Curator & Author: Larry H. Bernstein, MD, FCAP
Leaders in Pharmaceutical Intelligence
Subtitle: Nitric Oxide, Peroxinitrite, and NO donors in Renal Function Loss
Summary: The criticality of renal function is traced to the emergence of animal forms from the sea to land. It also becomes acutely and/or chronically dysfunctional in metabolic, systemic inflammatory and immunological diseases of man. We have already described the key role that nitric oxide and the NO synthases play in reduction of oxidative stress, and we have seen that a balance has to be struck between pro- and anti-oxidative as well as inflammatory elements for avoidance of diseases, specifically involving the circulation, but effectively not limited to any organ system. In this discussion we shall look at kidney function, NO and NO donors. This is an extension of a series of posts on NO and NO related disorders.
__________________________________________________________________________________________________________________________________________________________
Part I. The evolution of kidney structure and Function Evolution of kidney function
In fish the nerves that activate breathing take a short journey from an ancient part of the brain, the brain stem, to the throat and gills. For the ancient tadpole, the nerve controlling a reflex related to hiccup in man served a useful purpose, allowing the entrance to the lung to remain open when breathing air but closing it off when gulping water – which would then be directed only to the gills.
For humans and other mammals it provides a bit of evidence of our common ancestry. DNA evidence has pinned iguanas and chameleons as the closest relatives to snakes. In utero, we develop three separate kidneys in succession, absorbing the first two before we wind up with the embryonic kidney that will become our adult kidney. The first two of these reprise embryonic kidneys of ancestral forms, and in the proper evolutionary order.
The pronephric kidney does not function in human and other mammalian embryos. It disappears and gives rise to the Mesonephric kidney. This kidney filters wastes from the blood and excretes them to the outside of the body via a pair of tubes called the mesonephric ducts (also “Wolffian ducts”). The mesonephric kidney goes on to develop into the adult kidney of fish and amphibians.
This kidney does function for a few weeks in the human embryo, but then disappears as our final kidney forms, which is the Metanephric kidney. This begins developing about five weeks into gestation, and consists of an organ that filters wastes from the blood and excretes them to the outside through a pair ureters. In the embryo, the wastes are excreted directly into the amniotic fluid. The metanephric kidney is the final adult kidney of reptiles, birds, and mammals.
The first two kidneys resemble, in order, those of primitive aquatic vertebrates (lampreys and hagfish) and aquatic or semiaquatic vertebrates (fish and amphibians): an evolutionary order.
The explanation, then, is that we go through developmental stages that show organs resembling those of our ancestors. Take a step back and we see that fresh water fish have glomerular filtration. Cardiac contraction provides the pressure to force the water, small molecules, and ions into the glomerulus as nephric filtrate. The essential ingredients are then reclaimed by the tubules, returning to the blood in the capillaries surrounding the tubules. The amphibian kidney also functions chiefly as a device for excreting excess water.
But the problem is to conserve water, not eliminate it. The frog adjusts to the varying water content of its surroundings by adjusting the rate of filtration at the glomerulus. When blood flow through the glomerulus is restricted, a renal portal system is present to carry away materials reabsorbed through the tubules. Bird kidneys function like those of reptiles (from which they are descended). Uric acid is also their chief nitrogenous waste. All mammals share our use of urea as their chief nitrogenous waste. Urea requires much more water to be excreted than does uric acid. Mammals produce large amounts of nephric filtrate but are able to reabsorb most of this in the tubules. But even so, humans lose several hundred ml each day in flushing urea out of the body.
In his hypothesis of the evolution of renal function Homer Smith proposed that the formation of glomerular nephron and body armor had been adequate for the appearance of primitive vertebrates in fresh water and that the adaptation of homoiotherms to terrestrial life was accompanied by the appearance of the loop of Henle.
In the current paper, the increase in the arterial blood supply and glomerular filtration rate and the sharp elevation of the proximal reabsorption are viewed as important mechanisms in the evolution of the kidney. The presence of glomeruli in myxines and of nephron loops in lampreys suggests that fresh water animals used the preformed glomerular apparatus of early vertebrates, while mechanisms of urinary concentration was associated with the subdivision of the kidney into the renal cortex and medulla. The principles of evolution of renal functions can be observed at several levels of organizations in the kidney.
Natochin YV. Evolutionary aspects of renal function. Kidney International 1996; 49: 1539–1542; doi:10.1038/ki.1996.220. Smith HW: From Fish to Philosopher. Boston, Little, Brown, 1953.
___________________________________________________________________________________________________________________________________________________________
The Kidney: Anatomy and Physiology
The kidney lies in the lower abdomen capped by the adrenal glands. It has an outer cortex and an inner medulla. The basic unit is the nephron, which filters blood at the glomerulus, and not only filters urine eliminating mainly urea, also uric acid, and other nitrogenous waste, but also reabsorbs Na+ in exchange for H+/(reciprocal K+) through the carbonic anhydrase of the epithelium. In addition, it serves as a endocrine organ and receptor through the renin-angiotensin/aldosterone system, sensitivity to water loss controlled by antidiuretic hormone, and is sensitive to the natriuretic peptides of the heart. The kidney is an elegant structure with a high concentration of glomeruli in the cortex, and in the medulla one finds a U-shaped tube that is critical in a countercurrent multiplier system with a descending limb, Loop of Henley, and ascending limb.
As the filtrate flows through the glomerulus into the descending limb, there is reabsorption of glucose and of H+ by the carbonic anhydrase conversion to water and CO2, except with serious acidemia, in which K+ is reabsorbed with H+ loss to the filtrate, resulting in a hyperkalemia. In the descending limb Na+ is absorbed into the interstitium, and the hypertonic interstitium draws water back for circulation, regulated by the action of ADH on the epithelium of the ascending limb. The result in terms of basic urinary clearance, the volume of urine loss is moderated by the amount needed for circulation (10 units of whole blood) without dehydration, and an amount sufficient for metabolite loss (including drug metabolites). The urine flows into the kidney pelvis and flow down the ureters.
The renal blood flow needs mention. The blood reaches the glomerulus by way of the afferent arteriole and leaves by way of the efferent arteriole. In a book by the Harvard Pathologist Shields Warren on diabetes he made a distinction between hypertension and diabetes in that efferent arteriolar sclerosis is present in both, but diabetes is uniquely identified by afferent arteriolar sclerosis. In diabetes you also have a typical glomerulosclerosis, which might be related to the same hyalinization found in the pancreatic islets – a secondary amyloidosis.
______________________________________________________________________________________________________________________________________________________________

English: Nephron, Diagram of the urine formation. The number inside tubular urin concentration in mOsm/l – when ADH acts Polski: Nefron, Schemat tworzenia moczu. Cyfry wewnątrz kanalików oznaczają lokalne stężenie w mOsm/l – gdy działa ADH (dochodzi do zagęszczania moczu). (Photo credit: Wikipedia)
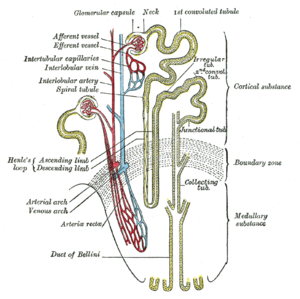
Loop of Henle (Grey’s Anatomy book) (Photo credit: Wikipedia)

Frontal section through the kidney (Photo credit: Wikipedia)
_____________________________________________________________________________________________________________________________________________________________
_ Part IIa. Nitric Oxide role in renal tubular epithelial cell function Tubulointerstitial Nephritides
As part of the exponential growth in our understanding of nitric oxide (NO) in health and disease over the past 2 decades, the kidney has become appreciated as a major site where NO may play a number of important roles. Although earlier work on the kidney focused more on effects of NO at the level of larger blood vessels and glomeruli, there has been a rapidly growing body of work showing critical roles for NO in tubulointerstitial disease. In this review we discuss some of the recent contributions to this important field.
Mattana J, Adamidis A, Singhal PC. Nitric oxide and tubulointerstitial nephritides. Seminars in Nephrology 2004; 24(4):345-353.
Nitric oxide donors and renal tubular (subepithelial) matrix
Nitric oxide (NO) and its metabolite, peroxynitrite (ONOO-), are involved in renal tubular cell injury. If NO/ONOO- has an effect to reduce cell adhesion to the basement membrane, does this effect contribute to tubular obstruction and would it be partially responsible for the harmful effect of NO on the tubular epithelium during acute renal failure (ARF)?
Wangsiripaisan A, et al. examined the effect of the NO donors
- [1] (z)-1-[2-(2-aminoethyl)-N-(2-ammonioethyl)amino]diazen-1- ium-1, 2-diolate (DETA/NO),
- [2] spermine NONOate (SpNO), and
- [3] the ONOO- donor 3-morpholinosydnonimine (SIN-1) on
cell-matrix adhesion to collagen types I and IV, and also fibronectin using three renal tubular epithelial cell lines:
- [1] LLC-PK1,
- [2] BSC-1, and
- [3] OK.
It was only the exposure to SIN-1 that caused a dose-dependent impairment in cell-matrix adhesion.
Similar results were obtained in the different cell types and matrix proteins. The effect of SIN-1 (500 microM) on LLC-PK1 cell adhesion was not associated with either cell death or alteration of matrix protein and was attenuated by either
- [1] the NO scavenger 2-(4-carboxyphenyl)-4,4,5,5-tetramethylimidazoline-1-oxyl-3-oxide,
- [2] the superoxide scavenger superoxide dismutase, or
- [3] the ONOO- scavenger uric acid in a dose-dependent manner.
These investigators concluded in this seminal paper that ONOO- generated in the tubular epithelium during ischemia/reperfusion has the potential to impair the adhesion properties of tubular cells, which then may contribute to the tubular obstruction in ARF.
Wangsiripaisan A, Gengaro PE, Nemenoff RA, Ling H, et al. Effect of nitric oxide donors on renal tubular epithelial cell-matrix adhesion. Kidney Int 1999; 55(6):2281-8.
Coexpressed Nitric Oxide Synthase and Apical β1 Integrins
In sepsis-induced acute renal failure, actin cytoskeletal alterations result in shedding of proximal tubule epithelial cells (PTEC) and tubular obstruction.
This study examined the hypothesis that inflammatory cytokines, released early in sepsis, cause PTEC cytoskeletal damage and alter integrin-dependent cell-matrix adhesion. The question of whether the intermediate nitric oxide (NO) modulates these cytokine effects was also examined. After exposure of human PTEC to tumor necrosis factor-α, interleukin-1α, and interferon-γ, the actin cytoskeleton was disrupted and cells became elongated, with extension of long filopodial processes.
Cytokines induced shedding of viable, apoptotic, and necrotic PTEC, which was dependent on NO synthesized by inducible NO synthase (iNOS) produced as a result of cytokine actions on PTEC. Basolateral exposure of polarized PTEC monolayers to cytokines induced maximal NO-dependent cell shedding, mediated in part through NO effects on cGMP. Cell shedding was accompanied by dispersal of basolateral β1 integrins and E-cadherin, with corresponding upregulation of integrin expression in clusters of cells elevated above the epithelial monolayer.
These cells demonstrated coexpression of iNOS and apically redistributed β1 integrins. These authors point out that the major ligand involved in cell anchorage was laminin, probably through interactions with the integrin α3β1.
This interaction was downregulated by cytokines but was not dependent on NO. They posulate a mechanism by which inflammatory cytokines induce PTEC damage in sepsis, in the absence of hypotension and ischemia.
Glynne PA, Picot J and Evans TJ. Coexpressed Nitric Oxide Synthase and Apical β1 Integrins Influence Tubule Cell Adhesion after Cytokine-Induced Injury. JASN 2001; 12(11): 2370-2383.
Potentiation by Nitric Oxide of Apoptosis in Renal Proximal Tubule Cells
Proximal tubular epithelial cells (PTEC) exhibit a high sensitivity to undergo apoptosis in response to proinflammatory stimuli and immunosuppressors and participate in the onset of several renal diseases. This study examined the expression of inducible nitric oxide (NO) synthase after challenge of PTEC with bacterial cell wall molecules and inflammatory cytokines and analyzed the pathways that lead to apoptosis in these cells by measuring changes in the mitochondrial transmembrane potential and caspase activation.
The data show that the apoptotic effects of proinflammatory stimuli mainly were due to the expression of inducible NO synthase. Cyclosporin A and FK506 inhibited partially NO synthesis.
However, both NO and immunosuppressors induced apoptosis, probably through a common mechanism that involved the irreversible opening of the mitochondrial permeability transition pore. Activation of caspases 3 and 7 was observed in cells treated with high doses of NO and with moderate concentrations of immunosuppressors.
The conclusion is that the cooperation between NO and immunosuppressors that induce apoptosis in PTEC might contribute to the renal toxicity observed in the course of immunosuppressive therapy.
Hortelano S, Castilla M, Torres AM, Tejedor A, and Bosca L. Potentiation by Nitric Oxide of Cyclosporin A and FK506- Induced Apoptosis in Renal Proximal Tubule Cells. J Am Soc Nephrol 2000; 11: 2315–2323.
___________________________________________________________________________________________________________________________________________________________
Part IIb. Related studies with ROS and/or RNS on nonrenal epithelial cells
Reactive nitrogen species block cell cycle re-entry Endogenous sources of reactive nitrogen species (RNS) act as second messengers in a variety of cell signaling events, whereas environmental sources of RNS like nitrogen dioxide (NO2) inhibit cell survival and growth through covalent modification of cellular macromolecules. Murine type II alveolar cells arrested in G0 by serum deprivation were exposed to either NO2 or SIN-1, a generator of RNS, during cell cycle re-entry.
In serum-stimulated cells, RNS blocked cyclin D1 gene expression, resulting in cell cycle arrest at the boundary between G0 and G1. Dichlorofluorescin diacetate (DCF) fluorescence indicated that RNS induced sustained production of intracellular hydrogen peroxide (H2O2), which normally is produced only transiently in response to serum growth factors.
Loading cells with catalase prevented enhanced DCF fluorescence and rescued cyclin D1 expression and S phase entry.
These studies indicate environmental RNS interfere with cell cycle re-entry through an H2O2-dependent mechanism that influences expression of cyclin D1 and progression from G0 to the G1 phase of the cell cycle.
Yuan Z, Schellekens H, Warner L, Janssen-Heininger Y, Burch P, Heintz NH. Reactive nitrogen species block cell cycle re-entry through sustained production of hydrogen peroxide. Am J Respir Cell Mol Biol. 2003;28(6):705-12. Epub 2003 Jan 10.
Peroxynitrite modulates MnSOD gene expression
Peroxynitrite (ONOO-) is a strong oxidant derived from nitric oxide (‘NO) and superoxide (O2.-), reactive nitrogen (RNS) and oxygen species (ROS) present in inflamed tissue. Other oxidant stresses, e.g., TNF-alpha and hyperoxia, induce mitochondrial, manganese-containing superoxide dismutase (MnSOD) gene expression. 3-morpholinosydnonimine HCI (SIN-1) (10 or 1000 microM) increased MnSOD mRNA, but did not change hypoxanthine guanine phosphoribosyl transferase (HPRT) mRNA. Authentic peroxynitrite (ONOO ) (100-500 microM) also increased MnSOD mRNA but did not change constitutive HPRT mRNA expression. ONOO stimulated luciferase gene expression driven by a 2.5 kb fragment of the rat MnSOD gene 5′ promoter region.
MnSOD gene induction due to ONOO- was
- [1] inhibited effectively by L-cysteine (10 mM) and
- [2] partially inhibited by N-acetyl cysteine (NAC)(50 mM) or
- [3] pyrrole dithiocarbamate (10 mM).
.NO from 1-propanamine, 3-(2-hydroxy-2-nitroso-1-propylhydrazine) (PAPA NONOate) (100 or 1000 microM) did not change MnSOD or HPRT mRNA, nor did either H202 or NO2-, breakdown products of SIN-1 and ONOO, have any effect on MnSOD mRNA expression; ONOO- and SIN-1 also did not increase detectable MnSOD protein content or increase MnSOD enzymatic activity.
Nevertheless, increased steady state [O2.-] in the presence of .NO yields ONOO , and ONOO has direct, stimulatory effects on MnSOD transcript expression driven at the MnSOD gene 5′ promoter region inhibited completely by L-cysteine and partly by N-acetyl cysteine in lung epithelial cells. This raises a question of whether the same effect is seen in renal tubular epithelium.
Jackson RM, Parish G, Helton ES. Peroxynitrite modulates MnSOD gene expression in lung epithelial cells. Free Radic Biol Med. 1998; 25(4-5):463-72.
Comparative impacts of glutathione peroxidase-1 gene knockout on oxidative stress
Selenium-dependent glutathione peroxidase-1 (GPX1) protects against reactive-oxygen-species (ROS)-induced oxidative stress in vivo, but its role in coping with reactive nitrogen species (RNS) is unclear. Primary hepatocytes were isolated from GPX1-knockout (KO) and wild-type (WT) mice to test protection of GPX1 against cytotoxicity of
- [1] superoxide generator diquat (DQ),
- [2]NO donor S-nitroso-N-acetyl-penicillamine (SNAP) and
- [3] peroxynitrite generator 3-morpholinosydnonimine (SIN-1).
Treating cells with SNAP in addition to DQ produced synergistic cytotoxicity that minimized differences in apoptotic cell death and oxidative injuries between the KO and WT cells. Less protein nitrotyrosine was induced by 0.05-0.5 mM DQ+0.25 mM SNAP in the KO than in the WT cells.
Total GPX activity in the WT cells was reduced by 65 and 25% by 0.5 mM DQ+0.1 mM SNAP and 0.5 mM DQ, respectively. Decreases in Cu,Zn-superoxide dismutase (SOD) activity and increases in Mn-SOD activity in response to DQ or DQ+SNAP were greater in the KO cells than in the WT cells.
The study indicates GPX1 was more effective in protecting hepatocytes against oxidative injuries mediated by ROS alone than by ROS and RNS together, and knockout of GPX1 did not enhance cell susceptibility to RNS-associated cytotoxicity. Instead, it attenuated protein nitration induced by DQ+SNAP.
To better understand the mechanism(s) underlying nitric oxide (. NO)-mediated toxicity, in the presence and absence of concomitant oxidant exposure, postmitotic terminally differentiated NT2N cells (which are incapable of producing . NO) were exposed to [1]PAPA-NONOate (PAPA/NO) and [2] 3-morpholinosydnonimine (SIN-1).
Exposure to SIN-1, which generated peroxynitrite (ONOO) in the range of 25-750 nM/min, produced a concentration- and time-dependent delayed cell death. In contrast, a critical threshold concentration (>440 nM/min) was required for . NO to produce significant cell injury. There is a largely necrotic lesion after ONOO exposure and an apoptotic-like morphology after . NO exposure.
Cellular levels of reduced thiols correlated with cell death, and pretreatment with N-acetylcysteine (NAC) fully protected from cell death in either PAPA/NO or SIN-1 exposure. NAC given within the first 3 h posttreatment further delayed cell death and increased the intracellular thiol level in SIN-1 but not . NO-exposed cells.
Cell injury from . NO was independent of cGMP, caspases, and superoxide or peroxynitrite formation. Overall, exposure of non-. NO-producing cells to . NO or peroxynitrite results in delayed cell death, which, although occurring by different mechanisms, appears to be mediated by the loss of intracellular redox balance.
Gow AJ, Chen Q, Gole M, Themistocleous M, Lee VM, Ischiropoulos H. Two distinct mechanisms of nitric oxide-mediated neuronal cell death show thiol dependency. Am J Physiol Cell Physiol. 2000; 278(6):C1099-107.
NO2 effect on phosphatidyl choline Nitrogen dioxide (NO2) inhalation affects the extracellular surfactant as well as the structure and function of type II pneumocytes.
The studies had differences in oxidant concentration, duration of exposure, and mode of NO2 application. This study evaluated the influence of the NO2 application mode on the phospholipid metabolism of type II pneumocytes. Rats were exposed to identical NO2 body doses (720 ppm x h), which were applied continuously (10 ppm for 3 d), intermittently (10 ppm for 8 h per day, for 9 d), and repeatedly (10 ppm for 3 d, 28 d rest, and then 10 ppm for 3 d). Immediately after exposure, type II cells were isolated and evaluated for cell yield, vitality, phosphatidylcholine (PC) synthesis, and secretion.
Type II pneumocyte cell yield was only increased from animals that had been continuously exposed to NO2, but vitality of the isolated type II pneumocytes was not affected by the NO2 exposure modes. Continuous application of 720 ppm x h NO2 resulted in increased activity of the cytidine-5-diphosphate (CDP)-choline pathway. After continuous NO2 application,
- [1] specific activity of choline kinase,
- [2] cytidine triphosphate (CTP):cholinephosphate cytidylyltransferase,
- [3] uptake of choline, and
- [4] pool sizes of CDP-choline and PC were significantly increased over those of controls.
Intermittent application of this NO2 body dose provoked less increase in PC synthesis and the synthesis parameters were comparable to those for cells from control animals after repeated exposure. Whereas PC synthesis in type II cells was stimulated by NO2, their secretory activity was reduced. Continuous exposure reduced the secretory activity most, whereas intermittent exposure nonsignificantly reduced this activity as compared with that of controls. The repeated application of NO2 produced no differences.
The authors conclude that…. type II pneumocytes adapt to NO2 atmospheres depending on the mode of its application, at least for the metabolism of PC and its secretion from isolated type II pneumocytes.
The reader asks whether this effect could also be found in renal epithelial cells, for which PC is not considered vital as for type II pneumocytes and possibly related to surfactant activity in the lung.
Müller B, Seifart C, von Wichert P, Barth PJ. Adaptation of rat type II pneumocytes to NO2: effects of NO2 application mode on phosphatidylcholine metabolism. Am J Respir Cell Mol Biol. 1998; 18(5): 712-20.
iNOS involved in immediate response to anaphylaxis
The generation of large quantities of nitric oxide (NO) is implicated in the pathogenesis of anaphylactic shock. The source of NO, however, has not been established and conflicting results have been obtained when investigators have tried to inhibit its production in anaphylaxis.
This study analyzed the expression of inducible nitric oxide synthase (iNOS) and endothelial nitric oxide synthase (eNOS) in a mouse model of anaphylaxis. BALB/c mice were sensitized and challenged with ovalbumin to induce anaphylaxis. Tissues were removed from the heart and lungs, and blood was drawn at different time points during the first 48 hours after induction of anaphylaxis. The Griess assay was used to measure nitric oxide generation.
Nitric oxide synthase expression was examined by reverse transcriptase polymerase chain reaction and immunohistochemistry. A significant increase in iNOS mRNA expression and nitric oxide production was evident as early as 10 to 30 minutes after allergen challenge in both heart and lungs.
In contrast, expression of eNOS mRNA was not altered during the course of the experiment. The results support involvement of iNOS in the immediate physiological response of anaphylaxis.
Sade K, Schwartz IF, Etkin S, Schwartzenberg S, et al. Expression of Inducible Nitric Oxide Synthase in a Mouse Model of Anaphylaxis. J Investig Allergol Clin Immunol 2007; 17(6):379-385.
________________________________________________________________________________________________________________________________________________________
Part IIc. Additional Nonrenal Related NO References
1. Nitrogen dioxide induces death in lung epithelial cells in a density-dependent manner. Persinger RL, Blay WM, Heintz NH, Hemenway DR, Janssen-Heininger YM. Am J Respir Cell Mol Biol. 2001 May;24(5):583-90. PMID: 11350828 [PubMed – indexed for MEDLINE] Free Article
2. Molecular mechanisms of nitrogen dioxide induced epithelial injury in the lung. Persinger RL, Poynter ME, Ckless K, Janssen-Heininger YM. Mol Cell Biochem. 2002 May-Jun;234-235(1-2):71-80. Review. PMID: 12162462 [PubMed – indexed for MEDLINE]
3. Nitric oxide and peroxynitrite-mediated pulmonary cell death. Gow AJ, Thom SR, Ischiropoulos H. Am J Physiol. 1998 Jan;274(1 Pt 1):L112-8. PMID: 9458808 [PubMed – indexed for MEDLINE] Free Article
4. Mitogen-activated protein kinases mediate peroxynitrite-induced cell death in human bronchial epithelial cells. Nabeyrat E, Jones GE, Fenwick PS, Barnes PJ, Donnelly LE. Am J Physiol Lung Cell Mol Physiol. 2003 Jun;284(6):L1112-20. Epub 2003 Feb 21. PMID: 12598225 [PubMed – indexed for MEDLINE] Free Article
5. Peroxynitrite inhibits inducible (type 2) nitric oxide synthase in murine lung epithelial cells in vitro. Robinson VK, Sato E, Nelson DK, Camhi SL, Robbins RA, Hoyt JC. Free Radic Biol Med. 2001 May 1;30(9):986-91. PMID: 11316578 [PubMed – indexed for MEDLINE]
6. Nitric oxide-mediated chondrocyte cell death requires the generation of additional reactive oxygen species. Del Carlo M Jr, Loeser RF. Arthritis Rheum. 2002 Feb;46(2):394-403. PMID: 11840442 [PubMed – indexed for MEDLINE]
7. Colon epithelial cell death in 2,4,6-trinitrobenzenesulfonic acid-induced colitis is associated with increased inducible nitric-oxide synthase expression and peroxynitrite production.
Yue G, Lai PS, Yin K, Sun FF, Nagele RG, Liu X, Linask KK, Wang C, Lin KT, Wong PY. J Pharmacol Exp Ther. 2001 Jun;297(3):915-25. PMID: 11356911 [PubMed – indexed for MEDLINE] Free Article
__________________________________________________________________________________________________________________________________________________________
Part IIIa. Acute renal failure Acute renal failure (ARF), characterized by sudden loss of the ability of the kidneys to [1] excrete wastes, [2] concentrate urine, [3] conserve electrolytes, and [4] maintain fluid balance, is a frequent clinical problem, particularly in the intensive care unit, where it is associated with a mortality of between 50% and 80%.
This clinical entity was described as an acute loss of kidney function that occurred in severely injured crush victims because of histological evidence for patchy necrosis of renal tubules at autopsy. In the clinical setting, the terms ATN and acute renal failure (ARF) are frequently used interchangeably. However, ARF does not include increases in blood urea due to [1] reversible renal vasoconstriction (prerenal azotemia) or [2] urinary tract obstruction (postrenal azotemia). Acute hemodialysis was first used clinically during the Korean War in 1950 to treat military casualties, and this led to a decrease in mortality of the ARF clinical syndrome from about 90% to about 50%. In the half century that has since passed, much has been learned about the pathogenesis of ischemic and nephrotoxic ARF in experimental models, but there has been very little improvement in mortality. This may be explained by changing demographics: [1] the age of patients with ARF continues to rise, and [2] comorbid diseases are increasingly common in this population. Both factors may obscure any increased survival related to improved critical care. Examining the incidence of ARF in several military conflicts does, however, provide some optimism. The incidence of ARF in seriously injured casualties decreased between World War II and the Korean War, and again between that war and the Vietnam War, despite the lack of any obvious difference in the severity of the injuries. What was different was the rapidity of the fluid resuscitation of the patients? Fluid resuscitation on the battlefield with the rapid evacuation of the casualties to hospitals by helicopter began during the Korean War and was optimized further during the Vietnam War. For seriously injured casualties the incidence of ischemic ARF was one in 200 in the Korean War and one in 600 in the Vietnam War. This historical sequence of events suggests that early intervention could prevent the occurrence of ARF, at least in military casualties. In experimental studies it has been shown that progression from an azotemic state associated with renal vasoconstriction and intact tubular function (prerenal azotemia) to established ARF with tubular dysfunction occurs if the renal ischemia is prolonged. Moreover, early intervention with fluid resuscitation was shown to prevent the progression from prerenal azotemia to established ARF. Diagnostic evaluation of ARF One important question, therefore, is how to assure that an early diagnosis of acute renal vasoconstriction can be made prior to the occurrence of tubular dysfunction, thus providing the potential to prevent progression to established ARF. In this regard, past diagnostics relied on observation of the patient response to a fluid challenge: [1] decreasing levels of blood urea nitrogen (BUN) indicated the presence of reversible vasoconstriction, [2] while uncontrolled accumulation of nitrogenous waste products, i.e., BUN and serum creatinine, indicated established ARF.
This approach, however, frequently led to massive fluid overload in the ARF patient with resultant
- [1] pulmonary congestion,
- [2] hypoxia, and
- [3] premature need for mechanical ventilatory support and/or hemodialysis.
On this background the focus turned to an evaluation of urine sediment and urine chemistries to differentiate between renal vasoconstriction with intact tubular function and established ARF.
It was well established that if tubular function was intact, renal vasoconstriction was associated with enhanced tubular sodium reabsorption. Specifically, the fraction of filtered sodium that is rapidly reabsorbed by normal tubules of the vasoconstricted kidney is greater than 99%.
Thus, when nitrogenous wastes, such as creatinine and urea, accumulate in the blood due to a fall in glomerular filtration rate (GFR) secondary to renal vasoconstriction with intact tubular function, the fractional excretion of filtered sodium (FENa = [(urine sodium × plasma creatinine) / (plasma sodium × urine creatinine)]) is less than 1%. An exception to this physiological response of the normal kidney to vasoconstriction is when the patient is receiving a diuretic, including mannitol, or has glucosuria, which decreases tubular sodium reabsorption and increases FENa.
It has recently been shown in the presence of diuretics that a rate of fractional excretion of urea (FEurea) of less than 35 indicates intact tubular function, thus favoring renal vasoconstriction rather than established ARF as a cause of the azotemia.
___________________________________________________________________________________________________________________________________________________________
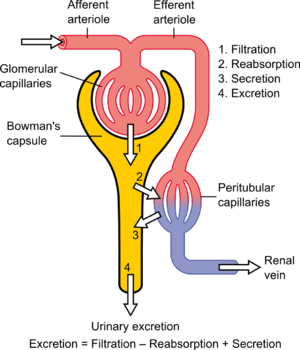
English: Physiology of Nephron (Photo credit: Wikipedia)

Structures of the kidney: 1.Renal pyramid 2.Interlobar artery 3.Renal artery 4.Renal vein 5.Renal hilum 6.Renal pelvis 7.Ureter 8.Minor calyx 9.Renal capsule 10.Inferior renal capsule 11.Superior renal capsule 12.Interlobar vein 13.Nephron 14.Minor calyx 15.Major calyx 16.Renal papilla 17.Renal column (no distinction for red/blue (oxygenated or not) blood, arteriole is between capilaries and larger vessels (Photo credit: Wikipedia)
_
______________________________________________________________________________________________________________________________________________________
Mechanisms of ARF
Based on the foregoing comments, this discussion of mechanisms of ARF will not include nitrogenous-waste accumulation due to renal vasoconstriction with intact tubular function (prerenal azotemia) or urinary tract obstruction (postrenal azotemia). The mechanisms of ARF involve both vascular and tubular factors. An ischemic insult to the kidney will in general be the cause of the ARF. While a decrease in renal blood flow with diminished oxygen and substrate delivery to the tubule cells is an important ischemic factor, it must be remembered that a relative increase in oxygen demand by the tubule is also a factor in renal ischemia.
Approximately 30–70% of these shed epithelial tubule cells in the urine are viable and can be grown in culture. Recent studies using cellular and molecular techniques have provided information relating to the structural abnormalities of injured renal tubules that occur both in vitro and in vivo. In vitro studies using chemical anoxia have revealed abnormalities in the proximal tubule cytoskeleton that are associated with translocation of Na+/K+-ATPase from the basolateral to the apical membrane.
A comparison of cadaveric transplanted kidneys with delayed versus prompt graft function has also provided important results regarding the role of Na+/K+-ATPase in ischemic renal injury. This study demonstrated that, compared with kidneys with prompt graft function, those with delayed graft function had a significantly greater cytoplasmic concentration of Na+/K+-ATPase and actin-binding proteins — spectrin (also known as fodrin) and ankyrin — that had translocated from the basolateral membrane to the cytoplasm.
Such a translocation of Na+/K+-ATPase from the basolateral membrane to the cytoplasm could explain the decrease in tubular sodium reabsorption that occurs with ARF. An important focus of research is the mechanisms whereby the critical residence of Na+/K+-ATPase in the basolateral membrane (which facilitates vectorial sodium transport) is uncoupled by hypoxia or ischemia. The actin-binding proteins,
serve as substrates for the calcium-activated cysteine protease calpain.
________________________________________________________________________________________________________________________________________________________
In vitro studies in proximal tubules have shown a rapid rise in cytosolic calcium concentration during acute hypoxia, which antedates the evidence of tubular injury as assessed by lactic dehydrogenase (LDH) release. There is further evidence to support the importance of the translocation of Na+/K+-ATPase from the basolateral membrane to the cytoplasm during renal ischemia/reperfusion.
Specifically, calpain-mediated breakdown products of the actin-binding protein spectrin occur with renal ischemia. Calpain activity was demonstrated to increase during hypoxia in isolated proximal tubules. Measurement of LDH release following calpain inhibition indicated attenuation of hypoxic damage to proximal tubules. There was no evidence of an increase in cathepsin, a (cysteine protease) in proximal tubules during hypoxia , but there is a calcium-independent pathway for calpain activation during hypoxia.
Calpastatin, an endogenous cellular inhibitor of calpain activation, was shown to be diminished during hypoxia in association with the rise in another cysteine protease, caspase.
This effect of diminished calpastatin activity could be reversed by caspase inhibition. Proteolytic pathways appear to be involved in calpain-mediated proximal tubule cell injury during hypoxia. Calcium activation of phospholipase A has also been shown to contribute to renal tubular injury during ischemia.
________________________________________________________________________________________________________________________________________________________
Tubular obstruction during ARF
The existence of proteolytic pathways involving cysteine proteases, namely calpain and caspases, may therefore explain
- the decrease in proximal tubule sodium reabsorption and
- increased FENa
secondary to proteolytic uncoupling of Na+/K+-ATPase from its basolateral membrane anchoring proteins.
This tubular perturbation alone, however, does not explain the fall in GFR that leads to nitrogenous-waste retention and thus the rise in BUN and serum creatinine. Decreased proximal tubule sodium reabsorption may lead to a decreased GFR during ARF. First of all, brush border membranes and cellular debris could provide the substrate for intraluminal obstruction in the highly resistant portion of distal nephrons.
In fact, microdissection of individual nephrons of kidneys from patients with ARF demonstrated obstructing casts in distal tubules and collecting ducts. This observation could explain the dilated proximal tubules that are observed upon renal biopsy of ARF kidneys. The intraluminal casts in ARF kidneys stain prominently for Tamm-Horsfall protein (THP), which is produced in the thick ascending limb. Importantly, THP is secreted into tubular fluid as a monomer but subsequently may become a polymer that forms a gel-like material in the presence of increased luminal Na+ concentration, as occurs in the distal nephron during clinical ARF with the decrease in tubular sodium reabsorption.
Thus, the THP polymeric gel in the distal nephron provides an intraluminal environment for distal cast formation involving viable, apoptotic, and necrotic cells.
The loss of the tubular epithelial cell barrier and/or the tight junctions between viable cells during acute renal ischemia could lead to a leak of glomerular filtrate back into the circulation. (If this occurs and normally non-reabsorbable substances, such as inulin, leak back into the circulation, then a falsely low GFR will be measured as inulin clearance. It should be noted, however, that the degree of extensive tubular damage observed in experimental studies demonstrating tubular fluid backleak is rarely observed with clinical ARF in humans). Moreover, dextran sieving studies in patients with ARF demonstrated that, at best, only a 10% decrease in GFR could be explained by backleak of filtrate. Cadaveric transplanted kidneys with delayed graft function, however, may have severe tubular necrosis, and thus backleak of glomerular filtration may be more important in this setting.
Inflammation and NO
There is now substantial evidence for the involvement of inflammation in the pathogenesis of the decreased GFR associated with acute renal ischemic injury. In this regard, there is experimental evidence that iNOS may contribute to tubular injury during ARF. Hypoxia in isolated proximal tubules has been shown to increase NO release, and there is increased iNOS protein expression in ischemic kidney homogenates. An antisense oligonucleotide was shown to block the upregulation of iNOS and afford functional protection against acute renal ischemia. Moreover, when isolated proximal tubules from iNOS, eNOS, and neuronal NO synthase (nNOS) knockout mice were exposed to hypoxia, only the tubules from the iNOS knockout mice were protected against hypoxia, as assessed by LDH release. The iNOS knockout mice were also shown to have lower mortality during ischemia/reperfusion than wild-type mice. The scavenging of NO by oxygen radicals produces peroxynitrite causing tubule damage during ischemia. While iNOS may contribute to ischemic injury of renal tubules, the vascular effect of eNOS in the glomerular afferent arteriole is protective against ischemic injury. In this regard, eNOS knockout mice are more sensitive to endotoxin-related injury than normal mice.
Moreover, the protective role of vascular eNOS may be more important than the deleterious effect of iNOS at the tubule level during renal ischemia. This is because treatment of mice with the nonspecific NO synthase (NOS) inhibitor L-NAME, which blocks both iNOS and eNOS, worsens renal ischemic injury. NO may downregulate eNOS and is a potent inducer of heme oxygenase-1, which has been shown to be cytoprotective against renal injury. The MAPK pathway also appears to be involved in renal oxidant injury. Activation of extracellular signal–regulated kinase (ERK) or inhibition of JNK ameliorates oxidant injury–induced necrosis in mouse renal proximal tubule cells in vitro. Upregulation of ERK may also be important in the effect of preconditioning whereby early ischemia affords protection against a subsequent ischemia/reperfusion insult. Alterations in cell cycling are also involved in renal ischemic injury. Upregulation of p21, which inhibits cell cycling, appears to allow cellular repair and regeneration, whereas homozygous p21 knockout mice demonstrate enhanced cell necrosis in response to an ischemic insult.
Prolonged duration of the ARF clinical course and the need for dialysis are major factors projecting a poor prognosis. Patients with ARF who require dialysis have a 50–70% mortality rate. Infection and cardiopulmonary complications are the major causes of death in patients with ARF. Excessive fluid administration in patients with established ARF may lead to pulmonary congestion, hypoxia, the need for ventilatory support, pneumonia, and multiorgan dysfunction syndrome, which has an 80–90% mortality rate. Until means to reverse the diminished host defense mechanisms in azotemic patients with clinical ARF are available, every effort should be made to avoid invasive procedures such as the placement of bladder catheters, intravenous lines, and mechanical ventilation. Over and above such supportive care, it may be that combination therapy will be necessary to prevent or attenuate the course of ARF. Such combination therapy must involve agents with potential beneficial effects on vascular tone, tubular obstruction, and inflammation.
Schrier RW, Wang W, Poole B, and Mitra A. Acute renal failure: definitions, diagnosis, pathogenesis, and therapy. The Journal of Clinical Investigation 2004; 114(1):5-14. http://www.jci.org
______________________________________________________________________________________________________________________________________________________
Part IIIb. Additional Related References on NO, oxidative stress and Kidney
Shelgikar PJ, Deshpande KH, Sardeshmukh AS, Katkam RV, Suryakarl AN. Role of oxidants and antioxidants in ARF patients undergoing hemodialysis. Indian J Nephrol 2005;15: 73-76.
Lee JW. Renal Dysfunction in Patients with Chronic Liver Disease. Electrolytes Blood Press 7:42-50, 2009ㆍdoi: 10.5049/EBP.2009.7.2.42.
Saadat H, et al. Endothelial Nitric Oxide Function and Tubular Injury in Premature Infants. Int J App Sci and Technol 2012; 7(6): 77-81. http://www.ijastnet.com.
Amerisan MS. Cardiovascular disease in chronic kidney disease. Indian J Nephrol 2005;15: 1-7.
___________________________________________________________________________________________________________________________________________________________
Traditional risk factors for CVD in CKD
- Hypertension
- Older Age
- Diabetes Mellitus
- Male gender
- High LDL
- White Race
- Low HDL
- Physical inactivity
- Smoking
- Menopause
- LVH
CKD Related CV Risk Factors
- Blood Pressure
- ? Homocysteinemia
- Anemia
- ? Inflammation
- Ca++ x P++
- ? NO synthesis
- Na+ Retention
- ? Lp (a)
- Hypervolemia
- ? Insulin Resistance
- Proteinuria & Hypoalbuminemia
- Iron over load
- ? Adeponectin
- ??Vit. C or E
- ? 5 Lipoxygenase
- ROS
- Genetic factors
- ADMA (Asymmetric Dimethyl Arginine)
S Vikrant, SC Tiwari. Essential Hypertension – Pathogenesis and Pathophysiology. J Indian Acad Clinical Medicine 2001; 2(3):141-161. Scheme for pathogenesis of salt dependent hypertension.
The hypothesis proposes that early hypertension is episodic and is mediated by a hyperactive sympathetic nervous system or activated renin-angiotensin system.
Cell membrane alterations
Hypotheses linking abnormal ionic fluxes to increased peripheral resistance through increase in cell sodium, calcium, or pH. The hypertension that is more common in obese people may arise in large part from the insulin resistance and resultant hyperinsulinaemia that results from the increased mass of fat. However, rather unexpectedly, insulin resistance may also be involved in hypertension in non-obese people.
Overall scheme for the mechanisms by which obesity, if predominantly upper body or visceral in location, could promote
________________________________________________________________________________________________________________________________________________________
- diabetes,
- dyslipidemia and
- hypertension via hyperinsulinemia.
The explanation for insulin resistance found in as many as half of nonobese hypertensive is not obvious and may involve one or more aspects of insulin’s action
__________________________________________________________________________________________________________________________________________________________
Proposed mechanisms by which insulin resistance and/or hyperinsulinemia may lead to increased blood pressure.
- Enhanced renal sodium and water reabsorption.
- Increased blood pressure sensitivity to dietary salt intake
- Augmentation of the pressure and
- aldosterone responses to AII
- Changes in transmembrane electrolyte transport
- a. Increased intracellular sodium
- b. Decreased Na+/K+ – ATPase activity
- c. Increased intracellular Ca2+ pump activity
- d. Increased intracellular Ca2+ accumulation
- e. Stimulation of growth factors
___________________________________________________________________________________________________________________________________________________________
Part IV. New Insights on NO donors
This study investigated the involvement of nitric oxide (NO) into the irradiation-induced increase of cell attachment. These experiments explored the cellular mechanisms of low-power laser therapy. HeLa cells were irradiated with a monochromatic visible-tonear infrared radiation (600–860 nm, 52 J/m2) or with a diode laser (820 nm, 8–120 J/m2) and the number of cells attached to a glass matrix was counted after 30 minute incubation at 37oC. The NO donors
- sodium nitroprusside (SNP),
- glyceryl trinitrate (GTN), or
- sodium nitrite (NaNO2)
were added to the cellular suspension before or after irradiation. The action spectra and the concentration and fluence dependencies obtained were compared and analyzed.
The well-structured action spectrum for the increase of the adhesion of the cells, with maxima at 619, 657, 675, 740, 760, and 820 nm, points to the existence of a photoacceptor responsible for the enhancement of this property (supposedly cytochrome c oxidase, the terminal respiratory chain enzyme), as well as signaling pathways between the cell mitochondria, plasma membrane, and nucleus.
Treating the cellular suspension with SNP before irradiation significantly modifies the action spectrum for the enhancement of the cell attachment property (band maxima at 642, 685, 700, 742, 842, and 856 nm). The action of SNP, GTN, andNaNO2 added before or after irradiation depends on their concentration and radiation fluence.
The NO donors added to the cellular suspension before irradiation eliminate the radiation induced increase in the number of cells attached to the glass matrix, supposedly by way of binding NO to cytochrome c oxidase. NO added to the suspension after irradiation can also inhibit the light-induced signal downstream. Both effects of NO depend on the concentration of the NO donors added.
The results indicate that NO can control the irradiation-activated reactions that increase the attachment of cells.
Karu TI, Pyatibrat LV, and Afanasyeva NI. Cellular Effects of Low Power Laser Therapy Can be Mediated by Nitric Oxide. Lasers Surg. Med 2005; 36:307–314.
IFNa-2b (IFN-a) effect on barrier function of renal tubular epithelium
IFNa treatment can be accompanied by impaired renal function and capillary leak. This study shows IFNa produced dose-dependent and time-dependent decrease in transepithelial resistance (TER) ameliorated by tyrphostin, an inhibitor of phosphotyrosine kinase with increased expression of occludin and E-cadherin. In conclusion, IFNa can directly affect barrier function in renal epithelial cells via ovewrexpression or missorting of the junctional proteins occludin and E-cadherin.
Lechner J, Krall M, Netzer A, Radmayr C, et al. Effects of interferon a-2b on barrier function and junctional complexes of renal proximal tubulat LLC-pK1 cells. Kidney Int 1999; 55:2178-2191.
Ischemia-reperfusion injury
The pathophysiology of acute renal failure (ARF) is complex and not well understood. Numerous models of ARF suggest that oxygen-derived reactive species are important in renal ischemia-reperfusion (I-R) injury, but the nature of the mediators is still controversial. Treatment with oxygen radical scavengers, antioxidants, and iron chelators such as
- superoxide dismutase,
- dimethylthiourea,
- allopurinol, and
- deferoxamine
are protective in some models, and suggest a role for the hydroxyl radical formation. However, these compounds are not protective in all models of I-R injury, and direct evidence for the generation of hydroxyl radical is absent. Furthermore, these inhibitors have another property in common.
They all directly scavenge or inhibit the formation of peroxynitrite (ONOO−), a highly toxic species derived from nitric oxide (NO) and superoxide. Thus, the protective effects seen with these inhibitors may be due in part to their ability to inhibit ONOO− formation. Even though reactive oxygen species are thought to participate in ischemia-reperfusion (I-R) injury, induction of and production of high levels of inducible nitric oxide (NO) also contribute to this injury.
NO combines with superoxide to form the potent oxidant peroxynitrite (ONOO−). NO and ONOO− were investigated in a rat model of renal I-R injury using the selective iNOS inhibitor L-N6-(1-iminoethyl)lysine (L-NIL).
I-R surgery significantly increased plasma creatinine levels to 1.9 ± 0.3 mg/dl (P < .05) and caused renal cortical necrosis. L-NIL administration (3 mg/kg) in animals subjected to I-R significantly decreased plasma creatinine levels to 1.2 ± 0.10 mg/dl (P < .05 compared with I-R) and reduced tubular damage.
ONOO− formation was evaluated by detecting 3-nitrotyrosine-protein adducts (3NTyPAs), a stable biomarker of ONOO− formation. The kidneys from I-R animals had increased levels of 3NTyPAs compared with control animals L-NIL-treated rats (3 mg/kg) subjected to I-R showed decreased levels of 3NTyPAs.
These results suggests that iNOS-generated NO mediates damage in I-R injury possibly through ONOO− formation.
______________________________________________________________________________________________________________________________________________________
In summary,
- 3-nitrotyrosine-protein adducts were detected in renal tubules after I-R injury.
- Selective inhibition of iNOS by L-NIL decreased injury, improved renal function, and decreased apparent ONOO− formation.
- Reactive nitrogen species should be considered potential therapeutic targets in the prevention and treatment of renal I-R injury.
_______________________________________________________________________________________________________________________________________________________
Walker LM, Walker PD, Imam SZ, et al. Evidence for Peroxynitrite Formation in Renal Ischemia-Reperfusion Injury: Studies with the Inducible Nitric Oxide Synthase InhibitorL-N6-(1-Iminoethyl)lysine1. 2000.
Role of TNFa independent of iNOS Renal failure is a frequent complication of sepsis, mediated by renal vasoconstrictors and vasodilators. Endotoxin induces several proinflammatory cytokines, among which tumor necrosis factor (TNF) is thought to be of major importance. Tumor necrosis factor (TNF) has been suggested to be a factor in the acute renal failure in sepsis or endotoxemia. Passive immunization by anti-TNFa prevented development of septic shock in animal experiments.The development of ARF involves excessive intrarenal vasoconstriction. Involvement of nitric oxide (NO), generated by inducible NO synthase (iNOS), is still a factor in the pathogenesis of endotoxin-induced renal failure. TNF-a leads to a decrease in glomerular filtration rate (GFR).
This study tested the hypothesis that the role of TNF-a in endotoxic shock related ARF is mediated by iNOS-derived NO. An injection of lipopolysaccharide (LPS) constituent of gram-negative bacteria to wild-type mice resulted in a 70% decrease in glomerular filtration rate (GFR) and in a 40% reduction in renal plasma flow (RPF) 16 hours after the injection. The results occurred independent of hypotension, morphological changes, apoptosis, and leukocyte accumulation. In mice pretreated with TNFsRp55, only a 30% decrease in GFR was observed without a significant change in RPF as compared with controls. Pretreatment with TNKsRp55 on renal function Wild-type mice were pretreated with TNFsRp55(10 mg/kg IP) for one hour before the administration of 5 mg/kg intraperitoneal endotoxin. GFR and RPF were determined 16 hours thereafter. Data are expressed as mean 6, SEM, N 5 6. *P , 0.05 vs. Control; §P , 0.05 vs. LPS, by ANOVA.
The serum NO concentration was significantly lower in endotoxemic wild-type mice pretreated with TNFsRp55, as compared with untreated endotoxemic wild-type mice. In LPS-injected iNOS knockout mice and wild-type mice treated with a selective iNOS inhibitor, 1400W, the development of renal failure was similar to that in wild-type mice. As in wild-type mice,TNFsRp55 significantly attenuated the decrease in GFR (a 33% decline, as compared with 75% without TNFsRp55) without a significant change in RPF in iNOS knockout mice given LPS. These results demonstrate a role of TNF in the early renal dysfunction (16 h) in a septic mouse model independent of iNOS,
- hypotension,
- apoptosis,
- leukocyte accumulation,and
- morphological alterations,
thus suggesting renal hypoperfusion secondary to an imbalance between, as yet to be defined renal vasoconstrictors and vasodilators.
Knotek M, Rogachev B, Wang W,….., Edelstein CL, Dinarello CA, and Schrier RW. Endotoxemic renal failure in mice: Role of tumor necrosis factor independent of inducible nitric oxide synthase. Kidney International 2001; 59:2243–2249
Ischemic acute renal failure
Inflammation plays a major role in the pathophysiology of acute renal failure resulting from ischemia. This review discusses the contribution of
- endothelial
- epithelial cells and
- leukocytes
to this inflammatory response. The roles of cytokines/chemokines in the injury and recovery phase are reviewed. The protection of mouse kidney prior to exposure to ischemia or urinary tract obstruction is a potential model to search for pharmacologic agents to protect the kidney against injury by inflammatory mediators produced by tubular epithelial cells and activated leukocytes in renal ischemia/reperfusion (I/R) injury. Tubular epithelia produce
- TNF-a,
- IL-1,
- IL-6,
- IL-8,
- TGF-b,
- MCP-1,
- ENA-78,
- RANTES, and
- fractalkines,
whereas leukocytes produce
- TNF-a,
- IL-1,
- IL-8,
- MCP-1,
- ROS, and
- eicosanoids.
The release of these chemokines and cytokines serve as effectors for a positive feedback pathway enhancing inflammation and cell injury, the cycle of tubular epithelial cell injury and repair following renal ischemia/reperfusion. Tubular epithelia are typically cuboidal in shape and apically-basally polarized; the Na+/K+-ATPase localizes to basolateral plasma membranes, whereas cell adhesion molecules, such as integrins localize basally. In response to ischemia reperfusion,
- the Na+/K+-ATPase appears apically, and
- integrins are detected on lateral and basal plasma membranes.
Some of the injured epithelial cells undergo necrosis and/or apoptosis detaching from the underlying basement membrane into the tubular space where they contribute to tubular occlusion. Viable cells that remain attached, dedifferentiate, spread, and migrate to repopulate the denuded basement membrane. With cell proliferation, cell-cell and cell-matrix contacts are restored, and the epithelium redifferentiates and repolarizes, forming a functional, normal epithelium Inflammation is a significant component of renal I/R injury, playing a considerable role in its pathophysiology.
Although significant progress has been made in defining the major components of this process, the complex cross-talk between endothelial cells, inflammatory cells, and the injured epithelium with each generating and often responding to cytokines and chemokines is not well understood. In addition, we have not yet taken full advantage of the large body of data on inflammation in other organ systems.
Furthermore, preconditioning the kidney to afford protection to subsequent bouts of ischemia may serve as a useful model challenging us to therapeutically mimic endogenous mechanisms of protection.
Understanding the inflammatory response prevalent in ischemic kidney injury will facilitate identification of molecular targets for therapeutic intervention.
Bonventre JV and Zuk A. Ischemic acute renal failure: An inflammatory disease? Forefronts in Nephrology 2002;.. :480-485
Gene expression profiles in renal proximal tubules In kidney disease renal proximal tubular epithelial cells (RPTEC) actively contribute to the progression of tubulointerstitial fibrosisby mediating both
- an inflammatory response and
- via epithelial-to-mesenchymal transition.
Using laser capture microdissection we specifically isolated RPTEC from cryosections of the healthy parts of kidneys removed owing to renal cell carcinoma and from kidney biopsies from patients with proteinuric nephropathies. RNA was extracted and hybridized to complementary DNA microarrays after linear RNA amplification. Statistical analysis identified 168 unique genes with known gene ontology association, which separated patients from controls. Besides distinct alterations in signal-transduction pathways (e.g. Wnt signalling), functional annotation revealed a significant upregulation of genes involved in
_________________________________________________________________________________________________________________________________________________________
- cell proliferation and cell cycle control (like insulin-like growth factor 1 or cell division cycle 34),
- cell differentiation (e.g. bone morphogenetic protein 7),
- immune response,
- intracellular transport and
- metabolism
__________________________________________________________________________________________________________________________________________________________
in RPTEC from patients.
The study also revealed differential expression of a number of genes responsible for cell adhesion (like BH-protocadherin) with a marked downregulation of most of these transcripts. In summary, the results obtained from RPTEC revealed a differential regulation of genes, which are likely to be involved in either pro-fibrotic or tubulo-protective mechanisms in proteinuric patients at an early stage of kidney disease.
Rudnicki M, Eder S, Perco P, Enrich J, et al. Gene expression profiles of human proximal tubular epithelial cells in proteinuric nephropathies. Kidney International 2006; xx:1-11. Kidney International advance online publication, 20 December 2006; doi:10.1038/sj.ki.5002043. http://www.kidney-international.org
Oxidative stress involved with diabetic nephropathy
Diabetic Nephropathy (DN) poses a major health problem. There is strong evidence for a potential role of the eNOS gene. This case control study investigated the possible role of genetic variants of the endothelial Nitric Oxide Synthase (eNOS) gene and oxidative stress in the pathogenesis of nephropathy in patients with diabetes mellitus. The study included 124 diabetic patients;
- 68 of these patients had no diabetic nephropathy (group 1) while
- 56 patients exhibited symptoms of diabetic nephropathy (group 2).
- Sixty two healthy non-diabetic individuals were also included as a control group.
Blood samples from subjects and controls were analyzed to investigate the eNOS genotypes and to estimate
- the lipid profile and
- markers of oxidative stress such as malondialdehyde (MDA) and nitric oxide (NO).
No significant differences were found in the frequency of eNOS genotypes between diabetic patients (either in group 1 or group 2) and controls (p >0.05). Also, no significant differences were found in the frequency of eNOS genotypes between group 1 and group 2 (p >0.05). Both group 1 and group 2 had significantly higher levels of nitrite and MDA when compared with controls (all p = 0.0001). Also group 2 patients had significantly higher levels of nitrite and MDA when compared with group 1 (p = 0.02, p = 0.001 respectively).
The higher serum level of the markers of oxidative stress in diabetic patients particularly those with diabetic nephropathy suggest that oxidative stress and not the eNOS gene polymorphism is involved in the pathogenesis of the diabetic nephropathy in this subset of patients
Badawy A, Elbaz R, Abbas AM, Ahmed Elgendy A, et al. Oxidative stress and not endothelial Nitric Oxide Synthase gene polymorphism involved in diabetic nephropathy. Journal of Diabetes and Endocrinology 2011; 2(3): 29-35.
Metformin in renal ischemia reperfusion
Renal ischemia plays an important role in renal impairment and transplantation. Metformin is a biguanide used in type 2 diabetes, it inhibits hepatic glucose production and increases peripheral insulin sensitivity. While the mode of action of metformin is incompletely understood, it appears to have anti-inflammatory and antioxidant effects involved in its beneficial effects on insulin resistance. Control, Sham, ischemia/reperfusion (I/R) and Metformin treated I /R groups A renal I/R injury was done by a left renal pedicle occlusion to induce ischemia for 45 min followed by 60 min of reperfusion with contralateral nephrectomy. Metformin pretreated I/R rats in a dose of 200 mg/kg/day for three weeks before ischemia induction.
- Nitric oxide (NO),
- tumor necrosis factor alpha (TNF α) ,
- catalase (CAT) and
- reduced glutathione (GSH) activities
were determined in renal tissue, while
- creatinine clearance (CrCl) ,
- blood urea nitrogen (BUN) were measured and
5 hour urinary volume and electrolytes were estimated . BUN and CrCl levels in the I/R group were significantly higher than in control rats (p<0.05) table (1).
__________________________________________________________________________________________________________________________________
Table 1: Creatinine clearance (Cr Cl) and blood urea nitrogen (BUN) levels in control and test groups.
(Mean ± SD)
| Groups |
CrCl (ml/min) |
BUN (mg/dl) |
| Control group |
1.30 ±0.11 |
14.30±0.25 |
| Sham group+ metformin |
1.27±0.09 |
15.70±0.19 |
| I/R group (P1) |
1.85±0.25 (<0.001 ) |
28.00±0.62 (<0.001) |
| I/R+ metformin group (P2,P3) |
1.55±0.22 (0.001, 0.028) |
18.10±1.00 (<0.001, <0.001) |
- P1: Statistical significance between control
group and saline treated I/R group.
- P2 Statistical significance between control
group and Metformin treated I/R group.
- P3 Statistical significance between saline treated
I/R group and Metformin treated I/R group
_______________________________________________________________________________________________________________________________________________________
When metformin was administered before I/R, BUN and CrCl levels were still significantly higher than control group but their elevation were significantly lower in comparison to I/R group alone (P<0.05). TNF α and NO levels were significantly higher in the I/R group than those of the control group (Table 2). Pre-treatment with metformin significantly lowered their levels in comparison to I/R group (P<0.05).
________________________________________________________________________________________________________________________________________________________
Table 2: Tumor necrosis factor α (TNF α) and inducible nitric oxide (iNO) levels in control and test groups.
(Mean ± SD)
| Groups |
TNF α (pmol/mg tissue) |
iNO (nmol/ mg tissue) |
| Control group 1 |
7.60 ±5.98 |
2.54 ± 0.82 |
| Sham group+ metformin |
16.70 ±5.50 |
2.35 ±0.80 |
| I/R group (P1) |
54. 00±6.02 (<0.001) |
4.50±0.89 (<0.001) |
| I/R+metformin group (P2,P3) |
39 ± 14.01 (<0.001, 0.006) |
3.53±0.95 (0.02, 0.03) |
- P1: Statistical significance between control group
and saline treated I/R group.
- P2 Statistical significance between control group
and Metformin treated I/R group.
- P3 Statistical significance between saline treated
I/R group and Metformin treated I/R group
_________________________________________________________________________________________________________________________________________________________
These results showed significant increase in NO,TNF α, BUN , CrCl and significant decrease in urinary volume , electrolytes, CAT and GSH activities in the I/R group than those in the control group. Metformin decreased significantly NO, TNF α, BUN and CrCl while increased urinary volume, electrolytes, CAT and GSH activities. Lipid peroxidation is related to I/R induced tissue injury. Production of inducible NO synthase (NOS) under lipid peroxidation and inflammatory conditions results in the induction of NO which react with O2 liberating peroxynitrite (OONO-). NO itself inactivates the antioxidant enzyme system CAT and GSH. Alteration in NO synthesis have been observed in other kidney injuries as nephrotoxicity and acute renal failure induced by endotoxins.
Treatment with iNOS inhibitors improved renal function and decreased peroxynitrite radical which is believed to be responsible for the shedding of proximal convoluted tubules in I/R. Metformin produced anti-inflammatory renoprotective effect on CrCl and diuresis in renal I/R injury.
Malek HA. The possible mechanism of action of metformin in renal ischemia reperfusion in rats. The Pharma Research Journal 2011; 6(1):42-49.
Possible role of NO donors in ARFThe L-arginine-nitric oxide (NO) pathway has been implicated in many physiological functions in the kidney, including
- regulation of glomerular hemodynamics,
- mediation of pressure-natriuresis,
- maintenance of medullary perfusion,
- blunting of tubuloglomerular feedback (TGF),
- inhibition of tubular sodium reabsorption and
- modulation of renal sympathetic nerve activity
Its net effect in the kidney is to promote natriuresis and diuresis, contributing to adaptation to variations of dietary salt intake and maintenance of normal blood pressure. Nitric oxide has been implicated in many physiologic processes that influence both acute and long-term control of kidney function. Its net effect in the kidney is to promote natriuresis and diuresis, contributing to adaptation to variations of dietary salt intake and maintenance of normal blood pressure. A pretreatment with nitric oxide donors or L-arginine may prevent the ischemic acute renal injury. In chronic kidney diseases, the systolic blood pressure is correlated with the plasma level of asymmetric dimethylarginine, an endogenous inhibitor of nitric oxide synthase. A reduced production and biological action of nitric oxide is associated with an elevation of arterial pressure, and conversely, an exaggerated activity may represent a compensatory mechanism to mitigate the hypertension.
JongUn Lee. Nitric Oxide in the Kidney : Its Physiological Role and Pathophysiological Implications. Electrolyte & Blood Pressure 2008; 6:27-34.
Renal Hypoxia and Dysoxia following Reperfusion
Acute renal failure (ARF) is a common condition which develops in 5% of hospitalized patients. Of the patients who develop ARF, ~10% eventually require renal replacement therapy. Among critical care patients who have acute renal failure and survive, 2%-10% develop terminal renal failure and require long-term dialysis. The kidneys are particularly susceptible to ischemic injury in many clinical conditions such as renal transplantation, treatment of suprarenal aneurysms, renal artery reconstructions, contrast-agent induced nephropathy, cardiac arrest, and shock. One reason for renal sensitivity to ischemia is that the kidney microvasculature is highly complex and must meet a high energy demand.
Under normal, steady state conditions, the oxygen (O2) supply to the renal tissues is well in excess of oxygen demand. Under pathological conditions, the delicate balance of oxygen supply versus demand is easily disturbed due to the unique arrangement of the renal microvasculature and its increasing numbers of diffusive shunting pathways.
The renal microvasculature is serially organized, with almost all descending vasa recta emerging from the efferent arterioles of the juxtamedullary glomeruli. Adequate tissue oxygenation is thus partially dependent on the maintenance of medullary perfusion by adequate cortical perfusion. This, combined with the low amount of medullary blood flow (~10% of total renal blood flow) in the U-shaped microvasculature of the medulla allows O2 shunting between the descending and ascending vasa recta and contributes to the high sensitivity of the medulla and cortico-medullary junction to decreased O2 supply.
Whereas past investigations have focused mainly on tubular injury as the main cause of ischemia-related acute renal failure, increasing evidence implicates alterations in the intra-renal microcirculation pathway and in the O2 handling. Indeed, although acute tubular necrosis (ATN) has classically been believed to be the leading cause of ARF, data from biopsies in patients with ATN have shown few or no changes consistent with tubular necrosis.
The role played by microvascular dysfunction, however, has generated increasing interest. The complex pathophysiology of ischemic ARF includes the inevitable
- reperfusion phase associated with oxidative stress,
- cellular dysfunction and
- altered signal transduction.
During this process, alterations in oxygen transport pathways can result in cellular hypoxia and/or dysoxia. In this context, the distinction between hypoxia and dysoxiais that
- cellular hypoxia refers to the condition of decreased availability of oxygen due to inadequate convective delivery from the microcirculation.
- Cellular dysoxia, in contrast, refers to a pathological condition where the ability of mitochondria to perform oxidative phosphorylation is limited, regardless of the amount of available oxygen.
_______________________________________________________________________________________________________________________________________________________
The latter condition is associated with mitochondrial failure and/or activation of alternative pathways for oxygen consumption. Thus, we would expect that an optimal balance between oxygen supply and demand is essential to reducing damage from renal ischemia-reperfusion (I/R) injury. Complex interactions exist between
- tubular injury,
- microvascular injury, and
- inflammation after renal I/R.
On the one hand, insults to the tubule cells promotes the liberation of a number of inflammatory mediators, such as TNF-á, IL-6, TGF-â, and chemotactic cytokines(RANTES, monocyte chemotactic protein-1, ENA-78, Gro-á, and IL-8). On the other hand, chemokine production can promote
- leukocyte-endothelium interactions and
- leukocyte activation,
resulting in…..
- renal blood flow impairment and
- the expansion of tubular damage
- impaired renal hemodynamics and
- electrolyte reabsorption
Adequate medullary tissue oxygenation, in terms of balanced oxygen supply and demand, is dependent on the maintenance of medullary perfusion by adequate cortical perfusion and also on the high rate of O2 consumption required for active electrolyte transport. Furthermore, renal blood flow is closely associated with renal sodium transport, mitochondrial activity and NO-mediated O2 consumption In addition to having a limited O2 supply due to the anatomy of the microcirculation anatomy, the sensitivity of the medulla to hypoxic conditions results from this high O2 consumption.
Renal sodium transport is the main O2-consuming function of the kidney and is closely linked to renal blood flow for sodium transport, particularly in the thick ascending limbs of the loop of Henle and the S3 segments of the proximal tubules. Medullary renal blood flow is also highly dependent on cortical perfusion, with almost all descending vasa recta emerging from the efferent arteriole of juxta medullary glomeruli. A profound reduction in cortical perfusion can disrupt medullary blood flow and lead to an imbalance between O2 supply and O2 consumption. On theother hand, inhibition of tubular reabsorption by diuretics increases medullary pO2 by decreasing the activity of Na+/K+-ATPases and local O2 consumption.
Mitochondrial activity and NO-mediated O2 consumption
The medulla has been found to be the main site of production of NO in the kidney. In addition to the actions described above, NO appears to be a key regulator of renal tubule cell metabolism by inhibiting the activity of the Na+-K+-2Cl- cotransporter and reducing Na+/H+ exchange. Since superoxide (O2-) is required to inhibit solute transport activity, it was assumed that these effects were mediated by peroxynitrite (OONO-). Indeed, mitochondrial nNOS upregulation, together with an increase in NO production, has been shown to increase mitochondrial peroxynitrite generation, which in turn, can induce cytochrome c release and promote apoptosis. NO has also been shown to directly compete with O2 at the mitochondrial level. These findings support the idea that NO acts as an endogenous regulator to match O2 supply to O2 consumption, especially in the renal medulla. NO reversibly binds to the O2 binding site of cytochrome oxidase, and acts as a potent, rapidMitochondrial activity and NO-mediated O2 consumption, and reversible inhibitor of cytochrome oxidase in competition with molecular O2. This inhibition could be dependent on the O2 level, since the IC50 (the concentration of NO that reduces the specified response by half) decreases with reduction in O2 concentration. The inhibition of electron flux at the cytochrome oxidase level switches the electron transport chain to a reduced state, and consequently leads to depolarization of the mitochondrial membrane potential and electron leakage.
To summarize, while the NO/O2 ratio can act as a regulator of cellular O2 consumption by matching decreases in O2 delivery to decreases in cellular O2 cellular, the inhibitory effect of NO on mitochondrial respiration under hypoxic conditions further impairs cellular aerobic metabolism. This leads to a state of “cytopathic hypoxia,” as described in the sepsis literature. Only cell-secreted NO competes with O2 and to regulate mitochondrial respiration. In addition to the 3 isoforms (eNOS, iNOS, cnNOS), an α-isoform of neuronal NOS, the mitochondrial isoform (mNOS) located in the inner mitochondrial membrane, has also been shown to regulate mitochondrial respiration. These data support a role for NO in the balanced regulation of renal O2 supply and O2 consumption after renal I/R However, the relationships between the determinants of O2 supply, O2 consumption, and renal function, and their relation to renal damage remain largely unknown.
Sustained endothelial activation Ischemic renal failure leads to persistent endothelial activation, mainly in the form of endothelium-leukocyte interactions and the activation of adhesion molecules. This persistent activation can compromise renal blood flow, prevent the recovery of adequate tissue oxygenation, and jeopardize tubular cell survival despite the initial recovery of renal tubular function. A 30-50% reduction in microvascular density was seen 40 weeks after renal ischemic injury in a rat model. Vascular rarefaction has been proposed to induce chronic hypoxia resulting in tubulointerstitial fibrosis via the molecular activation of fibrogenic factors such as transforming growth factor (TGF)-β, collagen, and fibronectin, all of which may play an important role in the progression of chronic renal disease.
Adaptation to hypoxia Over the last decade, the role of hypoxia-inducible factors (HIFs) in O2 supply and adaptation to hypoxic conditions has found increasing support. HIFs are O2-sensitive transcription factors involved in O2-dependent gene regulation that mediate cellular adaptation to O2 deprivation and tissue protection under hypoxic conditions in the kidney. NO generation can promote HIF-1α accumulation in a cGMP-independent manner. However, Hagen et al. (2003) showed that NO may reduce the activation of HIF in hypoxia via the inhibitory effect of NO on cytochrome oxidase.
Therefore, it seems that NO has pleiotropic effects on HIF expression, with various responses related to different pathways. HIF-1α upregulates a number of factors implicated in cytoprotection, including angiogenic growth factors, such as vascular endothelial growth factors (VEGF), endothelial progenitor cell recruitment via the endothelial expression of SDF-1, heme-oxygenase-1 (HO-1), and erythropoietin (EPO), and vasomotor regulation.
HO-1 produces carbon monoxide (a potent vasodilator) while degrading heme, which may preserve tissue blood flow during reperfusion. Thus, it has been suggested that the induction of HO-1 can protect the kidney from ischemic damage by decreasing oxidative damage and NO generation.
Finally, in addition to its anti-apoptotic properties, EPO may protect the kidney from ischemic damage by restoring the renal microcirculation by stimulating the mobilization and differentiation of progenitor cells toward an endothelial phenotype and by inducing NO release from eNOS.
Pharmacological interventions
Use of pharmacological interventions which act at the microcirculatory level may be a successful strategy to overcome ischemia-induced vascular damage and prevent ARF. Activated protein C (APC), an endogenous vitamin K-dependent serine protease with multiple biological activities, may meet these criteria. Along with antithrombotic and profibrinolytic properties, APC can reduce the chemotaxis and interactions of leukocytes with activated endothelium.
However, renal dysfunction was not improved in the largest study published so far. In addition, APC has been discontinued by Lilly for the use intended in severe sepsis. Moreover, neither drugs with renal vasodilatory effects (i.e., dopamine, fenoldopam, endothelin receptors blockers, adenosine antagonists) nor agents that decrease renal oxygen consumption (i.e., loop diuretics) have been shown to protect the kidney from ischemic damage. We have to bear in mind that a magic bullet to treat the highly complex condition of which is renal I/R is not in sight.
We can expect that understanding the balance between O2 delivery and O2 consumption, as well as the function of O2-consuming pathways (i.e., mitochondrial function, reactive oxygen species generation) will be central to this treatment strategy.
Take home point
The deleterious effects of NO are thought to be associated with the NO generated by the induction of iNOS and its contribution to oxidative stress both resulting in vascular dysfunction and tissue damage. Ischemic injury also leads to structural damage to the endothelium and leukocyte infiltration. Consequently, renal tissue hypoxia is proposed to promote the initial tubular damage, leading to acute organ dysfunction. Comment: I express great appreciation for refeering to this work, which does provide enormous new insights into hypoxia-induced acute renal failure, and ties together the anatomy, physiology, and gene regulation through signaling pathways.
Ince C, Legrand M, Mik E , Johannes T, Payen D. Renal Hypoxia and Dysoxia following Reperfusion of the Ischemic Kidney. Molecular Medicine (Proof) 2008; pp36. http://www.molmed.org
Nitric oxide and non-hemodynamic functions of the kidney
One of the major scientific advances in the past decade in understanding of the renal function and disease is the prolific growth of literature incriminating nitric oxide (NO) in renal physiology and pathophysiology. NO was first shown to be identical with endothelial derived relaxing factor (EDRF) in 1987 and this was followed by a rapid flurry of information defining the significance of NO in not only vascular physiology and hemodynamics but also in neurotransmission, inflammation and immune defense systems. Although most actions of NO are mediated by cyclic guanosine monophosphate (cGMP) signaling, S-nitrosylation of cysteine residues in target proteins constitutes another well defined non-cGMP dependent mechanism of NO effects. Recent years have witnessed a phenomenal scientific interest in the vascular biology, particularly the relevance of nitric oxide (NO) in cardiovascular and renal physiology and pathophysiology. Although hemodynamic actions of NO received initial attention, a variety of non-hemodynamic actions are now known to be mediated by NO in the normal kidney, which include
- tubular transport of electrolyte and water,
- maintenance of acid-base homeostasis,
- modulation of glomerular and interstitial functions,
- renin-angiotensin activation and
- regulation of immune defense mechanism in the kidney.
____________________________________________________________________________________________________________________________________________________________
Table 1 : Functions of NO in the kidney
- 1. Renal macrovascular and microvascular dilatation (afferent > efferent)
- 2. Regulation of mitochondrial respiration.
- 3. Modulation renal medullary blood flow
- 4. Stimulation of fluid, sodium and HCO3 – reabsorption in the proximal tubule
- 5. Stimulation of renal acidification in proximal tubule by stimulation of NHE activity
- 6. Inhibition of Na+, Cl- and HCO3 – reabsorption in the mTALH
- 7. Inhibition of Na+ conductance in the CCD
- 8. Inhibition of H+-ATPase in CCD
_____________________________________________________________________________________________________________________________________________________________
One of the renal regulatory mechanisms related to maintenance of arterial blood pressure involves the phenomenon of pressure-natriuresis in response to elevation of arterial pressure. This effect implies inhibition of tubular sodium reabsorption resulting in natriuresis, in an effort to lower arterial pressure. Experimental evidence from indicates that intra-renal NO modulates pressure natriuresis.
Furthermore many studies have confirmed the role of intra renal NO in mediating tubulo-glomerular feedback (TGF). In vivo micropuncture studies have shown that NO derived from nNOS in macula densa specifically inhibits the TGF responses leading to renal afferent arteriolar vasoconstriction in response to sodium reabsorption in the distal tubule. Other recent studies support the inhibitory role of NO from eNOS and iNOS in mTALH segment on TGF effects.
Recent observations in vascular biology have yielded new information that endothelial dysfunction early in the course might contribute to the pathophysiology of acute renal failure. Structural and functional changes in the vascular endothelium are demonstrable in early ischemic renal failure. Altered NO production and /or decreased bioavailability of NO comprise the endothelial function in acute renal failure.
Several studies have indicated imbalance of NOS activity with enhanced expression and activity of iNOS and decreased eNOS in ischemic kidneys.
The imbalance results from enhanced iNOS activity and attenuated eNOS activity in the kidney.
Many experimental studies support a contributory role for NO in glomerulonephritis (GN). Evidence from recent studies pointed out that NO may be involved in peroxynitrite formation, pro-inflammatory chemokines and signaling pathways in addition to direct glomerular effects that promote albumin permeability in GN. Although originally macrophages and other leukocytes were first considered as the source renal NO production in GN, it is now clear iNOS derived NO from glomerular mesangial cells are the primary source of NO in GN.
In most pathological states, the role of NO is dependent by the stage of the disease, the nitric oxide synthase (NOS) isoform involved and the presence or absence of other modifying intrarenal factors. Additionally NO may have a dual role in several disease states of the kidney such as acute renal failure, inflammatory nephritides, diabetic nephropathy and transplant rejection.
A rapidly growing body of evidence supports a critical role for NO in tubulointerstitial nephritis (TIN). In the rat model of autoimmune TIN, Gabbai et al. demonstrated increased iNOS expression in the kidney and NO metabolites in urine and plasma. However the effects of iNOS on renal damage in TIN seem to have a biphasic effect- since iNOS specific inhibitors (eg. L-Nil) are renoprotective in the acute phase while they actually accelerated the renal damage in the chronic phase.
Thus chronic NOS inhibition is used to induce chronic tubulointerstitial injury and fibrosis along with mild glomerulosclerosis and hypertension.
Major pathways of L-arginine metabolism.
L-arginine may be metabolized by the urea cycle enzyme arginase to L-ornithine and urea by arginine decarboxylase to agmatine and CO2 or by NOS to nitric oxide (NO) and L-citrulline.
Adapted from Klahr S: Can L-arginine manipulation reduce renal disease? Semin Nephrol 1999; 61:304-309.
It is obvious that kidney is not only a major source of arginine and nitric oxide but NO plays an important role in the water and electrolyte balance and acid-base physiology and many other homeostatic functions in the kidney. Unfortunately we are far from a precise understanding of the significance of NO alterations in various disease states primarily due to conflicting data from the existing literature.
Therapeutic potential for manipulation of L-arginine- nitric oxide axis in renal disease states has been discussed. More studies are required to elucidate the abnormalities in NO metabolism in renal diseases and to confirm the therapeutic potential of L-arginine.
Sharma SP. Nitric oxide and the kidney. Indian J Nephrol 2004;14: 77-84
Inhibition of Constitutive Nitric Oxide Synthase
Excess NO generation plays a major role in the hypotension and systemic vasodilatation characteristic of sepsis. Yet the kidney response to sepsis is characterized by vasoconstriction resulting in renal dysfunction. We have examined the roles of inducible nitric oxide synthase (iNOS) and endothelial NOS (eNOS) on the renal effects of lipopolysaccharide administration by comparing the effects of specific iNOS inhibition, L-N6-(1-iminoethyl)lysine (L-NIL), and 2,4-diamino-6-hydroxy-pyrimidine vs. nonspecific NOS inhibitors (nitro-L-arginine-methylester). cGMP responses to carbamylcholine (CCh) (stimulated, basal) and sodium nitroprusside in isolated glomeruli were used as indices of eNOS and guanylate cyclase (GC) activity, respectively. LPS significantly decreased blood pressure and GFR (P =0.05) and inhibited the cGMP response to CCh.
GC activity was reciprocally increased. L-NIL and 2,4-diamino-6-hydroxy-pyrimidine administration prevented the decrease in GFR, restored the normal response to CCh, and GC activity was normalized. In vitro application of L-NIL also restored CCh responses in LPS glomeruli. Neuronal NOS inhibitors verified that CCh responses reflected eNOS activity.
L-NAME, a nonspecific inhibitor, worsened GFR, a reduction that was functional and not related to glomerular thrombosis, and eliminated the CCh response. No differences were observed in eNOS mRNA expression among the experimental groups. Selective iNOS inhibition prevents reductions in GFR, whereas nonselective inhibition of NOS further decreases GFR.
These findings suggest that the decrease in GFR after LPS is due to local inhibition of eNOS by iNOS, possibly via NO autoinhibition.
Schwartz D, Mendonca M, Schwartz I, Xia Y, et al. Inhibition of Constitutive Nitric Oxide Synthase (NOS) by Nitric Oxide Generated by Inducible NOS after Lipopolysaccharide Administration Provokes Renal Dysfunction in Rats. J. Clin. Invest. 1997; 100:439–448.
Salt-Sensitivity and Hypertension Renin-angiotensin system (RAS) plays a key role in the regulation of renal function, volume of extracellular fluid and blood pressure. The activation of RAS also induces oxidative stress, particularly superoxide anion (O2-) formation.
Although the involvement of O2 – production in the pathology of many diseases is known for long, recent studies also strongly suggest its physiological regulatory function of many organs including the kidney. However, a marked accumulation of O2- in the kidney alters normal regulation of renal function and may contribute to the development of salt-sensitivity and hypertension.
In the kidney, O2- acts as vasoconstrictor and enhances tubular sodium reabsoption. Nitric oxide (NO), another important radical that exhibits opposite effects than O2 -, is also involved in the regulation of kidney function. O2- rapidly interacts with NO and thus, when O2- production increases, it diminishes the bioavailability of NO leading to the impairment of organ function. As the activation of RAS, particularly the enhanced production of angiotensin II, can induce both O2- and NO generation, it has been suggested that physiological interactions of
provide a coordinated regulation of kidney function. The imbalance of these interactions is critically linked to the pathophysiology of salt-sensitivity and hypertension.
Kopkan L, Červenka L. Renal Interactions of Renin-Angiotensin System, Nitric Oxide and Superoxide Anion: Implications in the Pathophysiology of Salt-Sensitivity and Hypertension. Physiol. Res. 2009; 58 (Suppl. 2): S55-S67.
Epicrisis
In this review I attempted to evaluate complex and still incomplete and conflicting conclusions from many studies. I thus broke the report into three major portions:
___________________________________________________________________________________________________________________________________________________________
- 1 The kidney and its anatomy, physiology, and ontogeny.
- 2 The pathological disease variation affecting the kidney
- a: a tie in to eNON and iNos, nitric oxide, cGMP and glutaminase – in acute renal failure, hypertension, chronic renal failure, dialysis the pathology of acute tubular necrosis, glomerular function, efferent arteriolar and kidney medullary circulatory impairment, and cast formation related to Tamm Horsfall protein
- b :The role of NO, eNOS and iNOS in disorders of the lund alveolar cell and subendothelial matrix, and of liver disease also affecting the kidney, and the heart. c Additional references
- 3. a Acute renal failure, oxidate stress, ischemia-reperfusion injury, tubulointerstitial chronic inflammation
- 3 b Additional references 4. Nitric oxide donors – opportunities for therapeutic targeting? As we see this in as full a context as possible, it is hard to distinguish the cart from the horse.
___________________________________________________________________________________________________________________________________________________________
We know that there is an unquestionable role of NO, and a competing balance to be achieved between eNOS, iNOS, an effect on tubular water and ion-cation reabsorptrion, a role of TNFa, and consequently an important role in essential/malignant hypertension, with the size of the effect related to the stage of disorder, the amount of interstitial fibrosis, the remaining nephron population, the hypertonicity of the medulla, the vasodilation of the medullary circulation, and the renin-angiotensin-aldosterone system. Substantial data and multiple patients with many factors per patient would be need to extract the best model using a supercomputer.
Like this:
Like Loading...
Read Full Post »