Heart Vasculature – Regeneration and Protection of Coronary Artery Endothelium and Smooth Muscle: A Concept-based Pharmacological Therapy of a Combination Three Drug Regimen including THYMOSIN
Author & Curator: Aviva Lev-Ari, PhD, RN
ABSTRACT
A concept-based original pharmacological therapy was developed for the research results presented in Cell by Wu, Fujiwara, Cibulsky et al. (2006), Moretti, Caron, Nakano, et al. (2006) and for the research results in Nature by Smart, Risebro, Melville, et al. (2007). We propose the following concept-based original pharmacological therapy design for Preoperative and Postoperative management of cardiac injury to heart tissue, smooth muscle, to aorta and coronary artery disease. This is a treatment for Coronary Vasculogenesis, Anti-hypertention (short-acting), Vascular Anti-inflammation (vasculitis), Neovascularization of ischemic tissue and release of adult epicardium from a quiescent state while restoring its pluripotency.
VIEW VIDEO
What are Induced Pluripotent Stem Cells? (iPS Cells)
http://www.youtube.com/watch?v=i-QSurQWZo0
Lasker Lecture: Dr. Shinya Yamanaka, 2 of 3:
Induced Pluripotent Stem Cells? (iPS Cells)
http://www.youtube.com/watch?v=DQNoyDwCPzM
Multipotent Embryonic Isl1^+ Progenitor Cells Lead to Cardiac, Smooth Muscle, and Endothelial Cell Diversification
Alessandra A Moretti, Leslie L Caron, Atsushi A Nakano, Jason T JT Lam, Alexandra A Bernshausen,Yinhong Y Chen, Yibing Y Qyang, Lei L Bu, Mika M Sasaki, Silvia S Martin-Puig, Yunfu Y Sun, Sylvia M SM Evans, Karl-Ludwig KL Laugwitz, Kenneth R KR Chien
Cell 127(6):15 (2006), PMID 17123592
Cardiogenesis requires the generation of endothelial, cardiac, and smooth muscle cells, thought to arise from distinct embryonic precursors. We use genetic fate-mapping studies to document that isl1^+ precursors from the second heart field can generate each of these diverse cardiovascular cell types in vivo. Utilizing embryonic stem (ES) cells, we clonally amplified a cellular hierarchy of isl1^+ cardiovascular progenitors, which resemble the developmental precursors in the embryonic heart. The transcriptional signature of isl1^+/Nkx2.5^+/flk1^+ defines a multipotent cardiovascular progenitor, which can give rise to cells of all three lineages. These studies document a developmental paradigm for cardiogenesis, where muscle and endothelial lineage diversification arises from a single cell-level decision of a multipotent isl1^+ cardiovascular progenitor cell (MICP). The discovery of ES cell-derived MICPs suggests a strategy for cardiovascular tissue regeneration via their isolation, renewal, and directed differentiation into specific mature cardiac, pacemaker, smooth muscle, and endothelial cell types.
http://pubget.com/paper/17123592/Multipotent_Embryonic_Isl1___Progenitor_Cells_Lead_to_Cardiac__Smooth_Muscle__and_Endothelial_Cell_Diversification
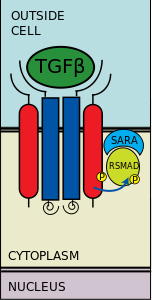
Thymosin beta 4 (Tβ4)
is a highly conserved, 43-amino acid acidic peptide (pI 4.6) that was first isolated from bovine thymus tissue over 25 years ago. It is present in most tissues and cell lines and is found in high concentrations in blood platelets, neutrophils, macrophages, and other lymphoid tissues. Tβ4 has numerous physiological functions, the most prominent of which being the regulation of actin polymerization in mammalian nucleated cells and with subsequent effects on actin cytoskeletal organization, necessary for cell motility, organogenesis, and other important cellular events.
Recently,
- Tβ4 was shown to be expressed in the developing heart and found to stimulate migration of cardiomyocytes and endothelial cells, promote survival of cardiomyocytes (Nature, 2004), and most recently
- to play an essential role in all key stages of cardiac vessel development: vasculogenesis, angiogenesis, and arteriogenesis (Nature 2006).
- These results suggest that Tβ4 may have significant therapeutic potential in humans to protect myocardium and promote cardiomyocyte survival in the acute stages of ischemic heart disease.
RegeneRx Biopharmaceuticals, Inc. is developing Tβ4 for the treatment of patients with acute myocardial infarction (AMI). Such efforts presented will include the formulation, development, and manufacture of a suitable drug product for use in the clinic, the performance of nonclinical pharmacology and toxicology studies, and the implementation of a phase 1 clinical protocol to assess the safety, tolerability, and the pharmacokinetics of Tβ4 in healthy volunteers.
http://onlinelibrary.wiley.com/doi/10.1196/annals.1415.051/abstract;jsessionid=BB7CC897572B7DDB60370EA64A81FC3F.d01t03?deniedAccessCustomisedMessage=&userIsAuthenticated=false
EXPLORATIONS with THYMOSIN beta4 for INDUCING ADULT EPICARDIAL PROGENETOR MOBILIZATION AND NEOVASCULARIZATION is presented in
Resident-cell-based Therapy in Human Ischaemic Heart Disease: Evolution in the PROMISE of Thymosin beta4 for Cardiac Repair
http://pharmaceuticalintelligence.com/2012/04/30/93/
EXPLORATIONS with THYMOSIN beta4 for INDUCTION of ARTERIOGENESIS, Prevention and repair of damaged cardiac tissue post MI and other CVD related research projects are presented in
Arteriogenesis and Cardiac Repair: Two Biomaterials – Injectable Thymosin beta4 and Myocardial Matrix Hydrogel
http://pharmaceuticalintelligence.com/2013/02/27/arteriogenesis-and-cardiac-repair-two-biomaterials-injectable-thymosin-beta4-and-myocardial-matrix-hydrogel/
Recent research results with THYMOSIN beta4 in use for Cardiovascular Disease
appeared in 2010:
Annals of the New York Academy of Sciences, May 2010 Volume 1194 Pages ix–xi, 1–230
http://onlinelibrary.wiley.com/doi/10.1111/nyas.2010.1194.issue-1/issuetoc
appeared in 2012:
- Thymosins in Health and Disease II: 3rd International Symposium on The Emerging Clinical Applications of Tymosin beta 4 in Cardiovascular Disease
Annals of the New York Academy of Sciences, October 2012 Volume 1270 Pages vii-ix, 1–121.
http://onlinelibrary.wiley.com/doi/10.1111/nyas.2012.1270.issue-1/issuetoc
Allan L. Goldstein, Enrico Garaci, Editors, Thymosins in Cardiovascular Disease, November 2012, Wiley-Blackwell (paperback)
http://www.wiley.com/WileyCDA/WileyTitle/productCd-1573319104.html?cid=RSS_WILEY2_LIFEMED
Selected for this article are the abstracts of the following research projects, all were presented at the 2nd International Symposium, May 2010:
Thymosin β4: structure, function, and biological properties supporting current and future clinical applications
Published studies have described a number of physiological properties and cellular functions of thymosin β4 (Tβ4), the major G-actin-sequestering molecule in mammalian cells. Those activities include the promotion of cell migration, blood vessel formation, cell survival, stem cell differentiation, the modulation of cytokines, chemokines, and specific proteases, the upregulation of matrix molecules and gene expression, and the downregulation of a major nuclear transcription factor. Such properties have provided the scientific rationale for a number of ongoing and planned dermal, corneal, cardiac clinical trials evaluating the tissue protective, regenerative and repair potential of Tβ4, and direction for future clinical applications in the treatment of diseases of the central nervous system, lung inflammatory disease, and sepsis. A special emphasis is placed on the development of Tβ4 in the treatment of patients with ST elevation myocardial infarction in combination with percutaneous coronary intervention, pp.179-189, May 2010.
Thymosin β4 and cardiac repair
Hypoxic heart disease is a predominant cause of disability and death worldwide. As adult mammals are incapable of cardiac repair after infarction, the discovery of effective methods to achieve myocardial and vascular regeneration is crucial. Efforts to use stem cells to repopulate damaged tissue are currently limited by technical considerations and restricted cell potential. We discovered that the small, secreted peptide thymosin β4 (Tβ4) could be sufficiently used to inhibit myocardial cell death, stimulate vessel growth, and activate endogenous cardiac progenitors by reminding the adult heart on its embryonic program in vivo. The initiation of epicardial thickening accompanied by increase of myocardial and epicardial progenitors with or without infarction indicate that the reactivation process is independent of injury. Our results demonstrate Tβ4 to be the first known molecule able to initiate simultaneous myocardial and vascular regeneration after systemic administration in vivo. Given our findings, the utility of Tβ4 to heal cardiac injury may hold promise and warrant further investigation, pp. 87-96, May 2010.
Thymosin β4 facilitates epicardial neovascularization of the injured adult heart
Ischemic heart disease complicated by coronary artery occlusion causes myocardial infarction (MI), which is the major cause of morbidity and mortality in humans
http://www.who.int/cardiovascular_diseases/resources/atlas/en/index.html
After MI the human heart has an impaired capacity to regenerate and, despite the high prevalence of cardiovascular disease worldwide, there is currently only limited insight into how to stimulate repair of the injured adult heart from its component parts. Efficient cardiac regeneration requires the replacement of lost cardiomyocytes, formation of new coronary blood vessels, and appropriate modulation of inflammation to prevent maladaptive remodeling, fibrosis/scarring, and consequent cardiac dysfunction. Here we show that thymosin β4 (Tβ4) promotes new vasculature in both the intact and injured mammalian heart. We demonstrate that limited EPDC-derived endothelial-restricted neovascularization constitutes suboptimal “endogenous repair,” following injury, which is significantly augmented by Tβ4 to increase and stabilize the vascular plexus via collateral vessel growth. As such, we identify Tβ4 as a facilitator of cardiac neovascularization and highlight adult EPDCs as resident progenitors which, when instructed by Tβ4, have the capacity to sustain the myocardium after ischemic damage, pp. 97-104, May 2010.
Thymosin β4: a key factor for protective effects of eEPCs in acute and chronic ischemia
Acute myocardial infarction is still one of the leading causes of death in the industrial nations. Even after successful revascularization, myocardial ischemia results in a loss of cardiomyocytes and scar formation. Embryonic EPCs (eEPCs), retroinfused into the ischemic region of the pig heart, provided rapid paracrine benefit to acute and chronic ischemia in a PI-3K/Akt-dependent manner. In a model of acute myocardial ischemia, infarct size and loss of regional myocardial function decreased after eEPC application, unless cell pre-treatment with thymosin β4 shRNA was performed. Thymosin ß4 peptide retroinfusion mimicked the eEPC-derived improvement of infarct size and myocardial function. In chronic ischemia (rabbit model), eEPCs retroinfused into the ischemic hindlimb enhanced capillary density, collateral growth, and perfusion. Therapeutic neovascularization was absent when thymosin ß4 shRNA was introduced into eEPCs before application. In conclusion, eEPCs are capable of acute and chronic ischemia protection in a thymosin ß4 dependent manner, pp. 105-111, May 2010.
Clinical Study Data of Thymosin beta 4 Presented
Published on October 3, 2009 at 5:10 AM
REGENERX BIOPHARMACEUTICALS, INC. (NYSE Amex:RGN) today reported on several clinical studies with Thymosin beta 4 (Tβ4) presented the Second International Symposium on Thymosins in Health and Disease, in Catania, Italy. The following are synopses of the presentations:
Myocardial Development of RGN-352 (Injectable Tβ4 Peptide)
David Crockford, RegeneRx’s vice president for clinical and regulatory affairs presented an overview of the biological properties that support Tβ4’s near term and long term clinical applications. Mr. Crockford noted that special emphasis is being placed on the development of RGN-352 for the systemic (injectable) treatment of patients with ST-elevation myocardial infarction (STEMI) in combination with percutaneous coronary intervention, the current standard of care in most western countries for this common type of heart attack. The goal with RGN-352 is to prevent or repair continued damage to cardiac tissue post-heart attack, when such tissue around the damaged site remains at risk.
Dr. Dennis Ruff, vice president and medical director of ICON, and principal investigator, presented the most current results on the Phase I safety study with RGN-352 entitled, “A Randomized, Double-blind, Placebo-controlled, Dose-response Phase I Study of the Safety and Tolerability of the Intravenous Administration of Thymosin Beta 4 and its Pharmacokinetics After Single and Multiple Doses in Healthy Volunteers.” Dr. Ruff discussed key aspects of the study and concluded with, “There were no dose limiting or serious adverse events throughout the dosing period. Synthetic Tβ4 administered intravenously up to 1260 mg, and for up to 14 days, appears to be well tolerated with low incidence of adverse events and no evidence of serious adverse events.”
http://www.news-medical.net/news/20091003/Clinical-study-data-of-Thymosin-beta-4-presented.aspx
RegeneRx Receives Notice of Allowance from Chinese Patent Office for Treatment and Prevention of Heart Disease
RegeneRx Receives Notice of Allowance from Chinese Patent Office for Treatment and Prevention of Heart Disease
February 7, 2013 — Rockville, MD
RegeneRx Biopharmaceuticals, Inc. (OTC Bulletin Board: RGRX) (“the Company” or “RegeneRx”) today announced that it has received a Notice of Allowance of a Chinese patent application for uses of Thymosin beta 4 (TB4) for treating, preventing, inhibiting or reducing heart tissue deterioration, injury or damage in a subject with heart failure disease. Claims also include uses for restoring heart tissue in those subjects. The patent will expire July 26, 2026 http://www.regenerx.com/wt/page/pr_1360265259
Theoretical treatment protocol differential between the Preoperative which may be between 3 to 6 month, and the Postoperative which may prolong to one year.
Proposal for Preoperative Treatment – Three drug combination involves
- Drug # 1: Thymosin fraction 5 (a sublingual composition)
- Drug # 2: Indomethacin (Nonsteroidal anti-inflammatory drugs (NSAID))
- Drug # 3: Clevidipine (blood pressure lowering drug, (no effect on heart rate))
Proposal for Postoperative Treatment – Three drug combination consists of
- Drug # 1: Thymosin fraction 5 (a sublingual composition)
- Drug # 4: ACEI (Captopril (50mg))
- Drug # 5: Beta Blocker and Diuretic (Metoprolol and hydrochlorothiazide (50 mg/25 mg)) Lopressor HCT
Unprecedented novel paradigm development in the scientific understanding of the origin of
- (a) myocardial cells
- (b) smooth muscle cells
- (c) endothelial cells
- (d) pace maker cells and
- (e) heart vasculature: aorta, pulmonary artery and coronary arteries, occurred in 2006.
In a seminal article in Cell, “Developmental Origin of a Bipotential Myocardial and Smooth Muscle Cell Precursor in the Mammalian Heart” Wu, et al., (2006), described their discovery as follows:
“Despite recent advances in delineating the mechanisms involved in cardiogenesis, cellular lineage specification remains incompletely understood.” To explore the relationship between developmental fate and potential.” They “isolated a cardiac-specific Nkx2.5+ cell population from the developing mouse embryo. The majority of these cells differentiated into cardiomyocytes and conduction system cells. Some, surprisingly, adopted a smooth muscle fate. To address the clonal origin of these lineages, we isolated Nkx2.5+ cells from in vitro differentiated murine embryonic stem cells and found ~28% of these cells expressed c-kit. These c-kit+ cells possessed the capacity for long-term in vitro expansion and differentiation into both cardiomyocytes and smooth muscle cells from a single cell.” They “confirmed these findings by isolating c-kit+Nkx2.5+ cells from mouse embryos and demonstrated their capacity for bipotential differentiation in vivo. Taken together, these results support the existence of a common precursor for cardiovascular lineages in the mammalian heart.”
Another breakthrough article in Cell, “Multipotent Embryonic Isl1+ Progenitor Cells Lead to Cardiac, Smooth Muscle, and Endothelial Cell Diversification” Moretti, et al., (2006) described their discovery as follows:
“Cardiogenesis requires the generation of endothelial, cardiac, and smooth muscle cells, thought to arise from distinct embryonic precursors.” They “use genetic fate-mapping studies to document that isl1+ precursors from the second heart field can generate each of these diverse cardiovascular cell types in vivo. Utilizing embryonic stem (ES) cells”, they “clonally amplified a cellular hierarchy of isl1+ cardiovascular progenitors, which resemble the developmental precursors in the embryonic heart. The transcriptional signature of isl1+/Nkx2.5+/flk1+ defines a multipotent cardiovascular progenitor, which can give rise to cells of all three lineages. These studies document a developmental paradigm for cardiogenesis, where muscle and endothelial lineage diversification arises from a single cell-level decision of a multipotent isl1+ cardiovascular progenitor cell (MICP). The discovery of ES cell-derived MICPs suggests a strategy for cardiovascular tissue regeneration via their isolation, renewal, and directed differentiation into specific mature cardiac, pacemaker, smooth muscle, and endothelial cell types.” (Moretti, et al., 2006).
Third scientific breakthrough was reported in Nature on the roles that Thymosin beta4 play in
- (a) coronary vessel development
- (b) induction of adult epicardial cell migration
- (c) cardiomyocyte survival by vascularization which is dependent on Thymosin beta4 and
- (d) identification of the pro-angiogenic tetrapeptide AcSDKP which is produced by endoproteinase activity of Thymosin beta4 (Smart, et al., 2007).
That new level of understanding has the potential to generate new pharmaco therapies to upregulate biological processes that underlie the function of the various compartments of the cardiovascular system, as new scientific explanations became available in 2006.
We have developed a methodology for discovery of concept-based original pharmacological therapy designs for combination of several drug regimens. We carry out two types of research strategy. Methodology Strategy Type One: we develop an original pharmacological therapy design specialized in addressing medical problems identified in targeted follow up studies on mortality and morbidity of cardiovascular patients. Methodology Strategy Type One is implemented in Lev-Ari & Abourjaily (2006a, 2006b, 2006c). We designed a specialized pharmaco therapy for the research results presented in NEJM, on “Circulating Endothelial Progenitor Cells and Cardiovascular Outcomes” (Werner, Kosiol, Schiegl, et al., 2005a) and the editorial interpretation of these research results by Rosenzweig (2005). We proposed the following concept-based original pharmacological therapy design for Endogenous Augmentation of circulating Endothelial Progenitor Cells for Reduction of Risk for Macrovascular Cardiac Events.
Proposal of Treatment – Three drug combination
- Inhibition of ET-1, ETA and ETA-ETB (Bosentan)
- Induction of NO production and stimulation of eNOS (Nebivolol)
- Stimulation of PPAR-gamma (substitute to Rosiglitazone)
Our Methodology Strategy Type Two involves discovery of concept-based original pharmacological therapy design for combination of several drug regimens for underlying biological processes discovered in the pursuit of basic researchers conducted in wet lab experiments by vascular biologists and molecular cardiologists. Here, we developed a concept-based original pharmacological therapy for the research results presented in Cell by Wu, Fujiwara, Cibulsky et al. (2006), Moretti, Caron, Nakano, et al. (2006) and for the research results in Nature by Smart, Risebro, Melville, et al. (2007). We propose the following concept-based original pharmacological therapy design for Preoperative and Postoperative management of cardiac injury to heart tissue, smooth muscle and to aorta and coronary artery disease. This is a treatment for Coronary Vasculogenesis, Anti-hypertention (short-acting), Vascular Anti-inflammation (vasculitis), Neovascularization of ischemic tissue and release of adult epicardium from a quiescent state and restoring its pluripotency.
Proposal for Preoperative Treatment – Three drug combination
Thymosin fraction 5 (a sublingual composition)
Indomethacin (Nonsteroidal anti-inflammatory drugs (NSAID))
Clevidipine (blood pressure lowering drug, no effect on heart rate)
Proposal for Postoperative Treatment – Three drugs combination
Thymosin fraction 5 (a sublingual composition)
ACEI (Captopril (50mg))
HCTBeta Blocker and Diuretic (Metoprolol and hydrochlorothiazide (50 mg/25 mg)) Lopressor
Thymosin beta4 Induces Adult Epicardial Progenitor Mobilization and Neovascularization
Smart et al. (2007) implicate Thymosine beta4 (Tb4) with the following functions: (a) Tb4 in regulating all three key stages of cardiac vessel development: coronary vasculogenesis, angiogenesis and arteriogenesis – collateral growth; (b) identify the adult epicardium as a potential source of vascular progenitors which, when stimulated by Tb4, migrate and differentiate into smooth muscle and endothelial cells; (c) the ability of Tb4 to promote coronary vascularization both during development and in the adult, enhances cardiomyocyte survival and contributes significantly towards Tb4-induced cardioprotection.
The reaction in the scientific community to these investigative results was most favorable.
“These results are very exciting because most humans suffering from ischemic cardiac events, either acutely or chronically, do not develop the collateral vessel growth necessary to preserve and restore heart tissue. If, in humans, we see the same effects as seen in mice, TB4 would be the first drug to prevent loss of (heart) muscle cells and restore blood flow in this manner and provide a new and much needed treatment modality for these patients,”
commented Deepak Srivastava, M.D., Professor and Director, Gladstone Institute of Cardiovascular Disease, University of California San Francisco, CA. Dr. Srivastava and his colleagues published the first paper on TB4’s effects on myocardial infarction in Nature in November 2004.
http://phx.corporate-ir.net/phoenix.zhtml?c=144396&p=irol-newsArticle&ID=932573&highlight=
VIEW VIDEO
http://www.youtube.com/watch?v=Vjj7LSuSMAo
Review of the Chemistry and the Mechanism of action supporting the process by which, N-acetyl-seryl-aspartyl-lysyl- proline (Ac-SDKP) stimulates endothelial cell differentiation from adult epicardium, is presented in
Resident-cell-based Therapy in Human Ischaemic Heart Disease: Evolution in the PROMISE of Thymosin beta4 for Cardiac Repair
http://pharmaceuticalintelligence.com/2012/04/30/93/
A Concept-based Pharmacologic Therapy of a Combined Three Drug Regimen for Regeneration and Protection of Coronary Artery Endothelium and Smooth Muscle.
This is a treatment for Coronary Vasculogenesis, Anti-hypertention (short-acting), Vascular Anti-inflammation (vasculitis), Neovascularization of ischemic tissue and release of adult epicardium from a quiescent state and restoring its pluripotency.
Preoperative Treatment – Three drugs
- Drug # 1:
- Thymosin fraction 5 (a sublingual composition)
- Drug # 2:
- Indomethacin (Nonsteroidal anti-inflammatory drugs (NSAID)) (25 mg PO bid)
- Drug # 3:
- Clevidipine (Blood pressure lowering drug, no effect on heart rate)
Postoperative Treatment – Three drugs
- Drug # 1:
- Thymosin fraction 5 (a sublingual composition)
- Drug # 4:
- ACEI (Captopril (50mg))
- Drug # 5:
- Beta Blocker and diuretic (Metoprolol and hydrochlorothiazide (50 mg/25 mg)) Lopressor HCT
Original Drug Therapy Combination Proposed
Drug # 1: Thymosin fraction 5
Drug # 2: Indomethacin
Drug # 3: Clevidipine
Thymosin beta4 is released from human blood platelets and attached by factor XIIIa (transglutaminase) to fibrin and collagen (Huff et al. 2002). They suggest that Thymosin beta4 cross-linking is mediated by factor XIIIa, a transglutaminase that is co-released from stimulated platelets. This provides a mechanism to increase the local concentration of Thymosin beta4 near sites of clots and tissue damage, where it may contribute to wound healing, angiogenesis and inflammatory responses (Al-Nedawi, et al., 2004). The beta-Thymosins constitute a family of highly conserved and extremely water-soluble 5 kDa polypeptides. Thymosin beta4 is the most abundant member; it is expressed in most cell types and is regarded as the main intracellular G-actin sequestering peptide. There is increasing evidence for extracellular functions of Thymosin beta4. For example, Thymosin beta4 increases the rate of attachment and spreading of endothelial cells on matrix components and stimulates the migration of human umbilical vein endothelial cells. They show that Thymosin beta4 can be cross-linked to proteins such as fibrin and collagen by tissue transglutaminase. Thymosin beta4 is not cross-linked to many other proteins and its cross-linking to fibrin is competed by another family member, Thymosin beta10 (Huff et al. 2002).
Rationale for selection of Sublingual compositions comprising Thymosin fraction 5
The actin binding motif of Thymosin beta4 is an essential site for its angiogenic activity (Philip, et al. (2003). Thymosin beta4 is presented in Smart, et al. (2007) in the Nature article as a single factor that can potentially couple myocardial and coronary vascular regeneration in failing mouse hearts. They have shown that cells in the heart’s outer layer can migrate deeper into a failing organ to carry out essential repairs. The migration of progenitor cells is controlled by the protein Thymosin beta 4, already known to help reduce muscle cell loss after a heart attack.
http://news.bbc.co.uk/2/hi/health/6143286.stm
The discovery opens up the possibility of using the protein to develop more effective treatments for heart disease. Previously it was thought that cells within the adult heart are in a state of permanent rest and that any progenitor cells that can contribute to heart tissue repair travel into the heart from the bone marrow. See 150 references on that perspective on cEPCs origin and roles, which was the scientific frontier on this topic, prior to the publication of Smart et al., (2007), in Lev-Ari & Abourjaily (2006a, 2006b, 2006c).
However, researchers at University College London have demonstrated that beneficial cells actually reside in the heart itself (Smart et al. (2007). This approach would bypass the risk of immune system rejection, a major problem with the use of stem cell transplants from another source. Allogenic rejection was the main reason for the selection of an endogenous augmentation method for cEPCs using drug therapy by Lev-Ari & Abourjaily (2006a, 2006b, 2006c). Closer examination revealed that without the Thymosin beta 4 protein, the progenitor cells failed to move deeper into the heart and change the cells needed to build healthy blood vessels and sustain muscle tissue.
http://www.irishhealth.com/clin/cholesterol/newsstory.php?id=10581
Nonsteroidal anti-inflammatory drugs (NSAIDs) are widely used for their anti-inflammatory effects and have been shown to have chemopreventive effects as well. NSAIDs inhibit cyclooxygenase (COX) activity to exert their anti-inflammatory effects, but it is not clear whether their antitumorigenic ability is through COX inhibition. Using subtractive hybridization, Jain et al. (2004) identified a novel member of the transforming growth factor- superfamily that has antitumorigenic activity from Indomethacin-treated HCT-116 human colorectal cancer cells. On further investigation of this library, they now report the identification of a new cDNA corresponding to the Thymosin beta-4 gene. Thymosin beta-4 is a small peptide that is known for its actin-sequestering function, and it is associated with the induction of angiogenesis, accelerated wound healing, and metastatic potential of tumor cells. However, only selective NSAIDs induce Thymosin beta-4 expression in a time- and concentration-dependent manner. For example,
Indomethacin and SC-560 [5-(4-chlorophenyl)-1-(4-methoxyphenyl)-3-(trifluoromethyl)-1H-pyrazole] induce Thymosin beta-4 expression whereas sulindac sulfide does not.
They show that selective NSAIDs induce actin cytoskeletal reorganization, a precursory step to many dynamic processes regulating growth and motility including tumorigenesis. This is the first report to link Thymosin beta-4 induction with NSAIDs. These data suggest that NSAIDs alter the expression of a diverse number of genes and provide new insights into the chemopreventive and biological activity of these drugs (Jain et al. 2004).
Rationale for Indomethacin selection
Inhibitor of prostaglandin synthesis. Inhibits cyclooxygenase (COX) 1 selective.
Suggested dosage: 25 mg PO bid.
Jain et al. (2004) report a link between Thymosin beta-4 induction with NSAIDs. We selected both drugs (drug classes) and anticipate strong synergistic therapeutic effects.
Clevidipine is the first third-generation calcium channel blocker, Dr. Papadakos said. It has what he called an “ultrashort” clinically relevant half-life of about one minute and then is rapidly metabolized. The effect on blood pressure is seen within one to two minutes.
http://www.medpagetoday.com/MeetingCoverage/SCCM/tb/5091
Clevidipine is an investigational agent undergoing late-stage clinical development to evaluate its potential as an innovative, targeted, fast acting intravenous product under investigation for lowering blood pressure before, during and after surgery.
http://www.themedicinescompany.com/products_Clevidipine.shtml
The Medicines Company entered into agreements with AstraZeneca PLC in March of 2002 for the development, licensing and commercialization of Clevidipine. If approved, the product could be an excellent fit with The Medicines Company’s emerging acute cardiovascular care franchise, which is led by Angiomax® (bivalirudin), an anticoagulant approved in the U.S. and other countries for use during coronary angioplasty procedures. If Clevidipine passes further clinical hurdles — phase III trials are under way — the drug may form a useful addition to the medications available to physicians in the perioperative setting
Mechanism of Action
Clevidipine belongs to a well-known class of drugs called dihydropyridine calcium channel antagonists. In vitro studies demonstrated that Clevidipine acts by selectively relaxing the smooth muscle cells that line small arteries, resulting in widening of the artery opening and reducing blood pressure within the artery (Levy, Huraux, Nordlander, 1997, 345-358).
Phase III Clinical Trials
The Medicines Company is currently sponsoring a Phase III clinical program of five studies to evaluate safety and efficacy of Clevidipine:
Early Development
The Medicines Company’s development program for Clevidipine follows upon the data sets generated by AstraZeneca, which completed clinical pharmacology, dose-finding and efficacy studies in almost 300 patients or volunteers. In clinical studies, Clevidipine has shown to provide the desired blood pressure lowering effect without causing an increase in heart rate (Kotrly, et al. 1984). Further studies demonstrate that reductions in blood pressure are dose-dependent, are not associated with an increase in heart rate and cease rapidly after stopping Clevidipine infusions (Ericsson, et al., 2000), (Schwieler, et al., 1999). In clinical studies Clevidipine was rapidly metabolized independent of the liver and the kidneys, allowing rapid clearance of the drug from the bloodstream (Ericsson, et al., 1999a), (Ericsson, et al., 1999b). Therefore, the effects of Clevidipine are short-lived, which translates into a rapid cessation of its effect on reducing blood pressure.
The two efficacy studies are known as ESCAPE-1 and ESCAPE-2. The primary objective of these studies is to determine the efficacy of Clevidipine injection versus placebo in treating pre-operative (ESCAPE-1) and post-operative (ESCAPE-2) high blood pressure. Three safety studies are collectively known as ECLIPSE. The primary objective is to establish the safety of Clevidipine in the treatment of perioperative high blood pressure, as measured by a comparison of the incidences of death, stroke, myocardial infarction and renal dysfunction between the Clevidipine and comparative treatment groups. The comparative treatments are nitroglycerin, sodium nitroprusside and nicardipine. The ECLIPSE trial randomized 589 patients at 40 centers in the U.S. to get either sodium nitroprusside or Clevidipine. Sodium nitroprusside was administered according to institutional practice; Clevidipine was begun at 2 mg/kg and doubled every 90 seconds until blood pressure was lowered. The primary endpoint was the difference in major clinical events — death, myocardial infarction, stroke, and renal dysfunction 30 days after surgery. The secondary endpoint was blood pressure control during the first 24 hours after surgery.
The study showed no significant differences in the elements of the primary endpoint, except for mortality, Dr. Papadakos said, where 1.7% of Clevidipine patients died, compared with 4.7 of those getting sodium nitroprusside. The difference was statistically significant at P<0.05, but Dr. Papadakos characterized the improvement as “slight.” On the other hand, the drug did show an important difference in blood pressure control over the first 24 hours, he said:
- Patients on Clevidipine spent an average of 4.37 minutes per hour outside the desired blood pressure range.
- Sodium nitroprusside patients spent, on average, 10.5 minutes per hour outside the desired range.
- The difference was statistically significant at P<0.003.
Dr. Papadakos concluded that Clevidipine is a new drug that is effective, safe, and easy to use. On 2/20/2007, Dr. Deutschman, who moderated the late-breaking session at which Dr. Papadakos spoke, said that a better comparison, would be intravenous nicardipine (Cardene IV), a second-generation calcium channel blocker that is also in wide use and is considered the standard of care. “We don’t know yet if this drug is going to be better than nicardipine,” he said.
http://www.medpagetoday.com/MeetingCoverage/SCCM/tb/5091
Rationale for Clevidipine selection
Clevidipine is an acute care product. Blood pressure management is a major component of care during the 13.4 million inpatient surgeries conducted in the U.S. each year. Blood pressure control, which is managed by an anesthesiologist, is often important in patients with both normal and high blood pressure undergoing surgery or other interventional procedures. Some of these patients require rapid, precise control of blood pressure to avoid compromising key organ function such as the heart, brain and kidney.
CONCLUSION
This is the first study to design a novel combination drug treatment for Coronary Vasculogenesis, Anti-hypertention (short-acting), Vascular Anti-inflammation (vasculitis), Neovascularization of ischemic tissue and release of adult epicardium from a quiescent state and restoring its pluripotency. This treatment is based on the new three paradigms that were presented in Cell (2006) and Nature (2007). This combination drug therapy of three drugs, one in current use (Indomethacin), and two in clinical trials (Thymosin beta4 & Clevidipine), has not been proposed before. It represents an original concept drug combination design by Lev-Ari & Abourjaily (2007). This combination represents the cutting edge conceptualization of the field of treatment of cardiac injury based on a protein produced in the heart cells, Thymosin beta4, which function as a tissue and artery healer. Its upregulation by drug therapy will revolutionize cardiology and treatment for cardiovascular disease. The combination drug therapy consists of the following drugs:
Thymosin fraction 5 (a sublingual composition)
Indomethacin (Nonsteroidal anti-inflammatory drugs (NSAID)) (25 mg PO bid)
Clevidipine (Blood pressure lowering drug, (no effect on heart rate))
REFERENCES
Al-Nedawi, K.N.I., Malgorzata, C., Bednarek, R., Szemraj, J., Swiatkowska, M., Cierniewska-Cieslak, A., Wyczolkowska, J., Cierniewski, C.S. (2004, February) “Thymosin 4 induces the synthesis of plasminogen activator inhibitor 1 in cultured endothelial cells and increases its extracellular expression.” Blood, 103, (4), 1319-1324.
Ericsson, H., Fakt. C., Hoglund. L., et al. (1999a). “Pharmacokinetics and pharmacodynamics of Clevidipine in healthy volunteers after intravenous infusion.” Eur J Clin Pharm., 55 (1), 61-67.
Ericsson, H., Fakt, C., Jolin-Mellgard A., et al. (1999b). “Clinical and pharmacokinetic results with a new ultrashort-acting calcium antagonist, Clevidipine, following gradually increasing intravenous doses to healthy volunteers.” Br J Clin Pharm., 47 (5), 531-538.
Ericsson, H., Bredberg, U., Eriksson, U., et al. (2000). “Pharmacokinetics and arteriovenous differences in Clevidipine concentration following a short and a long-term intravenous infusion in healthy volunteers.” Anesthesiology, 92 (4), 993-1001.
Fleming, I. (2006). “Signaling by the Angiotensin-Converting Enzyme” Circulation Research, 98, 887.
Heart can carry out own repairs, 16.11.2006
Retrieved on 3/1/2007
http://www.medicalprogress.org/benefits/heartdis/news.cfm?news_id=478
Retrieved on 3/1/2007
http://news.bbc.co.uk/2/hi/health/6143286.stm
Heart may be able to repair itself
Retrieved on 3/1/2007
http://www.irishhealth.com/clin/cholesterol/newsstory.php?id=10581
Huff, T., Otto, A., Muller, C.S.G., Meier, M., Hannappel, E. (2002). “Thymosin ß4 is released from human blood platelets and attached by factor XIIIa (transglutaminase) to fibrin and collagen.”The FASEB Journal, 16, 691-696.
Jain, A.K., Moore, S.M., Yamaguchi, K., Eling, T.E., Baek, S.J. (2004, August). “Selective Nonsteroidal Anti-Inflammatory Drugs Induce Thymosin beta-4 and Alter Actin Cytoskeletal Organization in Human Colorectal Cancer Cells.” Journal of Pharmacology and Experimental Therapeutics, 311 (3) 885-891.
Kotrly, K. J., Ebert, T. J., Vucins, E. et al. (1984). “Baroreceptor reflex control of heart rate during isoflurane anesthesia in humans.” Anesthesiology, 60, 173-179.
Lev-Ari, A. & Abourjaily, P. (2006a) “An Investigation of the Potential of circulating Endothelial Progenitor Cells (cEPC) as a Therapeutic Target for Pharmacologic Therapy Design for Cardiovascular Risk Reduction.” Part I: Macrovascular Disease – Therapeutic Potential of cEPCs – Reduction methods for CV risk. Unpublished manuscript.
Lev-Ari, A. & Abourjaily, P. (2006b) “An Investigation of the Potential of circulating Endothelial Progenitor Cells (cEPC) as a Therapeutic Target for Pharmacologic Therapy Design for Cardiovascular Risk Reduction.” Part II: Therapeutic Strategy for cEPCs Endogenous Augmentation: A Concept-based Treatment Protocol for a Combined Three Drug Regimen. Unpublished manuscript.
Lev-Ari, A. & Abourjaily, P. (2006c) “An Investigation of the Potential of circulating Endothelial Progenitor Cells (cEPC) as a Therapeutic Target for Pharmacological Therapy Design for Cardiovascular Risk Reduction.” Part III: Biomarker for Therapeutic Targets of Cardiovascular Risk Reduction by cEPCs Endogenous Augmentation using New Combination Drug Therapy of Three Drug Classes and Several Drug Indications. A Theoretical Design for Quantification of the Endogenous EPCs Augmentation for Differential Level of CV Risk Reduction and Diagnostic Device Design for Drug Delivery. Unpublished manuscript.
Lev-Ari, A. & Abourjaily, P. (2007). Heart Vasculature – Regeneration and Protection of Coronary Artery Endothelium and Smooth Muscle: A Concept-based Pharmacological Therapy of a Combined Three Drug Regimen. Unpublished manuscript.
Levy, J. H., Huraux, C., Nordlander, M. (1997). “Treatment of perioperative hypertension.” In: Epstein M, Ed. Chapter in Calcium Antagonists in Clinical Medicine. Philadelphiea: Hanely & Belfus, pp. 345-358.
Liu, J-M, Lawrence, F., Kovacevic, M., Bignon, J., Papadimitriou, E., Lallemand, J-Y., Katsoris, P., Potier, P., Fromes, Y., Wdzieczak-Bakala, J. (2003, April) “The tetrapeptide AcSDKP, an inhibitor of primitive hematopoietic cell proliferation, induces angiogenesis in vitro and in vivo.” Blood, 101 (8), 3014-3020
Moretti, A., Caron, L., Nakano, A., Lam, J.T., Bernshausen, A., Chen, Y., Qyang, Y., Bu, L., Sasaki, M., Martin-Puig, S., Sun, Y., Evans, S.M., Laugwitz, K-L, Chien, K.R. (2006, December) “Multipotent Embryonic Isl1+ Progenitor Cells Lead to Cardiac, Smooth Muscle, and Endothelial Cell Diversification.” Cell, 127, 1151-1165.
Protein Discovered That Can Tell Human Heart to Heal Itself
Retrieved 3/1/2007
http://www.cathlabdigest.com/displaynews.cfm?newsid=1122065
Philp D, Huff T, Gho YS, Hannappel E, Kleinman HK. (2003). “The actin binding site on Thymosin beta4 promotes angiogenesis.” FASEB Journal, published on line 9/18/2003.
Retrieved 3/1/2007
http://www.fasebj.org/cgi/reprint/03-0121fjev1.pdf
Putting the art in heart research, 15 February 2007
Retrieved on 3/1/2007
http://www.ich.ucl.ac.uk/pressoffice/pressrelease_00498
Rosenzweig A., (2005). Circulating Endothelial Progenitors – Cells as Biomarkers. NEJM, 353 (10), 1055-1057.
Schwieler, J.H., Ericsson, H., Lofdahl, P., et al. (1999). “Circulatory effects and pharmacology of Clevidipine, a novel ultra short acting and vascular selective calcium antagonist, in hypertensive humans.” J Cardiovasc Pharmacology, 34 (2), 268-274.
Smart, N., Risebro, C.A., Melville, A.D., Moses, K., Schwartz, R.J., Chien, K.R., Riley, P.R. (2007, January) “Thymosin Beta4 induces adult epicardial progenitor mobilization and neovascularization.” Nature, 445, 177-182.
Sublingual compositions comprising Thymosin fraction 5 and methods for administration
Retrieved 3/1/2007
http://www.pharmcast.com/Patents100/Yr2004/May2004/051104/6733791_Sublingual051104.htm
TMSB4X Thymosin, beta 4, X-linked
Retrieved on 3/1/2007
http://www.ihop-net.org/UniPub/iHOP/gs/92756.html
Waeckel, L., Jérôme Bignon, J., Jian-Miao Liu, J-M., Markovits, D., Ebrahimian, T.G., Vilar, J., Mees, B., Blanc-Brude, O., Barateau, V., Sophie Le ricousse-Roussanne. S., Duriez, M. Tobelem, G., Wdzieczak-Bakala, J., Bernard I Lévy, B.I., Silvestre, J-S. (2006) “Tetrapeptide AcSDKP Induces Postischemic Neovascularization Through Monocyte Chemoattractant Protein-1 Signaling.” Arteriosclerosis, Thrombosis, and Vascular Biology, 26, 773
Wang, D., Oscar A. Carretero, O.A.,Yang, X-Y., Rhaleb, N-E., Liu, Y-H., Liao, T-D., Yang, X-P. (2004). “N-acetyl-seryl-aspartyl-lysyl-proline stimulates angiogenesis in vitro and in vivo.” Am J Physiol Heart Circ Physiol., 287, H2099-H2105.
Werner N, Junk S, Laufs L, Link A, Walenta K, Bohm M, Nickenig G., (2003). Intravenous transfusion of endothelial progenitor cells reduces neointima formation after vascular injury. Circ Res., 93, e17– e24.
Werner N, Kosiol S, Schiegl T, Ahlers P, Walenta K, Link A, Böhm M, Nickenig G. (2005a). Circulating Endothelial Progenitor Cells and Cardiovascular Outcomes, NEJM, 353, 999-1007
Werner, N. & Nickenig, G. (2005b). Authors Reply to Correspondence to the Editor on Circulating Endothelial Progenitor Cells. NEJM, 353 (24), 2613-2616
Wu, S.M., Fujiwara, Y., Cibulsky, S.M., Clapham, D.E., Lien, C., Schultheiss, T.M., Orkin, S.H. (2006, December). “Developmental Origin of a Bipotential Myocardial and Smooth Muscle Cell Precursor in the Mammalian Heart.” Cell, 127, 1137-1150.
Other related articles on this Open Access Online Scientific Journal, include the following:
Saha, S. (2012b) Innovations in Bio instrumentation for Measurement of Circulating Progenetor Endothelial Cells in Human Blood.
http://pharmaceuticalintelligence.com/2012/07/08/innovations-in-bio-instrumentation-for-measurement-of-circulating-progenitor-endothelial-cells-in-human-blood/
Saha, S. (2012c) Endothelial Differentiation and Morphogenesis of Cardiac Precursor
http://pharmaceuticalintelligence.com/2012/07/17/endothelial-differentiation-and-morphogenesis-of-cardiac-precursors/
Saha, S. (2012e). Human Embryonic-Derived Cardiac Progenitor Cells for Myocardial Repair
http://pharmaceuticalintelligence.com/2012/08/01/human-embryonic-derived-cardiac-progenitor-cells-for-myocardial-repair/
Lev-Ari, A. 12/29/2012. Coronary artery disease in symptomatic patients referred for coronary angiography: Predicted by Serum Protein Profiles
http://pharmaceuticalintelligence.com/2012/12/29/coronary-artery-disease-in-symptomatic-patients-referred-for-coronary-angiography-predicted-by-serum-protein-profiles/
Bernstein, HL and Lev-Ari, A. 11/28/2012. Special Considerations in Blood Lipoproteins, Viscosity, Assessment and Treatment
http://pharmaceuticalintelligence.com/2012/11/28/special-considerations-in-blood-lipoproteins-viscosity-assessment-and-treatment/
Lev-Ari, A. 11/13/2012 Peroxisome proliferator-activated receptor (PPAR-gamma) Receptors Activation: PPARγ transrepression for Angiogenesis in Cardiovascular Disease and PPARγ transactivation for Treatment of Diabetes
http://pharmaceuticalintelligence.com/2012/11/13/peroxisome-proliferator-activated-receptor-ppar-gamma-receptors-activation-pparγ-transrepression-for-angiogenesis-in-cardiovascular-disease-and-pparγ-transactivation-for-treatment-of-dia/
Lev-Ari, A. 10/19/2012 Clinical Trials Results for Endothelin System: Pathophysiological role in Chronic Heart Failure, Acute Coronary Syndromes and MI – Marker of Disease Severity or Genetic Determination?
http://pharmaceuticalintelligence.com/2012/10/19/clinical-trials-results-for-endothelin-system-pathophysiological-role-in-chronic-heart-failure-acute-coronary-syndromes-and-mi-marker-of-disease-severity-or-genetic-determination/
Lev-Ari, A. 10/4/2012 Endothelin Receptors in Cardiovascular Diseases: The Role of eNOS Stimulation
http://pharmaceuticalintelligence.com/2012/10/04/endothelin-receptors-in-cardiovascular-diseases-the-role-of-enos-stimulation/
Lev-Ari, A. 10/4/2012 Inhibition of ET-1, ETA and ETA-ETB, Induction of NO production, stimulation of eNOS and Treatment Regime with PPAR-gamma agonists (TZD): cEPCs Endogenous Augmentation for Cardiovascular Risk Reduction – A Bibliography
http://pharmaceuticalintelligence.com/2012/10/04/inhibition-of-et-1-eta-and-eta-etb-induction-of-no-production-and-stimulation-of-enos-and-treatment-regime-with-ppar-gamma-agonists-tzd-cepcs-endogenous-augmentation-for-cardiovascular-risk-reduc/
Lev-Ari, A. 8/28/2012 Cardiovascular Outcomes: Function of circulating Endothelial Progenitor Cells (cEPCs): Exploring Pharmaco-therapy targeted at Endogenous Augmentation of cEPCs
http://pharmaceuticalintelligence.com/2012/08/28/cardiovascular-outcomes-function-of-circulating-endothelial-progenitor-cells-cepcs-exploring-pharmaco-therapy-targeted-at-endogenous-augmentation-of-cepcs/
Lev-Ari, A. 8/27/2012 Endothelial Dysfunction, Diminished Availability of cEPCs, Increasing CVD Risk for Macrovascular Disease – Therapeutic Potential of cEPCs
http://pharmaceuticalintelligence.com/2012/08/27/endothelial-dysfunction-diminished-availability-of-cepcs-increasing-cvd-risk-for-macrovascular-disease-therapeutic-potential-of-cepcs/
Lev-Ari, A. 8/24/2012 Vascular Medicine and Biology: CLASSIFICATION OF FAST ACTING THERAPY FOR PATIENTS AT HIGH RISK FOR MACROVASCULAR EVENTS Macrovascular Disease – Therapeutic Potential of cEPCs
http://pharmaceuticalintelligence.com/2012/08/24/vascular-medicine-and-biology-classification-of-fast-acting-therapy-for-patients-at-high-risk-for-macrovascular-events-macrovascular-disease-therapeutic-potential-of-cepcs/
Lev-Ari, A. 7/30/2012 Biosimilars: Intellectual Property Creation and Protection by Pioneer and by Biosimilar Manufacturers
http://pharmaceuticalintelligence.com/2012/07/30/biosimilars-intellectual-property-creation-and-protection-by-pioneer-and-by-biosimilar-manufacturers/
Lev-Ari, A. 7/29/2012 Biosimilars: Financials 2012 vs. 2008
http://pharmaceuticalintelligence.com/2012/07/30/biosimilars-financials-2012-vs-2008/
Lev-Ari, A. 7/29/2012 Biosimilars: CMC Issues and Regulatory Requirements
http://pharmaceuticalintelligence.com/2012/07/29/biosimilars-cmc-issues-and-regulatory-requirements/
Lev-Ari, A. 7/19/2012 Cardiovascular Disease (CVD) and the Role of agent alternatives in endothelial Nitric Oxide Synthase (eNOS) Activation and Nitric Oxide Production
http://pharmaceuticalintelligence.com/2012/07/19/cardiovascular-disease-cvd-and-the-role-of-agent-alternatives-in-endothelial-nitric-oxide-synthase-enos-activation-and-nitric-oxide-production/
Lev-Ari, A. 4/30/2012 Resident-cell-based Therapy in Human Ischaemic Heart Disease: Evolution in the PROMISE of Thymosin beta4 for Cardiac Repair
http://pharmaceuticalintelligence.com/2012/04/30/93/
Lev-Ari, A. 5/29/2012 Triple Antihypertensive Combination Therapy Significantly Lowers Blood Pressure in Hard-to-Treat Patients with Hypertension and Diabetes
http://pharmaceuticalintelligence.com/2012/05/29/445/
Lev-Ari, A. 7/2/2012 Macrovascular Disease – Therapeutic Potential of cEPCs: Reduction Methods for CV Risk
http://pharmaceuticalintelligence.com/2012/07/02/macrovascular-disease-therapeutic-potential-of-cepcs-reduction-methods-for-cv-risk/
Lev-Ari, A. 7/9/2012 Mitochondria Dysfunction and Cardiovascular Disease – Mitochondria: More than just the “powerhouse of the cell”
http://pharmaceuticalintelligence.com/2012/07/09/mitochondria-more-than-just-the-powerhouse-of-the-cell/
Lev-Ari, A. 7/16/2012 Bystolic’s generic Nebivolol – positive effect on circulating Endothelial Proginetor Cells endogenous augmentation
http://pharmaceuticalintelligence.com/2012/07/16/bystolics-generic-nebivolol-positive-effect-on-circulating-endothilial-progrnetor-cells-endogenous-augmentation/
Like this:
Like Loading...
Read Full Post »




![Steps for the development of imaging biomarkers (adapted from [5])](https://i0.wp.com/pharmaceuticalintelligence.com/wp-content/uploads/2013/03/fig-1.gif?resize=500%2C363)