What is the Future for Genomics in Clinical Medicine?
Author and Curator: Larry H Bernstein, MD, FCAP

WordCloud Image Produced by Adam Tubman
Introduction
This is the last in a series of articles looking at the past and future of the genome revolution. It is a revolution indeed that has had a beginning with the first phase discovery leading to the Watson-Crick model, the second phase leading to the completion of the Human Genome Project, a third phase in elaboration of ENCODE. But we are entering a fourth phase, not so designated, except that it leads to designing a path to the patient clinical experience.
What is most remarkable on this journey, which has little to show in treatment results at this time, is that the boundary between metabolism and genomics is breaking down. The reality is that we are a magnificent “magical” experience in evolutionary time, functioning in a bioenvironment, put rogether like a truly complex machine, and with interacting parts. What are those parts – organelles, a genetic message that may be constrained and it may be modified based on chemical structure, feedback, crosstalk, and signaling pathways. This brings in diet as a source of essential nutrients, exercise as a method for delay of structural loss (not in excess), stress oxidation, repair mechanisms, and an entirely unexpected impact of this knowledge on pharmacotherapy. I illustrate this with some very new observations.
Gutenberg Redone
The first is a recent talk on how genomic medicine has constructed a novel version of the “printing press”, that led us out of the dark ages.
In our series The Creative Destruction of Medicine, I’m trying to get into critical aspects of how we can Schumpeter or reboot the future of healthcare by leveraging the big innovations that are occurring in the digital world, including digital medicine.
For example, there was a drug trial for melanoma and the mutation of BRAF, which is the gene that is found in about 60% of people with malignant melanoma. When that trial was done, there was a placebo control, and there was a big ethical charge asking whether it is justifiable to have a body count. This was a matched drug for the biology underpinning metastatic melanoma, which is essentially a fatal condition within 1 year, and researchers were giving some individuals a placebo.
I shall however, use some new information that gives real cause for joy.
Reprogramming Alters Cells’ Fate
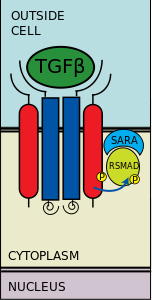
Transforming growth factor beta (TGF-β) is a secreted protein that controls proliferation, cellular differentiation, and other functions in most cells. http://en.wikipedia.org/wiki/TGFbeta (Photo credit: Wikipedia)
Pioneer Transcription Factors
A Nontranscriptional Role for HIF-1α as a Direct Inhibitor of DNA Replication
ME Hubbi, K Shitiz, DM Gilkes, S Rey,….GL Semenza. Johns Hopkins University SOM
Sci. Signal 2013; 6(262) 10pgs. [DOI: 10.1126/scisignal.2003417] http:dx.doi.org/10.1126/scisignal.2003417
http://SciSignal.com/A Nontranscriptional Role for HIF-1α as a Direct Inhibitor of DNA Replication/
Many of the cellular responses to reduced O2 availability are mediated through the transcriptional activity of hypoxia-inducible factor 1 (HIF-1). We report a role for the isolated HIF-1α subunit as an inhibitor of DNA replication, and this role was independent of HIF-1β and transcriptional regulation. In response to hypoxia, HIF-1α bound to Cdc6, a protein that is essential for loading of the mini-chromosome maintenance (MCM) complex (which has DNA helicase activity) onto DNA, and promoted the interaction between Cdc6 and the MCM complex. The binding of HIF-1α to the complex decreased phosphorylation and activation of the MCM complex by the kinase Cdc7. As a result, HIF-1α inhibited firing of replication origins, decreased DNA replication, and induced cell cycle arrest in various cell types. To whom correspondence should be addressed. E-mail: gsemenza@jhmi.edu
Citation: M. E. Hubbi, Kshitiz, D. M. Gilkes, S. Rey, C. C. Wong, W. Luo, D.-H. Kim, C. V. Dang, A. Levchenko, G. L. Semenza, A Nontranscriptional Role for HIF-1α as a Direct Inhibitor of DNA Replication. Sci. Signal. 6, ra10 (2013).
Identification of a Candidate Therapeutic Autophagy-inducing Peptide
Nature 2013;494(7436). http://nature.com/Identification_of_a_candidate_therapeutic_autophagy-inducing_peptide/ http://www.ncbi.nlm.nih.gov/pubmed/23364696
http://www.readcube.com/articles/10.1038/nature11866
Beth Levine and colleagues have constructed a cell-permeable peptide derived from part of an autophagy protein called beclin 1. This peptide is a potent inducer of autophagy in mammalian cells and in vivo in mice and was effective in the clearance of several viruses including chikungunya virus, West Nile virus and HIV-1.
Could this small autophagy-inducing peptide may be effective in the prevention and treatment of human diseases?
PR-Set7 Is a Nucleosome-Specific Methyltransferase that Modifies Lysine 20 of
Histone H4 and Is Associated with Silent Chromatin
K Nishioka, JC Rice, K Sarma, H Erdjument-Bromage, …, D Reinberg. Molecular Cell, Vol. 9, 1201–1213, June, 2002, Copyright 2002 by Cell Press http://www.cell.com/molecular-cell/abstract/S1097-2765(02)00548-8
http://www.sciencedirect.com/science/article/pii/S1097276502005488 http://www.ncbi.nlm.nih.gov/pubmed/12086618
http://www.cienciavida.cl/publications/b46e8d324fa4aefa771c4d6ece4d2e27_PR-Set7_Is_a_Nucleosome-Specific.pdf
We have purified a human histone H4 lysine 20methyl-transferase and cloned the encoding gene, PR/SET07. A mutation in Drosophila pr-set7 is lethal: second in-star larval death coincides with the loss of H4 lysine 20 methylation, indicating a fundamental role for PR-Set7 in development. Transcriptionally competent regions lack H4 lysine 20 methylation, but the modification coincided with condensed chromosomal regions polytene chromosomes, including chromocenter euchromatic arms. The Drosophila male X chromosome, which is hyperacetylated at H4 lysine 16, has significantly decreased levels of lysine 20 methylation compared to that of females. In vitro, methylation of lysine 20 and acetylation of lysine 16 on the H4 tail are competitive. Taken together, these results support the hypothesis that methylation of H4 lysine 20 maintains silent chromatin, in part, by precluding neighboring acetylation on the H4 tail.
Next-Generation Sequencing vs. Microarrays
Shawn C. Baker, Ph.D., CSO of BlueSEQ
GEN Feb 2013
With recent advancements and a radical decline in sequencing costs, the popularity of next generation sequencing (NGS) has skyrocketed. As costs become less prohibitive and methods become simpler and more widespread, researchers are choosing NGS over microarrays for more of their genomic applications. The immense number of journal articles citing NGS technologies it looks like NGS is no longer just for the early adopters. Once thought of as cost prohibitive and technically out of reach, NGS has become a mainstream option for many laboratories, allowing researchers to generate more complete and scientifically accurate data than previously possible with microarrays.
Gene Expression
Researchers have been eager to use NGS for gene expression experiments for a detailed look at the transcriptome. Arrays suffer from fundamental ‘design bias’ —they only return results from those regions for which probes have been designed. The various RNA-Seq methods cover all aspects of the transcriptome without any a priori knowledge of it, allowing for the analysis of such things as novel transcripts, splice junctions and noncoding RNAs. Despite NGS advancements, expression arrays are still cheaper and easier when processing large numbers of samples (e.g., hundreds to thousands).
Methylation
While NGS unquestionably provides a more complete picture of the methylome, whole genome methods are still quite expensive. To reduce costs and increase throughput, some researchers are using targeted methods, which only look at a portion of the methylome. Because details of exactly how methylation impacts the genome and transcriptome are still being investigated, many researchers find a combination of NGS for discovery and microarrays for rapid profiling.
Diagnostics
They are interested in ease of use, consistent results, and regulatory approval, which microarrays offer. With NGS, there’s always the possibility of revealing something new and unexpected. Clinicians aren’t prepared for the extra information NGS offers. But the power and potential cost savings of NGS-based diagnostics is alluring, leading to their cautious adoption for certain tests such as non-invasive prenatal testing.
Cytogenetics
Perhaps the application that has made the least progress in transitioning to NGS is cytogenetics. Researchers and clinicians, who are used to using older technologies such as karyotyping, are just now starting to embrace microarrays. NGS has the potential to offer even higher resolution and a more comprehensive view of the genome, but it currently comes at a substantially higher price due to the greater sequencing depth. While dropping prices and maturing technology are causing NGS to make headway in becoming the technology of choice for a wide range of applications, the transition away from microarrays is a long and varied one. Different applications have different requirements, so researchers need to carefully weigh their options when making the choice to switch to a new technology or platform. Regardless of which technology they choose, genomic researchers have never had more options.
Sequencing Hones In on Targets
Greg Crowther, Ph.D.
GEN Feb 2013
Cliff Han, PhD, team leader at the Joint Genome Institute in the Los Alamo National Lab, was one of a number of scientists who made presentations regarding target enrichment at the “Sequencing, Finishing, and Analysis in the Future” (SFAF) conference in Santa Fe, which was co-sponsored by the Los Alamos National Laboratory and DOE Joint Genome Institute. One of the main challenges is that of target enrichment: the selective sequencing of genomic or transcriptomic regions. The polymerase chain reaction (PCR) can be considered the original target-enrichment technique and continues to be useful in contexts such as genome finishing. “One target set is the unique gaps—the gaps in the unique sequence regions. Another is to enrich the repetitive sequences…ribosomal RNA regions, which together are about 5 kb or 6 kb.” The unique-sequence gaps targeted for PCR with 40-nucleotide primers complementary to sequences adjacent to the gaps, did not yield the several-hundred-fold enrichment expected based on previously published work. “We got a maximum of 70-fold enrichment and generally in the dozens of fold of enrichment,” noted Dr. Han.
“We enrich the genome, put the enriched fragments onto the Pacific Biosciences sequencer, and sequence the repeats,” continued Dr. Han. “In many parts of the sequence there will be a unique sequence anchored at one or both ends of it, and that will help us to link these scaffolds together.” This work, while promising, will remain unpublished for now, as the Joint Genome Institute has shifted its resources to other projects.
At the SFAF conference Dr. Jones focused on going beyond basic target enrichment and described new tools for more efficient NGS research. “Hybridization methods are flexible and have multiple stop-start sites, and you can capture very large sizes, but they require library prep,” said Jennifer Carter Jones, Ph.D., a genomics field applications scientist at Agilent. “With PCR-based methods, you have to design PCR primers and you’re doing multiplexed PCR, so it’s limited in the size that you can target. But the workflow is quick because there’s no library preparation; you’re just doing PCR.” She discussed Agilent’s recently acquired HaloPlex technology, a hybrid system that includes both a hybridization step and a PCR step. Because no library preparation is required, sequencing results can be obtained in about six hours, making it suitable for clinical uses. However, the hybridization step allows capture of targets of up to 5 megabases—longer than purely PCR-based methods can deliver. The Agilent talk also provided details on the applications of SureSelect, the company’s hybridization technology, to Methyl-Seq and RNA-Seq research. With this technology, 120-mer baits hybridize to targets, then are pulled down with streptavidin-coated magnetic beads.
These are selections from the SFAF conference, which is expected to be a boost to work on the microbiome, and lead to infectious disease therapeutic approaches.
Summary
We have finished a breathtaking ride through the genomic universe in several sessions. This has been a thorough review of genomic structure and function in cellular regulation. The items that have been discussed and can be studied in detail include:
- the classical model of the DNA structure
- the role of ubiquitinylation in managing cellular function and in autophagy, mitophagy, macrophagy, and protein degradation
- the nature of the tight folding of the chromatin in the nucleus
- intramolecular bonds and short distance hydrophobic and hydrophilic interactions
- trace metals in molecular structure
- nuclear to membrane interactions
- the importance of the Human Genome Project followed by Encode
- the Fractal nature of chromosome structure
- the oligomeric formation of short sequences and single nucletide polymorphisms (SNPs)and the potential to identify drug targets
- Enzymatic components of gene regulation (ligase, kinases, phosphatases)
- Methods of computational analysis in genomics
- Methods of sequencing that have become more accurate and are dropping in cost
- Chromatin remodeling
- Triplex and quadruplex models not possible to construct at the time of Watson-Crick
- sequencing errors
- propagation of errors
- oxidative stress and its expected and unintended effects
- origins of cardiovascular disease
- starvation and effect on protein loss
- ribosomal damage and repair
- mitochondrial damage and repair
- miscoding and mutational changes
- personalized medicine
- Genomics to the clinics
- Pharmacotherapy horizons
- driver mutations
- induced pluripotential embryonic stem cell (iPSCs)
- The association of key targets with disease
- The real possibility of moving genomic information to the bedside
- Requirements for the next generation of electronic health record to enable item 29
Related articles
- Retrovirus in the human genome is active in pluripotent stem cells (sciencedaily.com)
- Learning from the linker: New study sheds light on cellular reprogramming (phys.org)
- A Quantitative System for Discriminating Induced Pluripotent Stem Cells, Embryonic Stem Cells and Somatic Cells (plosone.org)
- Stem Cell Therapies May Improve Following Finding That Retrovirus In The Human Genome Is Active In Pluripotent Stem Cells (medicalnewstoday.com)
- Leprosy spreads by reprogramming nerve cells into migratory stem cells (guardian.co.uk)
- Harvard Researchers: Embryonic Cells Can Cause Cancer (salem-news.com)
Other Related articles on this Open Access Online Scientific Journal, include the following:
http://pharmaceuticalintelligence.com/2012/10/22/blood-vessel-generating-stem-cells-discovered/ RSaxena
http://pharmaceuticalintelligence.com/2012/12/09/naotech-therapy-for-breast-cancer/ TBarliya


