Brain Development
Larry H. Bernstein, MD, FCAP, Curator
LPBI
Updated 11/22/2015
Single Gene Found to Play Huge Role in Brain Development

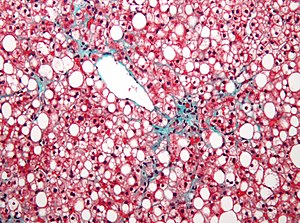
Figure 1: Cells in which NeuroD1 is turned on are reprogrammed to become neurons. Cell nuclei are shown in blue (Höchst stain) and neurons are shown in red (stained with neuronal marker TUJ1). [A. Pataskar,J. Jung, V. Tiwari]
Researchers at the Institute of Molecular Biology (IMB) in Mainz, Germany say they have unraveled a complex regulatory mechanism that explains how a single gene can drive the formation of brain cells. Their study (“NeuroD1 reprograms chromatin and transcription factor landscapes to induce the neuronal program”), published in The EMBO Journal, is an important step toward a better understanding of how the brain develops. It also harbors potential for regenerative medicine, according to the scientists.
Neurodegenerative disorders, such as Parkinson’s disease, are often characterized by an irreversible loss neurons. Unlike many other cell types in the body, neurons are generally not able to regenerate by themselves, so if the brain is damaged, it stays damaged. One hope of developing treatments for this kind of damage is to understand how the brain develops in the first place, and then try to imitate the process. However, the brain is also one of the most complex organs in the body, and very little is understood about the molecular pathways that guide its development.

Figure 2: Diagram showing how NeuroD1 influences the development of neurons. During brain development, expression of NeuroD1 marks the onset of neurogenesis. NeuroD1 accomplishes this via epigenetic reprogramming: neuronal genes are switched on, and the cells develop into neurons. TF: transcription factor; V: ventricle; P: pial surface. [A. Pataskar, J. Jung, V. Tiwari]
ijay Tiwari, Ph.D, and his group have been investigating a central gene in brain development, NeuroD1. This gene is expressed in the developing brain and marks the onset of neurogenesis.
In their research article, Dr. Tiwari and his colleagues have shown that during brain development NeuroD1 is not only expressed in brain stem cells but acts as a master regulator of a large number of genes that cause these cells to develop into neurons. They used a combination of neurobiology, epigenetics, and computational biology approaches to show that these genes are normally turned off in development, but NeuroD1 activity changes their epigenetic state in order to turn them on. Strikingly, the researchers show that these genes remain switched on even after NeuroD1 is later switched off. They further show that this is because NeuroD1 activity leaves permanent epigenetic marks on these genes that keep them turned on, in other words it creates an epigenetic memory of neuronal differentiation in the cell.
“Our research has shown how a single factor, NeuroD1, has the capacity to change the epigenetic landscape of the cell, resulting in a gene expression program that directs the generation of neurons,” wrote the screenplay investigators.
“This is a significant step toward understanding the relationship between DNA sequence, epigenetic changes and cell fate. It not only sheds new light on the formation of the brain during embryonic development but also opens up novel avenues for regenerative therapy,” says Dr. Tiwari.
NEUROD1 neuronal differentiation 1 [ Homo sapiens (human) ]
Official Symbol NEUROD1 provided by HGNC
Official Full Name neuronal differentiation 1 provided by HGNC
Primary source HGNC:HGNC:7762 See related Ensembl:ENSG00000162992; HPRD:03428; MIM:601724; Vega:OTTHUMG00000132583
Gene type protein coding
RefSeq status REVIEWED
OrganismHomo sapiens
Lineage Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Euteleostomi; Mammalia; Eutheria; Euarchontoglires; Primates; Haplorrhini; Catarrhini; Hominidae; Homo
Also known asBETA2; BHF-1; MODY6; NEUROD; bHLHa3
Summary This gene encodes a member of the NeuroD family of basic helix-loop-helix (bHLH) transcription factors. The protein forms heterodimers with other bHLH proteins and activates transcription of genes that contain a specific DNA sequence known as the E-box. It regulates expression of the insulin gene, and mutations in this gene result in type II diabetes mellitus. [provided by RefSeq, Jul 2008]
https://en.wikipedia.org/wiki/NEUROD1
Neurogenic differentiation 1 (NeuroD1), also called β2,[1] is a transcription factor of the NeuroD-type. It is encoded by the human gene NEUROD1.
It is a member of the NeuroD family of basic helix-loop-helix (bHLH) transcription factors. The protein forms heterodimers with other bHLH proteins and activates transcription of genes that contain a specific DNA sequence known as the E-box. It regulates expression of the insulin gene, and mutations in this gene result in type II diabetes mellitus.[2]
Contents [hide]
NeuroD1 induces terminal neuronal differentiation in olfactory neurogenesis
Camille Boutin, Olaf Hardt, Antoine de Chevigny, Nathalie Coré, Sandra Goebbels, Ralph Seidenfaden, Andreas Bosio and Harold Cremer
PNAS Jan 19, 2010; 107(3): 1201–1206. http://dx.doi.org:/10.1073/pnas.0909015107
After their generation and specification in periventricular regions, neuronal precursors maintain an immature and migratory state until their arrival in the respective target structures. Only here are terminal differentiation and synaptic integration induced. Although the molecular control of neuronal specification has started to be elucidated, little is known about the factors that control the latest maturation steps. We aimed at identifying factors that induce terminal differentiation during postnatal and adult neurogenesis, thereby focusing on the generation of periglomerular interneurons in the olfactory bulb. We isolated neuronal precursors and mature neurons from the periglomerular neuron lineage and analyzed their gene expression by microarray. We found that expression of the bHLH transcription factor NeuroD1 strikingly coincides with terminal differentiation. Using brain electroporation, we show that overexpression of NeuroD1 in the periventricular region in vivo leads to the rapid appearance of cells with morphological and molecular characteristics of mature neurons in the subventricular zone and rostral migratory stream. Conversely, shRNA-induced knockdown of NeuroD1 inhibits terminal neuronal differentiation. Thus, expression of a single transcription factor is sufficient to induce neuronal differentiation of neural progenitors in regions that normally do not show addition of new neurons. These results suggest a considerable potential of NeuroD1 for use in cell-therapeutic approaches in the nervous system.
Determination of neuronal subtypes is an early event that coincides with cell cycle exit (1, 2). However, after their generation, new neurons have to remain immature for prolonged periods, allowing their migration to final destinations where terminal differentiation occurs (3). Little is known about the factors that maintain the precursor state or induce terminal differentiation.
Olfactory neurogenesis is particularly suited to approach these late steps in neuronal differentiation. Here, stem cell populations first located in the ventricular zone and after the establishment of an ependymal layer positioned in subventricular zone (SVZ) generate migratory neuroblasts throughout life (4). These perform long-distance chain migration via the rostral migratory stream (RMS) into the olfactory bulb (OB), where they migrate into the granule cell layer (GCL) and the glomerular layer (GL) to differentiate into GABA- and dopaminergic neurons (4, 5). Thus, in this system, generation of neurons is permanent and the consecutive steps in the neurogenic sequence are spatially separated.
Determination of newly generated neurons has been studied intensively over the past years. For example, it has been demonstrated that defined areas surrounding the lateral ventricle contain predetermined stem cells that give rise to defined subsets of interneurons (6, 7). Several transcription factors have been implicated in the specification of the different neuronal populations. The zinc finger transcription factor sp8, for instance, appears to be involved in the generation of interneurons expressing calretinin (8), and analysis of Sall3 mutant mice (9) points to a role of this factor in the dopaminergic, tyrosine hydroxylase–positive lineage (10). Furthermore, it appears that interneuron diversity relies on the combinatorial expression of such transcription factors. This is exemplified by Pax6 and Dlx2, which have been shown to interact in the determination of adult generated neuronal precursors toward a dopaminergic fate (9, 11, 12). All of these transcriptional regulators are expressed early during the neurogenic process and remain present until terminal differentiation occurs.
We aimed at the identification of transcription factors that induce terminal differentiation of postnatal generated neurons in the OB. To do so we isolated neuronal precursors and differentiated interneurons from the periglomerular lineage of the OB and compared their gene expression by microarray. We established that the expression of NeuroD1, a bHLH transcription factor that has been implicated in neuronal differentiation in several experimental systems (13–17), coincides with the passage from neuronal precursor to mature interneurons. Functionally, we show that premature expression of NeuroD1 in vitro and in vivo induced highly efficiently the differentiation of forebrain progenitors. In vivo, this leads to the transitory appearance of ectopic neurons in the SVZ, RMS and striatum. Conversely, knockdown of NeuroD1 specifically inhibits terminal maturation of periglomerular neurons in the OB. Thus, NeuroD1 is both necessary and sufficient to induce key steps in terminal neuronal differentiation.
NeuroD1 Is Specifically Expressed in Mature GL Interneurons.
Subpopulations of neuronal precursors destined for the GCL and GL of the OB are generated by regionally defined stem cell populations in the periventricular region but migrate intermingled in the RMS to the OB. Once there, cells resegregate: granule cell precursors terminate their migration in the GCL, whereas the smaller population of periglomerular neuron precursors traverses this layer and the mitral cell layer (MCL) to invade the peripherally located GL (Fig. 1A). Thus, at a given time point, the GL contains both mature periglomerular neurons and their specific progenitors. Based on this spatial organization we isolated these two populations, concurrently depleting glial cells.
We devised a three-step strategy based on the following: (i) microdissection followed by enzymatic dissociation of the postnatal GL, (ii) depletion of contaminating glial cells by magnetic activated cell sorting (MACS) using an A2B5 specific antibody (18), (iii) separation of PSA-NCAM expressing cells (19) from the remaining fraction containing the mature neurons (Fig. 1B). The same purification strategy was applied to tissue microdissected from the P2 periventricular region (18). Characterization of the different cell population after sorting was performed via immunocytochemistry using the markers used for sorting (A2B5 and PSA-NCAM) as well as the differentiation marker Gad65 (18) (Fig. S1). Thus, as starting material we obtained highly enriched mature OB periglomerular interneurons (PGN), their immature progenitors (PGP), as well as a mixed population of generic progenitors (GP) from the SVZ/RMS.

Fig. 1.
Expression of NeuroD1 in the olfactory neurogenic system (A) DAPI-stained coronal section through the olfactory bulb of P5. (B) Strategy to isolate neuronal populations at different steps of their maturation. (C) Relative changes in gene expression for selected genes. Expression in GP was considered baseline, and changes are expressed as fold difference. (D–F) NeuroD1 in situ hybridization on sections from P5 mouse brain. No signal was detected along the lateral ventricle or in the RMS (D). In the olfactory bulb, individual NeuroD1+ cells were present in the GCL, whereas the MCL and the GL contained higher amounts (E, high magnification in F). A similar expression pattern was found after β-gal reaction on NeuroD1-lacZ-knockin tissue (G). (Scale bar: 200 μm in A; 100 μm inD and E; 20 μm in F and G).
Based on the purified and characterized cell populations, we performed microarray analyses to gain insight into the changes in gene expression during the neurogenic process. Investigation of expression dynamics of genes associated with either the precursor status or neuronal differentiation (Fig. S2 A and B) were used to validate the approach. Furthermore, these data were compared with those from an already available Serial Analysis of Gene Expression (SAGE) study (20).
Serial Analysis of Microarray (SAM) demonstrated the presence of groups of genes with comparable expression patterns (Fig. S2 C–E). Interestingly, only a relatively small fraction of genes were absent in the immature cell populations GP and PGP but highly represented in mature PGN (Fig. S2E). One of the genes showing such a pattern was NeuroD1, which was expressed more than 50-fold higher in PGN than in the immature populations (Fig. 1C). This was in agreement with the above-cited SAGE data, showing that NeuroD1 expression was below the detection level in neuronal precursors of the adult SVZ (20). Thus, expression of NeuroD1 was absent from precursors but coincided with terminal neuronal differentiation.
This late expression of NeuroD1 was in contrast to that of factors that have been functionally implicated in the specification of PGN, including Pax6, Sp8 and Sall3, which were expressed in both the immature populations and in the mature neurons (Fig. 1C; in situ hybridization for Pax6 in Fig. S3). Only Dlx2 showed a moderate increase in the PGN lineage outgoing, however, from an already considerable baseline level in migrating precursors (12) (Fig. 1C).
Next we analyzed the expression of NeuroD1 using in situ hybridization on P5 forebrain sections. Strong expression was found in the GL, whereas weaker expression was observed in the GCL and MCL (Fig. 1 Eand F). The transcript was undetectable in the periventricular region and the RMS (Fig. 1D). This staining was confirmed using NeuroD1-lacZ knockin mice (21) (Fig. 1G). In conclusion, these data demonstrated the absence of NeuroD1 from immature cells of the system and its strong expression in mature PGN. This pattern was coherent with a function in terminal neuronal differentiation.
NeuroD1 Induces Neuronal Differentiation in Vitro.
We studied the neurogenic potential of NeuroD1 in primary cultured neural stem cells using the neurosphere assay. In parallel to NeuroD1, we performed all experiments under the same conditions using the transcription factor Pax6, a well-described neurogenic signal in the system (9, 11, 12), to control for specificity of the observed effects. Neurosphere cells were coelectroporated with NeuroD1 or Pax6 expression vectors and GFP immediately before plating in differentiation conditions. One week after transfection, in the control condition, 14 ± 1% of the GFP-positive cells coexpressed the early neuronal marker Tuj1 (Fig. S4 A and D) whereas NeuroD1 induced Tuj1 expression in virtually all cells (98.0 ± 2%, Fig. S4 B and D). Pax6 gain-of-function led to an intermediate value (60.0 ± 3%, Fig. S4 C and D). NeuN, a later neuronal marker (22), was expressed by 21.1 ± 1% of the Tuj1-positive cells in the control situation (Fig. S4 E and H) but induced by NeuroD1 in almost all cells (93.9 ± 2%; Fig. S4 F and H). Surprisingly, Pax6 expression led to nearly complete disappearance of NeuN (1.7 ± 0.3%; Fig. S4 G and H). We investigated the induction of subtype specific markers by NeuroD1. Whereas tyrosine hydroxylase showed no augmentation, we found a 20% increase in calretinin labeling, in agreement with previous findings (23).
Next we investigated morphological parameters like process length as well as density and length of filopodia. Both NeuroD1 and Pax6 induced a significant, greater than 2-fold increase in process length (Fig. S4 I and L). We analyzed dendritic filopodia, structures that are believed to be precursors of dendritic spines (24). Expression of NeuroD1 induced a doubling in density and length of filopodia (Fig. S4 N, P, and Q). Interestingly, Pax6 reduced filopodia density to a level significantly below that of controls (Fig. S4 O and P), whereas length of the few remaining filopodia was not affected (7.0 ± 0.4 μm; Fig. S4Q).
Thus, the expression of NeuroD1 in neurosphere amplified neural stem cells induced neuronal commitment as well as morphological characteristics of mature neurons. Like NeuroD1, Pax6 favored neuronal commitment but appeared to actively suppress certain characteristics of terminal neuronal differentiation.
NeuroD1 Induces Ectopic Neurons in Vivo.
We asked whether NeuroD1 was also sufficient to induce neuronal differentiation in vivo. We used postnatal forebrain electroporation, an approach that allows efficient genetic manipulation of neural stem cells along the lateral ventricles and, consequently, of all transitory or permanent cell populations that are generated in the olfactory neurogenic process (25). The NeuroD1 expression vector or empty control plasmids were coelectroporated together with a GFP-containing vector that allowed visualization of transfected cells and their progeny at high resolution. Consequences of NeuroD1 gain-of-function were analyzed at 2, 4, 6, 8, and 15 days postelectroporation (dpe). As for the in vitro studies, results were compared with the effects of Pax6 gain-of-function.
At 2 dpe of a control vector into the lateral wall of the forebrain ventricle, 9.8 ± 1.3% (Fig. 2 A and K) of the GFP-expressing cells were localized in the VZ and had the morphology of radial glia (RG) (25). The majority of the GFP + cells, representing mainly neuronal precursors, were localized in the SVZ. Electroporation of a NeuroD1 expression vector induced a loss of GFP-positive RG cells (3.7 ± 0.5%; Fig. 2 B and K). The remaining cells in the VZ showed lower GFP levels than in controls (Fig. 2 A and B asterisks).

Fig. 2.
NeuroD1 induces neuronal morphology in vivo. Effect of NeuroD1 gain-of-function at different time points postelectroporation. (A and B) Coronal forebrain sections at the level of the lateral ventricle at 2 dpe. In the control condition, strongly GFP labeled RG are present in the VZ (A, asterisk). Expression of NeuroD1 induced a relative loss of radial glia and fainter GFP label (B, asterisk). (C and D) Coronal sections at the level of the lateral ventricle at 4 dpe. NeuroD1 expression induced an accumulation of transfected cells in the SVZ (D) and the almost total disappearance of radial glia (D). (E–F′) Sagittal sections of the RMS at 4 dpe. In the control situation, cells migrated toward the OB and presented the bipolar morphology specific of migrating precursors (E, E′, arrowheads). NeuroD1 electroporation induced loss of tangential orientation, induction of complex branching (F, F′, arrowhead), and invasion of the surrounding tissues (F, arrowheads). (G and H) Coronal section at the level of the olfactory bulb at 4 dpe. Although the majority of cells have reached the OB in the control situation (G), only a few cells were located in the center of the OB in the presence of NeuroD1 (H). (I and I′) Examples of cells presenting neuronal morphology in the SVZ at 4 dpe. (J) High magnification showing the presence of filopodia covering NeuroD1-expressing cells (arrowheads). (K) Quantification of GFP-positive cells presenting radial glia cell morphology along the lateral ventricle at 2 and 4 dpe. Control: 9.8 ± 1.3% (n= 6) at 2 dpe; 24 ± 11.8% at 4 dpe (n = 3); NeuroD1: 3.7 ± 0.5% at 2 dpe (n = 6); 1.6 ± 0.7% at 4 dpe (n = 3). (l) Distribution of the GFP-positive cells along the rostrocaudal axis. NeuroD1 expressing cells accumulated in proximal parts of the system. (M) Morphological analysis of cells in the SVZ/RMS. Three different classes were defined: (i) bipolar cells presenting tangential orientation, (ii) spherical cells, and (iii) branched cells presenting multiple processes in various directions (compare I). NeuroD1-expressing cells presented a highly branched morphology. Control: bipolar, 80.4%; spherical, 19.5%; branched, 0% (n = 133 cells). NeuroD1: bipolar, 5%; spherical, 16.8%; branched, 78% (n = 119 cells). Statistics: Mann-Whitney test. ns, not significant. **P < 0.01; ***P < 0.005. (Scale bar: 100 μm in E, F, G,and H; 25 μm in A, B, C, D,E, and F’; 10 μm in I; 5 μm in J.)
At 4 dpe, in the control situation, considerable amounts of strongly GFP+ RG cells were still present in the VZ (Fig. 2C asterisks), whereas NeuroD1 expression induced an almost complete loss of RG cells (Fig. 2 Dand K). At this time point, control cells were found along the entire SVZ and RMS. They showed generally tangential orientation and the typical morphology of migratory neuronal precursors. Large amounts of such cells were also found in the center of the OB (Fig. 2 G and L). NeuroD1 expression induced an accumulation of GFP-labeled cells in the SVZ (Fig. 2 D and L) at the expense of cells in the RMS (Fig. 2H,quantified in Fig. 2L). The accumulating cells did not have the appearance of migrating precursors but displayed complex multibranched morphologies (Fig. 2 F and F′, examples in Fig. 2 I and I′, quantified inFig. 2M). All principal processes of these cells were covered with small protrusions resembling filopodia (Fig. 2J). Such morphologically complex cells, strongly resembling neurons, were also predominant in and along proximal parts of the RMS (Fig. 2F′). Interestingly, considerable amounts of multibranched cells were found outside of the periventricular region and the RMS, invading neighboring structures such as the striatum (Fig. 2F, arrows). There was a clear correlation between the quantity of transgene expression, as visualized by GFP fluorescence, and the above parameters. Thus, NeuroD1 induced dose-dependently a neuron-like morphology in cells in the SVZ, RMS, and surrounding tissues.
We characterized the NeuroD1 induced neuron-like cell population in the periventricular region using neuronal and glial markers (Fig. 3; examples in Fig. S5). Doublecortin (DCX), a microtubule-associated protein expressed in migratory neuronal precursors (26), was seen in 75.2 ± 4.5% of the cells in the control situation but showed a significant increase after expression of NeuroD1 (91.7 ± 2.2%). NeuN, a marker for most mature neuronal cell types in the brain (22) was low in controls (5.2 ± 1.4%, n = 8) but strongly induced by NeuroD1 (65.9 ± 4.5%, n = 9). Map2, a later generic neuronal marker (27), was also rare in control cells (14.1 ± 1.4%, n = 3) but highly expressed in the NeuroD1 condition (61.9 ± 2.7%, n = 3). GFAP and Olig2 did not show significant alterations due to NeuroD1 expression. Thus, the NeuroD1-induced ectopic cells with neuronal morphology in the SVZ and RMS showed molecular characteristics of neurons.

Fig. 3.
NeuroD1 induces generic neuronal markers in vivo Molecular phenotype of the cells located in the periventricular region (level 4 in Fig. 2l). Quantification representing the percentage of GFP-positive cells expressing the respective markers. DCX: control, 75.2 ± 4.5%, n = 5; NeuroD1, 91.7 ± 2.2%, n = 5. NeuN: control, 5.2 ± 1.4%, n = 8; NeuroD1, 65.9 ± 4.5%, n = 9. Map2: control, 14.1 ± 1.4%, n = 3; NeuroD1, 61.9 ± 2.7%, n = 3. Olig2: control, 6.8 ± 5%, n = 3; NeuroD1, 2.5 ± 0.5%, n = 3. GFAP: control, 0%, n = 3; NeuroD1, 0%, n = 2. Errors bars indicate SEM. Statistics: DCX and Map2, unpaired ttest; NeuN, Mann-Whitney test. ns, not significant. *P < 0.05; **P < 0.01; ***P < 0.005.
HighWire Press-hosted articles citing this article
-
……..
NeuroD1 Is Necessary For OB Interneuron Differentiation in Vivo.
Next we asked whether NeuroD1 is essential for the generation of PGN. Given that NeuroD1 deficiency in mice is generally associated with perinatal lethality (14, 15, 21), we used a strategy based on RNAi in concert with postnatal in vivo electroporation to knock down NeuroD1 in the olfactory bulb neurogenic system. For validation, three different NeuroD1 specific shRNA vectors were cotransfected with a NeuroD1 expression construct into COS-7 cells. Western blot analysis demonstrated that two of the shRNAs, sh775 and sh776, efficiently inhibited production of the NeuroD1 protein, whereas sh777 induced a less efficient downregulation (sh775, 94.6%; sh776, 96.9%; sh777, 78.4%; corrected for loading against αtubulin; Fig. 4A). All three shRNAs were used for further in vivo studies.

Fig. 4.
In vivo terminal neuronal differentiation of PGC is impaired in absence of NeuroD1. (A) Western blot analysis of protein extracts from cos-7 cells transfected with NeuroD1 or in combination with different NeuroD1 specific shRNAs. sh775 and sh776 strongly repressed NeuroD1 protein expression (94.6% and 96.9%, respectively), whereas sh777 repressed NeuroD1 by 74.8%. (B–H′′) Consequences of loss-of-function of NeuroD1 via in vivo postnatal electroporation at 4 and 15 dpe. (B–E) No differences were observed at the level of the lateral ventricle or in the RMS at 4 dpe. (F) Cell distribution along the rostro-caudal axis was normal (definition of levels in Fig. 2l). (G and H′′) Consequences of NeuroD1 knockdown on PGN morphology at 15 dpe. (G) Whereas shRNAs showing a strong effect on NeuroD1 expression strongly inhibited morphological differentiation, the weakly active shRNA 777 had only a minor effect compared with control. (H) Examples of cells that served for classification of PGN. Class1 cells present primary and secondary branching. Dendritic spines (arrowheads) indicate their synaptic integration in OB circuitry. Class 2 cells present a single primary branch. Class 3 cells present a spherical morphology and no branching. Errors bars indicate SEM. Statistics, unpaired t test. ns, not significant. **P < 0.01; ***P < 0.005. (Scale bar: 100 μm in B–E; 20 μm in H.
………
When the two highly active NeuroD1-specific shRNAs sh775 and sh776 were electroporated, the vast majority of cells in the GL showed simple morphologies with few or no processes (classes 2 and 3), whereas cells with complex neuronal morphologies were sparse (Fig. 4G). When the less-efficient shRNA sh777 was expressed, an intermediate degree of neuronal maturation was observed (Fig. 4G), suggesting a dose-dependent action of NeuroD1 under these conditions. Comparable results were obtained for the GCL. As in the PGL, knockdown of NeuroD1 induced a dose-dependent inhibition of terminal neuronal differentiation (Fig. S9 A and B).
Thus, knockdown of NeuroD1 did not notably interfere with early steps of interneuron generation, but induced a specific defect in the acquisition of the differentiated neuronal phenotype in the OB.
Discussion
Although considerable information is available concerning the generation, specification, and migration of neurons, little is known concerning the factors and regulatory cascades that maintain the immature neuronal precursor status or induce the exit from this state and trigger terminal differentiation. Using a systematic approach, we identified NeuroD1 as a candidate for the latter function and validated this role using gain- and loss-of function approaches.
In Xenopus, a late function of NeuroD1 has been suggested based on two lines of evidence (13). First, NeuroD1 is transitorily expressed in territories where neuronal differentiation occurs. Second, misexpression of NeuroD1 causes the premature differentiation of neuronal precursors into neurons. However, the observation that NeuroD1 could also convert presumptive epidermal cells into neurons pointed toward a determination function. Therefore, a doubtless discrimination between a proneural and a terminal differentiation function was not possible.
The above-cited pioneering work in the frog has been extended through the analysis of mice with mutations in the NeuroD1 gene (14, 15, 21). In the hippocampal dentate gyrus of such animals, granule cell precursors are generated correctly in the neuroepithelium and invade the hippocampal anlage. However, in the target structure, precursors show a severe deficit in proliferation, and a defined dentate gryus is not formed (15). In the mutant cerebellum, generation and migration of early precursors appear not to be affected. Nevertheless, once these cells become postmitotic, massive cell death is observed and the cerebellum is severely affected (14). Thus, in these systems a late function of NeuroD1 is already suggested. However, because of the complexity of the models and the relatively low level of resolution, the available information is still fragmentary.
We attempted to clarify the role of NeuroD1 in neuronal differentiation by analyzing its function during olfactory neurogenesis. Using SAGE, microarray, in situ hybridization, and lacZ knockin into the NeuroD1 locus, we have demonstrated that NeuroD1 is expressed in mature neurons of the OB but is absent from immature stages. These findings are in contrast to recent expression data based on a NeuroD1 antibody, suggesting expression of the transcription factor already in the SVZ and RMS (23, 29). However, our loss-of-function approach based on RNAi shows that NeuroD1 is dispensable for generation and migration of precursors but is necessary for their transition into neurons in the target layer. These findings are in agreement with those of a recent study based on conditional NeuroD1 mutants, which showed a comparable defect in the OB (29).
……
This work demonstrates that expression of a single transcription factor can induce massive ectopic neuronal differentiation of neural stem cells in the vertebrate forebrain. The existence of postnatal and adult neurogenesis holds potential for the treatment of neurodegenerative diseases (34). However, in many experimental paradigms, transplanted or recruited cells fail to undergo differentiation into neurons and either transdifferentiate into glia or remain immature precursors (18, 35). It appears conceivable to combine such approaches with the strong neuronal differentiation inducing activity of NeuroD1.
Scientists Unveil Critical Mechanism of Memory Formation
http://www.dddmag.com/news/2015/11/scientists-unveil-critical-mechanism-memory-formation