaty, MD, PhD†, Christian Fuchsberger, PhD‡, Matthias Olden, PhD‡, Louise V. Wain, PhD§, Paul Elliott, MB, PhD§, Nicholas L. Smith, PhD‖, Janine F. Felix, MD, PhD‖, Jeanette Erdmann, PhD¶, Joseph A. Vita, MD, Kim Sutton-Tyrrell, PhD, Eric J.G. Sijbrands, MD, PhD, Serena Sanna, PhD, Lenore J. Launer, MS, PhD, Tim De Meyer, PhD, Andrew D. Johnson, MD, Anna F.C. Schut, MD, PhD, David M. Herrington, MD, MHS, Fernando Rivadeneira, MD, PhD, Manuela Uda, PhD, Ian B. Wilkinson, MA, BM, FRCP, Thor Aspelund, PhD, Thierry C. Gillebert, MD, PhD, Luc Van Bortel, MD, PhD, Emelia J. Benjamin, MD, MSc, Ben A. Oostra, PhD, Jingzhong Ding, MD, PhD, Quince Gibson, MBA, André G. Uitterlinden, PhD, Gonçalo R. Abecasis, PhD, John R. Cockcroft, BSc, MB, ChB, FRCP, Vilmundur Gudnason, MD, PhD, Guy G. De Backer, MD, PhD, Luigi Ferrucci, MD, Tamara B. Harris, MD, MS, Alan R. Shuldiner, MD, Cornelia M. van Duijn, PhD, Daniel Levy, MD*, Edward G. Lakatta, MD* and Jacqueline C.M. Witteman, PhD*
Correspondence to Gary F. Mitchell, MD, Cardiovascular Engineering, Inc, 1 Edgewater Dr, Suite 201A, Norwood, MA 02062. E-mail GaryFMitchell@mindspring.com
↵* These authors contributed equally.
Abstract
Background—Carotid-femoral pulse wave velocity (CFPWV) is a heritable measure of aortic stiffness that is strongly associated with increased risk for major cardiovascular disease events.
Methods and Results—We conducted a meta-analysis of genome-wide association data in 9 community-based European ancestry cohorts consisting of 20 634 participants. Results were replicated in 2 additional European ancestry cohorts involving 5306 participants. Based on a preliminary analysis of 6 cohorts, we identified a locus on chromosome 14 in the 3′-BCL11B gene desert that is associated with CFPWV (rs7152623, minor allele frequency=0.42, β=−0.075±0.012 SD/allele, P=2.8×10−10; replication β=−0.086±0.020 SD/allele, P=1.4×10−6). Combined results for rs7152623 from 11 cohorts gave β=−0.076±0.010 SD/allele, P=3.1×10−15. The association persisted when adjusted for mean arterial pressure (β=−0.060±0.009 SD/allele, P=1.0×10−11). Results were consistent in younger (<55 years, 6 cohorts, n=13 914, β=−0.081±0.014 SD/allele, P=2.3×10−9) and older (9 cohorts, n=12 026, β=−0.061±0.014 SD/allele, P=9.4×10−6) participants. In separate meta-analyses, the locus was associated with increased risk for coronary artery disease (hazard ratio=1.05; confidence interval=1.02–1.08; P=0.0013) and heart failure (hazard ratio=1.10, CI=1.03–1.16, P=0.004).
Conclusions—Common genetic variation in a locus in the BCL11B gene desert that is thought to harbor 1 or more gene enhancers is associated with higher CFPWV and increased risk for cardiovascular disease. Elucidation of the role this novel locus plays in aortic stiffness may facilitate development of therapeutic interventions that limit aortic stiffening and related cardiovascular disease events.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 81-90
Published online before print November 8, 2011,
doi: 10.1161/ CIRCGENETICS.111.959817
Genetic Variation in PEAR1 is Associated with Platelet Aggregation and Cardiovascular Outcomes
Joshua P. Lewis1, Kathleen Ryan1, Jeffrey R. O’Connell1, Richard B. Horenstein1, Coleen M. Damcott1, Quince Gibson1, Toni I. Pollin1, Braxton D. Mitchell1, Amber L. Beitelshees1, Ruth Pakzy1, Keith Tanner1, Afshin Parsa1, Udaya S. Tantry2, Kevin P. Bliden2, Wendy S. Post3, Nauder Faraday3, William Herzog4, Yan Gong5, Carl J. Pepine6, Julie A. Johnson5, Paul A. Gurbel2 and Alan R. Shuldiner7*
Author Affiliations
1University of Maryland School of Medicine, Baltimore, MD
2Sinai Hospital of Baltimore, Baltimore, MD
3Johns Hopkins University School of Medicine, Baltimore, MD
4Sinai Hospital of Baltimore & Johns Hopkins University School of Medicine, Baltimore, MD
5University of Florida College of Pharmacy, Gainesville, FL
6University of Florida College of Medicine, Gainesville, FL
7University of Maryland School of Medicine & Veterans Administration Medical Center, Baltimore, MD
↵* University of Maryland School of Medicine & Veterans Administration Medical Center, Baltimore, MD ashuldin@medicine.umaryland.edu
Abstract
Background-Aspirin or dual antiplatelet therapy (DAPT) with aspirin and clopidogrel is standard therapy for patients at increased risk for cardiovascular events. However, the genetic determinants of variable response to aspirin (alone and in combination with clopidogrel) are not known.
Methods and Results-We measured ex-vivo platelet aggregation before and after DAPT in individuals (n=565) from the Pharmacogenomics of Antiplatelet Intervention (PAPI) Study and conducted a genome-wide association study (GWAS) of drug response. Significant findings were extended by examining genotype and cardiovascular outcomes in two independent aspirin-treated cohorts: 227 percutaneous coronary intervention (PCI) patients, and 1,000 patients of the International VErapamil SR/trandolapril Study (INVEST) GENEtic Substudy (INVEST-GENES). GWAS revealed a strong association between single nucleotide polymorphisms on chromosome 1q23 and post-DAPT platelet aggregation. Further genotyping revealed rs12041331 in the platelet endothelial aggregation receptor-1 (PEAR1) gene to be most strongly associated with DAPT response (P=7.66×10-9). In Caucasian and African American patients undergoing PCI, A-allele carriers of rs12041331 were more likely to experience a cardiovascular event or death compared to GG homozygotes (hazard ratio = 2.62, 95%CI 0.96-7.10, P=0.059 and hazard ratio = 3.97, 95%CI 1.10-14.31, P=0.035 respectively). In aspirin-treated INVEST-GENES patients, rs12041331 A-allele carriers had significantly increased risk of myocardial infarction compared to GG homozygotes (OR=2.03, 95%CI 1.01-4.09, P=0.048).
Conclusions-Common genetic variation in PEAR1 may be a determinant of platelet response and cardiovascular events in patients on aspirin, alone and in combination with clopidogrel.
Clinical Trial Registration Information-clinicaltrials.gov; Identifiers: NCT00799396 and NCT00370045
SOURCE:
CIRCGENETICS.112.964627
Published online before print February 7, 2013,
doi: 10.1161/ CIRCGENETICS.111.964627
Association of Genome-Wide Variation With Highly Sensitive Cardiac Troponin-T Levels in European Americans and Blacks
A Meta-Analysis From Atherosclerosis Risk in Communities and Cardiovascular Health Studies
Bing Yu, MD, MSc, Maja Barbalic, PhD, Ariel Brautbar, MD, Vijay Nambi, MD, Ron C. Hoogeveen, PhD, Weihong Tang, PhD, Thomas H. Mosley, PhD, Jerome I. Rotter, MD, Christopher R. deFilippi, MD, Christopher J. O’Donnell, MD, Sekar Kathiresan, MD, Ken Rice, PhD, Susan R. Heckbert, MD, PhD, Christie M. Ballantyne, MD, Bruce M. Psaty, MD, PhD and Eric Boerwinkle, PhD on behalf of the CARDIoGRAM Consortium
Author Affiliations
From the Human Genetic Center, University of Texas Health Science Center at Houston, Houston, TX (B.Y., M.B., E.B.); Deptartment of Medicine (A.B., V.N., R.C.H., C.M.B.), and Human Genome Sequencing Center (E.B.), Baylor College of Medicine, Houston, TX; Department of Epidemiology, University of Minnesota, Minneapolis, MN (W.T.); Division of Geriatrics, University of Mississippi Medical Center, Jackson, MS (T.H.M.); Medical Genetics Institute, Cedars-Sinai Medical Center, Los Angeles, CA (J.I.R.); School of Medicine, University of Maryland, Baltimore, MD (C.R.D.); National Heart, Lung, and Blood Institute and Framingham Heart Study, National Institutes of Health, Bethesda, MD (C.J.O.D.); Center for Human Genetic Research & Cardiovascular Research Center, Massachusetts General Hospital and Department of Medicine, Harvard Medical School, Boston, MA (S.K.); Department of Biostatistics (K.R.), and Cardiovascular Health Research Unit & Department of Epidemiology (S.R.H.), University of Washington, Seattle, WA; and Cardiovascular Health Research Unit, Departments of Medicine, Epidemiology, and Health Services, University of Washington & Group Health Research Institute, Group Health Cooperative, Seattle, WA (B.M.P.).
Correspondence to Eric Boerwinkle, PhD, Human Genetic Center, University of Texas School of Public Health, 1200 Herman Pressler E-447, Houston, TX 77030. E-mail Eric.Boerwinkle@uth.tmc.edu
Abstract
Background—High levels of cardiac troponin T, measured by a highly sensitive assay (hs-cTnT), are strongly associated with incident coronary heart disease and heart failure. To date, no large-scale genome-wide association study of hs-cTnT has been reported. We sought to identify novel genetic variants that are associated with hs-cTnT levels.
Methods and Results—We performed a genome-wide association in 9491 European Americans and 2053 blacks free of coronary heart disease and heart failure from 2 prospective cohorts: the Atherosclerosis Risk in Communities Study and the Cardiovascular Health Study. Genome-wide association studies were conducted in each study and race stratum. Fixed-effect meta-analyses combined the results of linear regression from 2 cohorts within each race stratum and then across race strata to produce overall estimates and probability values. The meta-analysis identified a significant association at chromosome 8q13 (rs10091374; P=9.06×10−9) near the nuclear receptor coactivator 2 (NCOA2) gene. Overexpression of NCOA2 can be detected in myoblasts. An additional analysis using logistic regression and the clinically motivated 99th percentile cut point detected a significant association at 1q32 (rs12564445; P=4.73×10−8) in the gene TNNT2, which encodes the cardiac troponin T protein itself. The hs-cTnT-associated single-nucleotide polymorphisms were not associated with coronary heart disease in a large case-control study, but rs12564445 was significantly associated with incident heart failure in Atherosclerosis Risk in Communities Study European Americans (hazard ratio=1.16; P=0.004).
Conclusions—We identified 2 loci, near NCOA2 and in the TNNT2 gene, at which variation was significantly associated with hs-cTnT levels. Further use of the new assay should enable replication of these results.
SOURCE:
Circulation: Cardiovascular Genetics.2013; 6: 82-88
Published online before print December 16, 2012,
doi: 10.1161/ CIRCGENETICS.112.963058
Genomics and Valvular Disease
Supravalvular Aortic Stenosis Elastin Arteriopathy
Giuseppe Merla, PhD, Nicola Brunetti-Pierri, MD, Pasquale Piccolo, PhD, Lucia Micale, PhD and Maria Nicla Loviglio, PhD, MSc
Author Affiliations
From the Medical Genetics Unit, IRCCS Casa Sollievo Della Sofferenza Hospital, San Giovanni Rotondo, Italy (G.M., L.M., M.N.L.); Telethon Institute of Genetics and Medicine, Napoli, Italy (N.B-P., P.P.); Department of Pediatrics, Federico II University of Naples, Naples, Italy (N.B-P.); and CIG Center for Integrative Genomics, University of Lausanne, Lausanne, Switzerland (M.N.L.).
Correspondence to Giuseppe Merla, PhD, Medical Genetics Unit, IRCCS Casa Sollievo della Sofferenza, viale Cappuccini, 71013 San Giovanni Rotondo, Italy. E-mail g.merla@operapadrepio.it
Abstract
Supravalvular aortic stenosis is a systemic elastin (ELN) arteriopathy that disproportionately affects the supravalvular aorta. ELN arteriopathy may be present in a nonsyndromic condition or in syndromic conditions such as Williams–Beuren syndrome. The anatomic findings include congenital narrowing of the lumen of the aorta and other arteries, such as branches of pulmonary or coronary arteries. Given the systemic nature of the disease, accurate evaluation is recommended to establish the degree and extent of vascular involvement and to plan appropriate interventions, which are indicated whenever hemodynamically significant stenoses occur. ELN arteriopathy is genetically heterogeneous and occurs as a consequence of haploinsufficiency of the ELN gene on chromosome 7q11.23, owing to either microdeletion of the entire chromosomal region or ELN point mutations. Interestingly, there is a prevalence of premature termination mutations resulting in null alleles among ELN point mutations. The identification of the genetic defect in patients with supravalvular aortic stenosis is essential for a definitive diagnosis, prognosis, and genetic counseling.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 692-696
doi: 10.1161/ CIRCGENETICS.112.962860
Genetic Loci for Coronary Calcification and Serum Lipids Relate to Aortic and Carotid Calcification
Daniel Bos, MD, M. Arfan Ikram, MD, PhD, Aaron Isaacs, PhD, Benjamin F.J. Verhaaren, MD, Albert Hofman, MD, PhD, Cornelia M. van Duijn, PhD, Jacqueline C.M. Witteman, PhD, Aad van der Lugt, MD, PhD and Meike W. Vernooij, MD, PhD
Author Affiliations
From the Departments of Radiology (D.B., M.A.I., B.F.J.V., A.v.d.L., M.W.V), Epidemiology (D.B., M.A.I., A.I., B.F.J.V., A.H., C.M.v.D., J.C.M.W., M.W.V.), and Genetic Epidemiology Unit (A.I., C.M.v.D.), Erasmus MC, Rotterdam, the Netherlands.
Correspondence to Meike W. Vernooij, MD, PhD, Department of Radiology, Erasmus MC, Gravendijkwal 230, PO Box 2040, 3000CA Rotterdam, the Netherlands. E-mail m.vernooij@erasmusmc.nl
Abstract
Background—Atherosclerosis in different vessel beds shares lifestyle and environmental risk factors. It is unclear whether this holds for genetic risk factors. Hence, for the current study genetic loci for coronary artery calcification and serum lipid levels, one of the strongest risk factors for atherosclerosis, were used to assess their relation with atherosclerosis in different vessel beds.
Methods and Results—From 1987 persons of the population-based Rotterdam Study, 3 single-nucleotide polymorphisms (SNPs) for coronary artery calcification and 132 SNPs for total cholesterol, low-density lipoprotein cholesterol, high-density lipoprotein cholesterol or triglycerides were used. To quantify atherosclerotic calcification as a marker of atherosclerosis, all participants underwent nonenhanced computed tomography of the aortic arch and carotid arteries. Associations between genetic risk scores of the joint effect of the SNPs and of all calcification were investigated. The joint effect of coronary artery calcification–SNPs was associated with larger calcification volumes in all vessel beds (difference in calcification volume per SD increase in genetic risk score: 0.15 [95% confidence interval, 0.11–0.20] in aorta, 0.14 [95% confidence interval, 0.10–0.18] in extracranial carotids, and 0.11 [95% confidence interval, 0.07–0.16] in intracranial carotids). The joint effect of total cholesterol SNPs, low-density lipoprotein SNPs, and of all lipid SNPs together was associated with larger calcification volumes in both the aortic arch and the carotid arteries but attenuated after adjusting for the lipid fraction and lipid-lowering medication.
Conclusions—The genetic basis for aortic arch and carotid artery calcification overlaps with the most important loci of coronary artery calcification. Furthermore, serum lipids share a genetic predisposition with both calcification in the aortic arch and the carotid arteries, providing novel insights into the cause of atherosclerosis.
SOURCE:
Circulation: Cardiovascular Genetics.2013; 6: 47-53
Published online before print December 16, 2012,
doi: 10.1161/ CIRCGENETICS.112.963934
Joint Associations of 61 Genetic Variants in the Nicotinic Acetylcholine Receptor Genes with Subclinical Atherosclerosis in American Indians
A Gene-Family Analysis
Jingyun Yang, PhD*, Yun Zhu, MS*, Elisa T. Lee, PhD, Ying Zhang, PhD, Shelley A. Cole, PhD, Karin Haack, PhD, Lyle G. Best, BS MD, Richard B. Devereux, MD, Mary J. Roman, MD, Barbara V. Howard, PhD and Jinying Zhao, MD, PhD
Author Affiliations
From the Tulane University School of Public Health and Tropical Medicine, New Orleans, LA (J.Y., Y. Zhu, J.Z.); Center for American Indian Health Research, University of Oklahoma Health Sciences Center, Oklahoma City, OK (E.T.L., Y. Zhang); Texas Biomedical Research Institute, San Antonio, TX (S.A.C., K.H.); Missouri Breaks Industries Research Inc, Timber Lake, SD (L.G.B.); The New York Hospital-Cornell Medical Center, New York, NY (R.B.D., M.J.R.); MedStar Health Research Institute, Hyattsville, MD (B.V.H.); and Georgetown and Howard Universities Centers for Translational Sciences, Washington, DC (B.V.H.).
Correspondence to Jinying Zhao, MD, PhD, Department of Epidemiology, School of Public Health and Tropical Medicine, Tulane University, 1440 Canal St, SL18, New Orleans, LA 70112. E-mail jzhao5@tulane.edu
↵* These authors contributed equally to this work.
Abstract
Background—Atherosclerosis is the underlying cause of cardiovascular disease, the leading cause of morbidity and mortality in all American populations, including American Indians. Genetic factors play an important role in the pathogenesis of atherosclerosis. Although a single-nucleotide polymorphism (SNP) may explain only a small portion of variability in disease, the joint effect of multiple variants in a pathway on disease susceptibility could be large.
Methods and Results—Using a gene-family analysis, we investigated the joint associations of 61 tag SNPs in 7 nicotinic acetylcholine receptor genes with subclinical atherosclerosis, as measured by carotid intima-media thickness and plaque score, in 3665 American Indians from 94 families recruited by the Strong Heart Family Study (SHFS). Although multiple SNPs showed marginal association with intima-media thickness and plaque score individually, only a few survived adjustments for multiple testing. However, simultaneously modeling of the joint effect of all 61 SNPs in 7 nicotinic acetylcholine receptor genes revealed significant association of the nicotinic acetylcholine receptor gene family with both intima-media thickness and plaque score independent of known coronary risk factors.
Conclusions—Genetic variants in the nicotinic acetylcholine receptor gene family jointly contribute to subclinical atherosclerosis in American Indians who participated in the SHFS. These variants may influence the susceptibility of atherosclerosis through pathways other than cigarette smoking per se.
SOURCE:
Circulation: Cardiovascular Genetics.2013; 6: 89-96
Published online before print December 22, 2012,
doi: 10.1161/ CIRCGENETICS.112.963967
Heredity of Cardiovascular Disorders Inheritance
A Clinical Approach to Common Cardiovascular Disorders When There Is a Family History
The Implications of Inheritance for Clinical Management
Srijita Sen-Chowdhry, MBBS, MD, FESC, Daniel Jacoby, MD and William J. McKenna, MD, DSc, FESC
Author Affiliations
From the Institute of Cardiovascular Science, University College London, London, United Kingdom (S.S-C., W.J.M.); Department of Epidemiology, Imperial College, London, London, United Kingdom (S.S-C.); Division of Cardiology, Yale School of Medicine, New Haven, CT (D.J., W.J.M.).
Correspondence to Professor William J. McKenna, MD, DSc, FESC, Institute of Cardiovascular Science, University College London, The Heart Hospital, 16-18 Westmoreland Street, London, E-mail william.mckenna@uclh.nhs.uk
Introduction
Since the advent of genotyping, recognition of heritable disease has been perceived as an opportunity for genetic diagnosis or new gene identification studies to advance understanding of pathogenesis. Until recently, however, clinical application of DNA-based testing was confined largely to Mendelian disorders. Even within this remit, predictive testing of relatives is cost-effective only in diseases in which the majority of families harbor mutations in known causal genes, such as adult polycystic kidney disease and hypertrophic cardiomyopathy, but not dilated cardiomyopathy. Confirmatory genetic testing of index cases with borderline clinical features may be economic in the still smaller subset of diseases with limited locus heterogeneity, such as Marfan syndrome. Furthermore, Mendelian diseases account for ≈5% of total disease burden.1 Genome-wide association studies have made headway in elucidating the genetic contribution to the more common, complex diseases, and high throughput techniques promise to facilitate integration of genetic analysis into clinical practice. Nevertheless, many genes remain to be identified and implementation of genomic profiling as a population screening tool would not be cost-effective at present. The implications of heredity, however, extend beyond serving as a platform for genetic analysis, influencing diagnosis, prognostication, and treatment of both index cases and relatives, and enabling rational targeting of genotyping resources. This review covers acquisition of a family history, evaluation of heritability and inheritance patterns, and the impact of inheritance on subsequent components of the clinical pathway.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 467-476
doi: 10.1161/ CIRCGENETICS.110.959361
Clinical Considerations of Heritable Factors in Common Heart Failure
Thomas P. Cappola, MD, ScM and Gerald W. Dorn II, MD
Author Affiliations
From the Department of Medicine, University of Pennsylvania, Philadelphia, PA (T.P.C.), and Center for Pharmacogenomics, Washington University School of Medicine, St Louis, MO (G.W.D.II.).
Correspondence to Gerald W. Dorn II, MD, Center for Pharmacogenomics, Washington University, 660 S Euclid Ave, Campus Box 8220, St Louis, MO 63110. E-mail gdorn@dom.wustl.edu
Introduction
Heart failure is a common condition responsible for at least 290 000 deaths each year in the United States alone.1 A small minority of heart failure cases are attributed to Mendelian or familial cardiomyopathies. The majority of systolic heart failure cases are not familial but represent the end result of 1 or many conditions that primarily injure the myocardium sufficiently to diminish cardiac output in the absence of compensatory mechanisms. Paradoxically, because they also injure the myocardium, it is the chronic actions of the compensatory mechanisms that in many instances contribute to the progression from simple cardiac injury to dilated cardiomyopathy and overt heart failure. Thus, the epidemiology of common heart failure appears to be just as sporadic as its major antecedent conditions (atherosclerosis, diabetes, hypertension, and viral myocarditis).
Familial trends in preclinical cardiac remodeling2 and risk of developing heart failure3 reveal an important role for genetic modifiers in addition to clinical and environmental factors. Candidate gene studies performed over the past 10 years have identified a few polymorphic gene variants that modify risk or progression of common heart failure.4 Whole-genome sequencing will lead to the discovery of other genetic modifiers that were not candidates.5 The imminent availability of individual whole-genome sequences at a cost competitive with available genetic tests for familial cardiomyopathy will no doubt further expand the list of putative genetic heart failure modifiers. Heart failure risk alleles along with traditional clinical factors will need to be considered by clinical cardiologists in their design of optimal disease surveillance and prevention programs and in individually tailoring heart failure management.
The use of individual genetic make-up is likely to have the earliest and greatest impact on managing patients with heart failure by tailoring available pharmacotherapeutics to optimize patient response and minimize adverse effects (ie, the area of pharmacogenetics). Modern heart failure management has been derived and directed by the results of large, randomized, multicenter clinical trials. When standard therapies are applied according to the selection criteria used in these trials, they prolong average survival across affected populations or decrease the incidence of heart failure in populations at risk.6 For this reason, standardized treatment guidelines prescribe heart failure therapies according to trial designs, aiming for the same target doses and general treatment approaches,7 and largely ignore individual characteristics. In this article, we review established and emerging knowledge of genetic influence on common heart failure and try to anticipate how these genetic factors may be best used to eschew the cookie-cutter approach to heart failure management and move toward implementing a personalized medicine approach for the treatment and prevention of this important and prevalent disease.
The Concept of Genotype-Directed Personal Medical Management in Heart Failure
Variation in clinical heart failure progression and therapeutic response (either benefits or side effects) supports the need for a more individualized approach to disease management. On the basis of clinical stratification (eg, by etiology of heart failure as ischemic versus nonischemic, functional status, comorbid disease), physicians try to match each patient’s specific heart failure syndrome with a therapeutic regime devised to provide the most benefit. Standard heart failure pharmacotherapy currently comprises a minimum of 3 medications (angiotensin-converting enzyme [ACE] inhibitors, β-blockers, and aldosterone antagonists), with consideration of additional medications (hydralazine/isosorbide, angiotensin receptor blockers) and diuretics. The recommended target dosages for these agents, derived from their respective clinical trials, is rarely achieved,8 partly because of untoward clinical side effects such as low blood pressure or renal dysfunction. Accordingly, the published guidelines most often are applied in each individual patient using ad hoc approaches derived from personal experience and the “art of medicine.”
Technological advances in human genomics promise a different approach and are bringing cardiology into an era of clinically applied pharmacogenetics9 (whether we want to or not). As sequencing costs decline, it is not hard to envision that patients will present having had their entire genome already sequenced. The imperative to apply genome information in clinical settings will increase, as demonstrated by recent proof-of-concept studies.10 Our field seems poorly prepared for this type of evolution in care; Roden et al9 identified 3 major barriers: First is the absence of rapidly available genotype information in the clinical workflow. This barrier is being overcome with whole-genome sequencing, which (with proper analysis) promises a permanent and largely immutable genetic roadmap for individual disease risk and drug response at a cost comparable to many other clinical tests.11 Second, we must have the knowledge to properly apply information on genetic variants for the diseases we are managing and the drugs we are using. As we describe, this knowledge is accumulating for heart failure and for other cardiac conditions, and the rate at which we are gaining additional information and developing further expertise appears to be accelerating.
The third and perhaps most formidable barrier is the lack of clinical evidence showing how real-time application of genetic information can best benefit patients. As has been broadly communicated to the medical community and lay public, common functional gene variants in CYP2C19 can impair the transformation of clopidogrel into its active metabolite, leading to increased risk of stent thrombosis after percutaneous coronary intervention.12 The relevant question thus becomes the following: If physicians have this information at the time of clinical care and reacted by adjusting clopidogrel dose or substituting prasugrel, which is unaffected by CYP2C19 genotype,13 would there be any improvement in clinical outcome? It is also important to consider whether any observed benefits justify the additional costs of genetic testing and for the alternate drug. Studies are currently examining these questions, and similar clinical trials will prospectively examine whether a genotype-guided strategy of warfarin dosing will be superior to the standard genotype-blinded approach in reaching target anticoagulation goals. At this time, there are no similar prospective, randomized, blinded trials of genotype-guided care for common heart failure.
Emerging Variants
The variants described here are established, but new ones are emerging. Although findings in heart failure genome-wide association studies have been limited, we can expect additional common heart failure variants to emerge as sample sizes increase.65 The CHARGE (Cohorts for Heart and Aging Research in Genomic Epidemiology) consortium published a genome-wide association study of incident heart failure that tested for associations between >2.4 million HapMap-imputed polymorphisms in >20 000 subjects.7 They identified 2 loci associated with heart failure, rs10519210 (15q22, containing USP3 encoding a ubiquitin-specific protease) in subjects of European ancestry and rs11172782 (12q14, containing LRIG3 encoding a leucine-rich, immunoglobulin-like domain-containing protein of uncertain function) in subjects of African ancestry.66 In a companion study using the same population and genotyping results, mortality analysis of the subgroup of individuals who developed heart failure implicated an intronic SNP in CMTM7 (CKLF-like MARVEL transmembrane domain-containing 7).67 These genetic associations require independent replication and further study to identify the underlying biological mechanisms.
A recently published genome-wide association study by a European consortium on dilated cardiomyopathy identified common variants in BAG3 (BCL2-associated athanogene 3) associated with heart failure57 and identified rare BAG3 missense and truncation mutations that segregate with familial cardiomyopathy. These findings were consistent with an earlier exome-sequencing study that identified BAG3 as a familial dilated cardiomyopathy gene and showed recapitulation of cardiomyopathy with BAG3 morpholino knockdown in zebra fish.68 Together, these studies convincingly support variation in BAG3 as a genetic risk factor of cardiomyopathy and heart failure. It is noteworthy that both common and rare functional variations were identified at this locus. A unifying hypothesis for these findings, which needs to be formally tested, is that common variants in BAG3 serve as proxies for rare functional BAG3 mutations with large effects. In this situation, the underlying genetic lesion is a rare variant with a large functional effect. This has recently been described for common variants in MYH6 that correlated with rare functional MYH6 variants to cause sick sinus syndrome.69 It is premature to speculate on the clinical applications of these newer findings.
Moving Knowledge to Practice
A small number of genomic variants have been identified that modify heart failure by affecting well-understood physiological systems. The principal barrier preventing their adoption in practice may be lack of evidence showing how application of this information can best be used for clinical benefit. Trials testing genotype targeting of antiplatelet therapy and anticoagulation will be completed in the coming years. The findings from these studies will likely determine the level of enthusiasm for conducting genotype-guided trials of β-blockers and RAAS antagonists in heart failure. Given that the lifetime risk of heart failure in the United States is estimated at 1 in 5, even a small favorable effect on heart failure prevention or outcome through use of genome-guided therapy has the potential for a large public health impact. We therefore believe that a near-term goal should be to conduct pharmacogenomic trials in heart failure based on our current understanding of heart failure variants.
Looking ahead, unbiased approaches will continue to reveal a large number heart failure-modifying variants (both common and rare). Based on experience in other complex phenotypes, such has height70 and plasma lipid levels,71 the underlying genetic mechanisms for many new heart failure variants will be completely unknown, and their sheer number will preclude detailed experimentation using murine models to figure them out. Leveraging these variants for clinical application is a challenge that we will be forced to confront.
As our ability to identify rare, disease-causing variants improves through personal genome sequencing, we will be faced with the additional problem of how best to estimate the disease risk conferred by a sequence variant for which there has been no biological validation. In probabilistic terms, because there are 3 billion nucleotides in the human genome and over twice that many humans on the planet, it is likely that a nucleotide substitution for every position is represented in someone. Obviously, it will be impossible to recombinantly express and functionally characterize every DNA variant that is going to be implicated in heart failure. Bioinformatics filters have been used to try and separate functionally significant from insignificant variants based on the likelihood of changing transcript expression or protein function. These tools are limited but will improve if we tailor their results to the known characteristics of each gene product. For example, current approaches to categorize amino acid substitutions as conservative or nonconservative based only on charge or side chains can be improved by molecular modeling that incorporates protein-specific structure-function information. This approach has been used to estimate the pathogenicity of myosin heavy chain (MHC) mutations in an effort to determine which mutations are likely to cause familial cardiomyopathy when linkage analysis is not feasible.72 In concept, this approach can be applied to any protein for which structure-function activities have been finely mapped to distinct domains.
A promising extension of this approach may be to use evolutionary genetics to infer disease causality. Again, using the MHC genes as examples, human genome data show a greater prevalence of nonsynonymous gene variants in MYH6, which encodes the minor cardiac α-MHC isoform, compared with the adjacent MYH7, which encodes the major β-MHC isoform. This disparity suggests a greater tolerance for protein changes in the α-MHC isoform and negative selection against these in β-MHC. We can infer, therefore, that amino acid changes are more likely to have adverse impacts in MYH7-encoded β-MHC. If this paradigm survives prospective testing, then the forthcoming explosion of individual genetic data not only will present a massive problem in interpretation, but also will provide the genetic information by which analyses of rare sequence variants across large unaffected populations can help to differentiate the tolerable variants from those that are more likely to alter disease risk.
Each Reference above is found in:
http://circgenetics.ahajournals.org/content/4/6/701.full
SOURCE:
Circulation: Cardiovascular Genetics.2011; 4: 701-709
doi: 10.1161/ CIRCGENETICS.110.959379
Pharmacogenomics
Hypertension Susceptibility Loci and Blood Pressure Response to Antihypertensives
Results From the Pharmacogenomic Evaluation of Antihypertensive Responses Study
Yan Gong, PhD, Caitrin W. McDonough, PhD, Zhiying Wang, MS, Wei Hou, PhD, Rhonda M. Cooper-DeHoff, PharmD, MS, Taimour Y. Langaee, PhD, Amber L. Beitelshees, PharmD, MPH, Arlene B. Chapman, MD, John G. Gums, PharmD, Kent R. Bailey, PhD, Eric Boerwinkle, PhD, Stephen T. Turner, MD and Julie A. Johnson, PharmD
Author Affiliations
From the Department of Pharmacotherapy and Translational Research (Y.G., C.W.M., R.M.C.-D., T.Y.L., J.G.G., J.A.J.), Department of Biostatistics, College of Medicine (W.H.), Division of Cardiovascular Medicine, College of Medicine (R.M.C.-D., J.A.J.), and Department of Community Health and Family Medicine (J.G.G.), University of Florida, Gainesville, FL; Division of Epidemiology, University of Texas at Houston, Houston, TX (Z.W., E.B.); Division of Endocrinology, Diabetes and Nutrition, University of Maryland, Baltimore, MD (A.L.B.); Renal Division, Emory University, Atlanta, GA (A.B.C.); and Division of Nephrology and Hypertension, Mayo Clinic, Rochester, MN (S.T.T.).
Correspondence to Yan Gong, PhD, Department of Pharmacotherapy and Translational Research, University of Florida, PO Box 100486, 1600 SW Archer Rd, Gainesville, FL 32610. E-mail gong@cop.ufl.edu.
Abstract
Background—To date, 39 single nucleotide polymorphisms (SNPs) have been associated with blood pressure (BP) or hypertension in genome-wide association studies in whites. Our hypothesis is that the loci/SNPs associated with BP/hypertension are also associated with BP response to antihypertensive drugs.
Methods and Results—We assessed the association of these loci with BP response to atenolol or hydrochlorothiazide monotherapy in 768 hypertensive participants in the Pharmacogenomics Responses of Antihypertensive Responses study. Linear regression analysis was performed on whites for each SNP in an additive model adjusting for baseline BP, age, sex, and principal components for ancestry. Genetic scores were constructed to include SNPs with nominal associations, and empirical P values were determined by permutation test. Genotypes of 37 loci were obtained from Illumina 50K cardiovascular or Omni1M genome-wide association study chips. In whites, no SNPs reached Bonferroni-corrected α of 0.0014, 6 reached nominal significance (P<0.05), and 3 were associated with atenolol BP response at P<0.01. The genetic score of the atenolol BP-lowering alleles was associated with response to atenolol (P=3.3×10–6 for systolic BP; P=1.6×10–6 for diastolic BP). The genetic score of the hydrochlorothiazide BP-lowering alleles was associated with response to hydrochlorothiazide (P=0.0006 for systolic BP; P=0.0003 for diastolic BP). Both risk score P values were <0.01 based on the empirical distribution from the permutation test.
Conclusions—These findings suggest that selected signals from hypertension genome-wide association studies may predict BP response to atenolol and hydrochlorothiazide when assessed through risk scoring.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 686-691
Published online before print October 19, 2012,
doi: 10.1161/ CIRCGENETICS.112.964080
Genetic Determinants of Statin-Induced Low-Density Lipoprotein Cholesterol Reduction
The Justification for the Use of Statins in Prevention: An Intervention Trial Evaluating Rosuvastatin (JUPITER) Trial
Daniel I. Chasman, PhD, Franco Giulianini, PhD, Jean MacFadyen, BA, Bryan J. Barratt, PhD, Fredrik Nyberg, MD, PhD, MPH and Paul M Ridker, MD, MPH
Author Affiliations
From the Center for Cardiovascular Disease Prevention (D.I.C., F.G., J.M., P.M.R.), JUPITER Trial Coordinating Center (D.I.C., F.G., J.M., P.M.R.), Brigham and Women’s Hospital and Harvard Medical School (D.I.C., P.M.R.), Boston, MA; Personalised Healthcare and Biomarkers, AstraZeneca Research and Development, Alderley Park, United Kingdom (B.J.B.); AstraZeneca Research and Development, Mölndal, Sweden (F.N.); and Unit of Occupational and Environmental Medicine, Department of Public Health and Community Medicine, Sahlgrenska Academy, University of Gothenburg, Gothenburg, Sweden (F.N.).
Correspondence to Daniel I. Chasman, PhD, Center for Cardiovascular Disease Prevention, Brigham and Women’s Hospital, 900 Commonwealth Ave E, Boston, MA 02215. E-mail dchasman@rics.bwh.harvard.edu
Abstract
Background—In statin trials, each 20 mg/dL reduction in cholesterol results in a 10–15% reduction of annual incidence rates for vascular events. However, interindividual variation in low-density lipoprotein cholesterol (LDL-C) response to statins is wide and may partially be determined on a genetic basis.
Methods and Results—A genome-wide association study of LDL-C response was performed among a total of 6989 men and women of European ancestry who were randomly allocated to either rosuvastatin 20 mg daily or placebo. Single nucleotide polymorphisms (SNPs) for genome-wide association (P<5×10−8) with LDL-C reduction on rosuvastatin were identified at ABCG2, LPA, and APOE, and a further association at PCSK9 was genome-wide significant for baseline LDL-C and locus-wide significant for LDL-C reduction. Median LDL-C reductions on rosuvastatin were 40, 48, 51, 55, 60, and 64 mg/dL, respectively, among those inheriting increasing numbers of LDL-lowering alleles for SNPs at these 4 loci (P trend=6.2×10−20), such that each allele approximately doubled the odds of percent LDL-C reduction greater than the trial median (odds ratio, 1.9; 95% confidence interval, 1.8–2.1; P=5.0×10−41). An intriguing additional association with sub–genome-wide significance (P<1×10-6) was identified for statin related LDL-C reduction at IDOL, which mediates posttranscriptional regulation of the LDL receptor in response to intracellular cholesterol levels. In candidate analysis, SNPs in SLCO1B1 and LDLR were confirmed as associated with LDL-C lowering, and a significant interaction was observed between SNPs in PCSK9 and LDLR.
Conclusions—Inherited polymorphisms that predominantly relate to statin pharmacokinetics and endocytosis of LDL particles by the LDL receptor are common in the general population and influence individual patient response to statin therapy.
SOURCE:
Circulation: Cardiovascular Genetics.2012; 5: 257-264
Published online before print February 13, 2012,
doi: 10.1161/ CIRCGENETICS.111.961144
Genetic Variation in the β2 Subunit of the Voltage-Gated Calcium Channel and Pharmacogenetic Association With Adverse Cardiovascular Outcomes in the INternational VErapamil SR-Trandolapril STudy GENEtic Substudy (INVEST-GENES)
Yuxin Niu, PhD*, Yan Gong, PhD*, Taimour Y. Langaee, PhD, Heather M. Davis, PharmD, Hazem Elewa, PhD, Amber L. Beitelshees, PharmD, MPH, James I. Moss, PhD, Rhonda M. Cooper-DeHoff, PharmD, Carl J. Pepine, MD and Julie A. Johnson, PharmD
Author Affiliations
From the Department of Pharmacotherapy and Translational Research and Center for Pharmacogenomics (Y.N., Y.G., T.Y.L., H.M.D., H.E., J.I.M., R.M.C.-D., J.A.J.), College of Pharmacy, University of Florida, Gainesville, Fla; Division of Endocrinology, Diabetes and Nutrition (A.L.B.), University of Maryland School of Medicine, Baltimore, Md; and Division of Cardiovascular Medicine (R.M.C.-D., C.J.P., J.A.J.), University of Florida College of Medicine, Gainesville, Fla.
Correspondence to Julie A. Johnson, PharmD, Department of Pharmacotherapy and Translational Research, College of Pharmacy, University of Florida, PO Box 100486, Gainesville, FL 32610. E-mail Johnson@cop.ufl.edu
↵* Drs Niu and Gong contributed equally to this work.
Abstract
Background— Single-nucleotide polymorphisms (SNPs) within the regulatory β2 subunit of the voltage-gated calcium channel (CACNB2) may contribute to variable treatment response to antihypertensive drugs and adverse cardiovascular outcomes.
Methods and Results— SNPs in CACNB2 from 60 ethnically diverse individuals were identified and characterized. Three common SNPs (rs2357928, rs7069292, and rs61839258) and a genome-wide association study-identified intronic SNP (rs11014166) were genotyped for a clinical association study in 5598 hypertensive patients with coronary artery disease randomized to a β-blocker (BB) or a calcium channel blocker (CCB) treatment strategy in the INternational VErapamil SR-Trandolapril STudy GENEtic Substudy (INVEST-GENES). Reporter gene assays were conducted on the promoter SNP, showing association with clinical outcomes. Twenty-one novel SNPs were identified. A promoter A>G SNP (rs2357928) was found to have significant interaction with treatment strategy for adverse cardiovascular outcomes (P for interaction, 0.002). In whites, rs2357928 GG patients randomized to CCB were more likely to experience an adverse outcome than those randomized to BB treatment strategy, with adjusted hazard ratio (HR) (CCB versus BB) of 2.35 (95% CI, 1.19 to 4.66; P=0.014). There was no evidence for such treatment difference in AG (HR, 1.16; 95% CI, 0.75 to 1.79; P=0.69) and AA (HR, 0.63; 95% CI, 0.36 to 1.11; P=0.11) patients. This finding was consistent in Hispanics and blacks. CACNB2 rs11014166 showed similar pharmacogenetic effect in Hispanics, but not in whites or blacks. Reporter assay analysis of rs2357928 showed a significant increase in promoter activity for the G allele compared to the A allele.
Conclusions— These data suggest that genetic variation within CACNB2 may influence treatment-related outcomes in high-risk patients with hypertension.
Clinical Trial Registration— URL: http://www.clinicaltrials.gov. Unique identifier: NCT00133692.
SOURCE:
Circulation: Cardiovascular Genetics.2010; 3: 548-555
doi: 10.1161/ CIRCGENETICS.110.957654
Hepatic Metabolism and Transporter Gene Variants Enhance Response to Rosuvastatin in Patients With Acute Myocardial Infarction
The GEOSTAT-1 Study
Kristian M. Bailey, MBChB, Simon P.R. Romaine, BSc, Beryl M. Jackson, RGN, Amanda J. Farrin, MSc, Maria Efthymiou, MSc, Julian H. Barth, MD, Joanne Copeland, BSc, Terry McCormack, MBBS, Andrew Whitehead, MSc, Marcus D. Flather, MBBS, Nilesh J. Samani, MD, FMedSci, Jane Nixon, PhD, Alistair S. Hall, MD, PhD, Anthony J. Balmforth, PhD and on behalf of the SPACE ROCKET Trial Group
Author Affiliations
From the Division of Cardiovascular and Diabetes Research (K.M.B., S.P.R.R., B.M.J., A.J.B.), and Division of Cardiovascular and Neuronal Remodelling (A.S.H.), Multidisciplinary Cardiovascular Research Centre, Leeds Institute of Genetics, Health and Therapeutics, University of Leeds, Leeds, United Kingdom; Clinical Trials Research Unit (A.J.F., M.E., J.C., J.N.), University of Leeds, Leeds, United Kingdom; Clinical Biochemistry (J.H.B.), Leeds General Infirmary, Leeds, United Kingdom; Whitby Group Practice (T.M.), Spring Vale Medical Centre, Whitby, North Yorkshire, United Kingdom; Pharmacy Department (A.W.), Leeds General Infirmary, Leeds, United Kingdom; Clinical Trials and Evaluation Unit (M.D.F.), Royal Brompton and Harefield NHS Trust and Imperial College, London, United Kingdom; and Department of Cardiovascular Sciences (N.J.S.), University of Leicester, Leicester, United Kingdom.
Correspondence to Alistair S. Hall, Clinical Cardiology, Multidisciplinary Cardiovascular Research Centre (MCRC), G Floor, Jubilee Building, Leeds General Infirmary, Leeds, LS1 3EX, United Kingdom. E-mail A.S.Hall@leeds.ac.uk
* Dr Bailey, Mr Romaine, Dr Hall, and Dr Balmforth contributed equally to this study.
Abstract
Background— Pharmacogenetics aims to maximize benefits and minimize risks of drug treatment. Our objectives were to examine the influence of common variants of hepatic metabolism and transporter genes on the lipid-lowering response to statin therapy.
Methods and Results— The Genetic Effects On STATins (GEOSTAT-1) Study was a genetic substudy of Secondary Prevention of Acute Coronary Events—Reduction of Cholesterol to Key European Targets (SPACE ROCKET) (a randomized, controlled trial comparing 40 mg of simvastatin and 10 mg of rosuvastatin) that recruited 601 patients after myocardial infarction. We genotyped the following functional single nucleotide polymorphisms in the genes coding for the cytochrome P450 (CYP) metabolic enzymes, CYP2C9*2 (430C>T), CYP2C9*3 (1075A>C), CYP2C19*2 (681G>A), CYP3A5*1 (6986A>G), and hepatic influx and efflux transporters SLCO1B1 (521T>C) and breast cancer resistance protein (BCRP; 421C>A). We assessed 3-month LDL cholesterol levels and the proportion of patients reaching the current LDL cholesterol target of <70 mg/dL (<1.81 mmol/L). An enhanced response to rosuvastatin was seen for patients with variant genotypes of either CYP3A5 (P=0.006) or BCRP (P=0.010). Furthermore, multivariate logistic-regression analysis revealed that patients with at least 1 variant CYP3A5 and/or BCRP allele (n=186) were more likely to achieve the LDL cholesterol target (odds ratio: 2.289; 95% CI: 1.157, 4.527; P=0.017; rosuvastatin 54.0% to target vs simvastatin 33.7%). There were no differences for patients with variants of CYP2C9, CYP2C19, or SLCO1B1 in comparison with their respective wild types, nor were differential effects on statin response seen for patients with the most common genotypes for CYP3A5 and BCRP (n=415; odds ratio: 1.207; 95% CI: 0.768, 1.899; P=0.415).
Conclusion— The LDL cholesterol target was achieved more frequently for the 1 in 3 patients with CYP3A5 and/or BCRP variant genotypes when prescribed rosuvastatin 10 mg, compared with simvastatin 40 mg.
Clinical Trial Registration— URL: http://isrctn.org. Unique identifier: ISRCTN 89508434.
SOURCE:
Circulation: Cardiovascular Genetics.2010; 3: 276-285
Published online before print March 5, 2010,
doi: 10.1161/ CIRCGENETICS.109.898502
Comprehensive Whole-Genome and Candidate Gene Analysis for Response to Statin Therapy in the Treating to New Targets (TNT) Cohort
John F. Thompson, PhD, Craig L. Hyde, PhD, Linda S. Wood, MS, Sara A. Paciga, MA, David A. Hinds, PhD, David R. Cox, MD, PhD, G. Kees Hovingh, MD, PhD and John J.P. Kastelein, MD, PhD
Author Affiliations
From the Helicos BioSciences (J.F.T.), Cambridge, Mass; Molecular Medicine (J.F.T., L.S.W., S.A.P.) and Statistical Applications (C.L.H.), Pfizer Global Research and Development, Groton, Conn; Perlegen Sciences (D.A.H., D.R.C.), Mountain View, Calif; and Department of Vascular Medicine (G.K.H., J.J.P.K.), Academic Medical Center, Amsterdam, The Netherlands.
Correspondence to John J.P. Kastelein, MD, PhD, Department of Vascular Medicine, Academic Medical Center, Meibergdreef 9, Room F4-159.2, 1105 AZ Amsterdam, The Netherlands. E-mail j.j.kastelein@amc.uva.nl or j.s.jansen@amc.uva.nl
Abstract
Background— Statins are effective at lowering low-density lipoprotein cholesterol and reducing risk of cardiovascular disease, but variability in response is not well understood. To address this, 5745 individuals from the Treating to New Targets (TNT) trial were genotyped in a combination of a whole-genome and candidate gene approach to identify associations with response to atorvastatin treatment.
Methods and Results— A total of 291 988 single-nucleotide polymorphisms (SNPs) from 1984 individuals were analyzed for association with statin response, followed by genotyping top hits in 3761 additional individuals. None was significant at the whole-genome level in either the initial or follow-up test sets for association with low-density lipoprotein cholesterol, high-density lipoprotein cholesterol, or triglyceride response. In addition to the whole-genome platform, 23 candidate genes previously associated with statin response were analyzed in these 5745 individuals. Three SNPs in apoE were most highly associated with low-density lipoprotein cholesterol response, followed by 1 in PCSK9 with a similar effect size. At the candidate gene level, SNPs in HMGCR were also significant though the effect was less than with those in apoE and PCSK9. rs7412/apoE had the most significant association (P=6×10−30), and its high significance in the whole-genome study (P=4×10−9) confirmed the suitability of this population for detecting effects. Age and gender were found to influence low-density lipoprotein cholesterol response to a similar extent as the most pronounced genetic effects.
Conclusions— Among SNPs tested with an allele frequency of at least 5%, only SNPs in apoE are found to influence statin response significantly. Less frequent variants in PCSK9 and smaller effect sizes in SNPs in HMGCR were also revealed.
SOURCE:
Circulation: Cardiovascular Genetics.2009; 2: 173-181
Published online before print February 12, 2009,
doi: 10.1161/ CIRCGENETICS.108.818062
Summary
Larry H. Bernstein, MD, FCAP
This review has examined a compendium of well regarded documents drawn from 248 articles in Circulation Cardiovascular Genetics from March 2010 to March 2013. The large amount of evidence obtained from large population studies identifying Genome Wide Analysis Studies (GWAS) examines a host of cardiac and vascular diseases in which there is association between specific single nucleotide peptides (SNPs), and gene loci, that may play or have no significant role in developing heart disease. It certainly is evidence of the role that the American Heart Association has is in supporting the leading research today for tomorrow’s patients. It is too early to sort them out, but it speaks to a large volume of discovery in this area.
It raises another issue that we have been confronted with mostly since the second half of the 20th century. What is that issue? The issue, it appears to me, is the vast improvements in analytical technology so that “imprecision” is far less likely to be a confounder in biological measurements and this lends access to far better accuracy? But from that question arises another! Accuracy only refers to what is measured, but does it give us better ability to explain a complex and dynamic process? In other words, what is what we are looking at representative of in manageable events? I think that this is the most important idea that should come out of the recent criticism of the trajectory that molecular genetics been on in the last 5 years.
It was still in an era that “BIG’ science was not the normal. One could spend an enormous effort at stepwise purification of a protein or enzyme, or other biomolecule starting with a slurry made from 100 lbs of “chicken heart”, for example. These separations were based on negative charges on the molecules and positive charges on the column, and the molecules of no interest were eluted by gradient elution. Much was learned about large scale preparation from small scale trials. But this work was not undertaken without the intent to carry out a number of investigations to understand the “functionality” of a link in a metabolic pathway. The studies that followed the purification required kinetic investigation with a coenzyme, or with a synthetically modified coenzyme, amino acid sequencing, NMR studies, etc. You could not put together a “mechanism” without having the minimum amount of necessary information for a reliable account. It is probably this requirement that led to today’s “BIG” science, that is founded upon multiple methods, now large data bases, and teams of investigators across institutions and continents. The acquisition of knowledge has been astounding, but the integration of knowledge has not caught up.
However, let’s see if we can sort out the most meaningful signals from what I too am beginning to call the “noisy channel”. As often happens, important areas of research are opened up that are followed by significant discovery and, in the long run, many other dead end publications that have no lasting significance. In order to do justice to the work, I’ll pick through documents I find interesting, keeping in mind there is a hidden layer of complexity of which only sufficient information leads to a better understanding. As much literature calls attention to, much of what ails us has nothing to do with classical Mendelian genetics, and has a postgenomic component.
The most fascinating aspect of this is the withering “dark matter” of the genome. While that component may be silent or expressed, the understanding comes at a higher observed order. The dark became light! The expression became subtle, like weak bond interactions. The underlying organization is a component of the adaptive ability of an organism or individual in an environment with plants and animals in a changing climate, at particular altitudes, with given water supplies, with disease vectors, and with endogenous sources of essential nutrients. This brings into focus the regulatory role of the genome as just as important a factor as transmission of the genetic code, especially in somatic cell populations.
The remainder of this discussion deals specifically with my observations on cardiovascular genomics. The following conclusion is appropriate, if incomplete, at this time on circulating miRNAs, particularly miR-133a:
- elevated levels of circulating miR-133a in patients with cardiovascular diseases originate mainly from the injured myocardium.
- Circulating miR-133a can be used as a marker for cardiomyocyte death, and
A number of articles that cite this article suggest that it may be useful for following disease progression:
Plasma microRNAs serve as biomarkers of therapeutic efficacy and disease progression in hypertension-induced heart failure Eur J Heart Fail 2013
MicroRNAs Within the Continuum of Postgenomics Biomarker Discovery Arterio. Thromb. Vasc. Bio. 2013;33:206-214
“Need for Rigor in Design, Reporting, and Interpretation of Transcriptomic Biomarker Studies” J. Clin. Microbiol.. 2012;50:4192-4193
Circulating microRNAs as diagnostic biomarkers for cardiovascular diseases. Am. J. Physiol. Heart Circ. Physiol.. 2012;303:H1085-H1095,
Circulating MicroRNAs: Novel Biomarkers and Extracellular Communicators in Cardiovascular Disease? Circ. Res.. 2012;110:483-495
Circulating MicroRNAs: Biomarkers or Mediators of Cardiovascular Diseases? Arterioscler. Thromb. Vasc. Bio. 2011;31:2383-2390,
Circulating MicroRNA-208b and MicroRNA-499 Reflect Myocardial Damage in Cardiovascular Disease MF Corsten, R Dennert, S Jochem, T Kuznetsova, et al.
The finding refers to an association that is related to the appearance of a miRNA in the circulation of patients with acute cardiac ischemia, and particular released into the circulation of patients from injured myocardium. This finding has to be distinguished from a finding of another miRNA released with acute injury. In the case of miR499 (and miR208b), there is a comparison with plasma cTnT, and an ROC curve is produced.
The List of this follows:
Circulation: Cardiovascular Genetics 2010; 3: 499-506
Strikingly, in plasma from
- acute myocardial infarction patients, cardiac myocyte–associated miR-208b and -499 were highly elevated, 1600-fold (P<0.005) and 100-fold (P<0.0005), respectively, as compared with control subjects. Receiver operating characteristic curve analysis revealed an area under the curve of 0.94 (P<10−10) for miR-208b and 0.92 (P<10−9) for miR-499. Both microRNAs correlated with plasma troponin T, indicating release of microRNAs from injured cardiomyocytes.
- In patients with acute heart failure, only miR-499 was significantly elevated (2-fold), whereas
- no significant changes in microRNAs studied could be observed in diastolic dysfunction.
Remarkably, plasma microRNA levels were not affected by a wide range of clinical confounders, including
- age,
- sex,
- body mass index,
- kidney function,
- systolic blood pressure, and
- white blood cell count.
This is miRNA with a different twist. It appears that there are 3 types found in AMI (133a, 208b, 409). But type 499 alone is increased with acute heart failure (no mention of chronic cardiomyopathy and no effect of estimated GFR, or of age).
If the problem was just of AMI, then we have to know what this brings to the table. As it is the hs-troponins have yet to be shown to effectively not only increase the high sensitivity of the tests, but to decrease the confusion generated by the elevation. The enormous improvement of a test that may be superior to the hs-ctn’s is for the patient with very indeterminiate shortness of breath, a nondefinitive ECG, and in a prodromal phase of AMI. This happened in the past, and it may happen now, and it may account for many cases of silent MI that were found at autopsy.
Cited by
Plasma microRNAs serve as biomarkers of therapeutic efficacy and disease progression in hypertension-induced heart failure Eur J Heart Fail. 2013;0:hft018v1-hft018,
Circulating microRNAs as diagnostic biomarkers for cardiovascular diseases Am. J. Physiol. Heart Circ. Physiol.. 2012;303:H1085-H1095,
Circulation Editors’ Picks: Most Read Articles in Cardiovascular Genetics Circulation. 2012;126:e163-e169,
MicroRNAs in Patients on Chronic Hemodialysis (MINOS Study) CJASN. 2012;7:619-623,
Novel techniques and targets in cardiovascular microRNA research Cardiovasc Res. 2012;93:545-554,
Microparticles: major transport vehicles for distinct microRNAs in circulation Cardiovasc Res. 2012;93:633-644,
Profiling of circulating microRNAs: from single biomarkers to re-wired networks Cardiovasc Res. 2012;93:555-562,
Small but smart–microRNAs in the centre of inflammatory processes during cardiovascular diseases, the metabolic syndrome, and ageing Cardiovasc Res. 2012;93:605-613,
Circulation: Heart Failure Editors’ Picks: Most Important Papers in Pathophysiology and Genetics Circ Heart Fail. 2012;5:e32-e49
Use of Circulating MicroRNAs to Diagnose Acute Myocardial Infarction Clin. Chem. 2012;58:559-567,
Circulating microRNAs to identify human heart failure Eur J Heart Fail. 2012;14:118-119,
Next Steps in Cardiovascular Disease Genomic Research–Sequencing, Epigenetics, and Transcriptomics Clin. Chem. 2012;58:113-126,
Most Read in Cardiovascular Genetics on Biomarkers, Inherited Cardiomyopathies and Arrhythmias, Metabolomics, and Genomics Circ Cardiovasc Genet. 2011;4:e24-e30,
MicroRNA-126 modulates endothelial SDF-1 expression and mobilization of Sca-1+/Lin- progenitor cells in ischaemia Cardiovasc Res. 2011;92:449-455,
The use of genomics for treatment is another matter, and has several factors, e.g., age, residual function after AMI, comorbidities
This is a lot of interesting work that opens as many questions as it answers. The observations are real, and they lead to questions relating to the heart and the circulation. Maybe it will generate answers to very tough issues concerning hypertension, renal disease and the heart. It is far too early to tell. It appears that we are about to hear a cacophony of miR’s in a symphony on cardiac and circulatory diseases not be be pieced together soon. But we have many more tools at our disposal than we did when Karmen discovered and made a distinction between
- Aspartate and Alanine aminotransferases in the late 1950s, followed in the 1960s by
- Creatine phosphokinase, the
- MB-isoenzyme of CK by Sobel, Shell and Kjeckshus,
- isoenzyme-1 of lactate dehydrogenase, and later the
- Troponins,
leading to the programs to “reduce the extent of infarct damage”. Then came the
- a- and b-type natriuretic peptides,
which are still not fully understood in their role in congestive heart failure and in renal disease.
One item strikes the imagination as a fruitful area of further study. Genetic Determinants of Potassium Sensitivity and Hypertension. Integrated Computational and Experimental Analysis of the Neuroendocrine Transcriptome in Genetic Hypertension Identifies Novel Control Points for the Cardiometabolic Syndrome
Essential hypertension, a common complex disease, displays substantial genetic influence. Contemporary methods to dissect the genetic basis of complex diseases such as the genomewide association study are powerful, yet a large gap exists betweens the fraction of population trait variance explained by such associations and total disease heritability.
The researchers
- developed a novel, integrative method (combining animal models, transcriptomics, bioinformatics, molecular biology, and trait-extreme phenotypes)
- to identify candidate genes for essential hypertension and the metabolic syndrome.
Method … transcriptome profiling on adrenal glands from blood pressure extreme mouse strains:
- the hypertensive BPH (blood pressure high) and
- hypotensive BPL (blood pressure low).
Results…. Microarray data clustering revealed
- underexpression of intermediary metabolism transcripts in HIGH BLOOD PRESSURE.
- The MITRA algorithm identified a conserved motif in the transcriptional regulatory regions of the underexpressed metabolic genes,
- They decide that regulation through this motif contributed to the global underexpression.
- Luciferase reporter assays demonstrated transcriptional activity of the motif through transcription factors
They finally hypothesized that genetic variation at HOXA3, SRY, and YY1 might predict blood pressure and other metabolic syndrome traits in humans. Tagging variants for each locus were associated with
- blood pressure in a human population blood pressure extreme sample with
- the most extensive associations for YY1 tagging single nucleotide polymorphism rs11625658 on
- systolic blood pressure,
- diastolic blood pressure,
- body mass index, and
- fasting glucose.
Meta-analysis extended the YY1 results into 2 additional large population samples with significant effects preserved on diastolic blood pressure, body mass index, and fasting glucose.
It will take much more of this beautiful integrative work to open up our imagination as to what physiological processes are occurring.
Like this:
Like Loading...
Read Full Post »
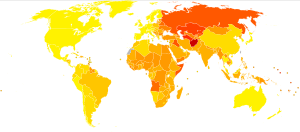

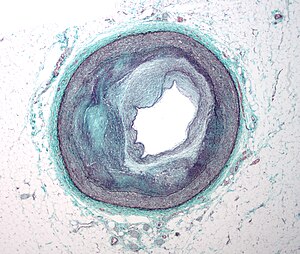
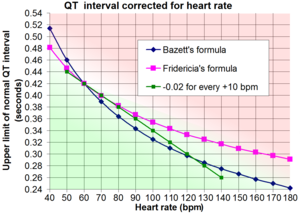









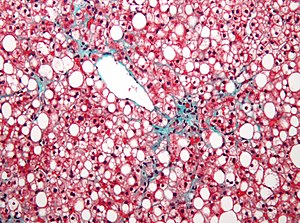

![Steps for the development of imaging biomarkers (adapted from [5])](https://i0.wp.com/pharmaceuticalintelligence.com/wp-content/uploads/2013/03/fig-1.gif?resize=500%2C363)