Hypoxia Inducible Factor 1 (HIF-1)
Writer and Curator: Larry H Bernstein, MD, FCAP
7.9 Hypoxia Inducible Factor 1 (HIF-1)
7.9.1 Hypoxia and mitochondrial oxidative metabolism
7.9.2 Hypoxia promotes isocitrate dehydrogenase-dependent carboxylation of α-ketoglutarate to citrate to support cell growth and viability
7.9.3 Hypoxia-Inducible Factors in Physiology and Medicine
7.9.4 Hypoxia-inducible factor 1. Regulator of mitochondrial metabolism and mediator of ischemic preconditioning
7.9.5 Regulation of cancer cell metabolism by hypoxia-inducible factor 1
7.9.6 Coming up for air. HIF-1 and mitochondrial oxygen consumption
7.9.7 HIF-1 mediates adaptation to hypoxia by actively downregulating mitochondrial oxygen consumption
7.9.8 HIF-1. upstream and downstream of cancer metabolism
7.9.9 In Vivo HIF-Mediated Reductive Carboxylation
7.9.10 Evaluation of HIF-1 inhibitors as anticancer agents
7.9.1 Hypoxia and mitochondrial oxidative metabolism
Solaini G1, Baracca A, Lenaz G, Sgarbi G.
Biochim Biophys Acta. 2010 Jun-Jul; 1797(6-7):1171-7
http://dx.doi.org/10.1016/j.bbabio.2010.02.011
It is now clear that mitochondrial defects are associated with a large variety of clinical phenotypes. This is the result of the mitochondria’s central role in energy production, reactive oxygen species homeostasis, and cell death. These processes are interdependent and may occur under various stressing conditions, among which low oxygen levels (hypoxia) are certainly prominent. Cells exposed to hypoxia respond acutely with endogenous metabolites and proteins promptly regulating metabolic pathways, but if low oxygen levels are prolonged, cells activate adapting mechanisms, the master switch being the hypoxia-inducible factor 1 (HIF-1). Activation of this factor is strictly bound to the mitochondrial function, which in turn is related with the oxygen level. Therefore in hypoxia, mitochondria act as [O2] sensors, convey signals to HIF-1directly or indirectly, and contribute to the cell redox potential, ion homeostasis, and energy production. Although over the last two decades cellular responses to low oxygen tension have been studied extensively, mechanisms underlying these functions are still indefinite. Here we review current knowledge of the mitochondrial role in hypoxia, focusing mainly on their role in cellular energy and reactive oxygen species homeostasis in relation with HIF-1 stabilization. In addition, we address the involvement of HIF-1 and the inhibitor protein of F1F0 ATPase in the hypoxia-induced mitochondrial autophagy.
Over the last two decades a defective mitochondrial function associated with hypoxia has been invoked in many diverse complex disorders, such as type 2 diabetes [1] and [2], Alzheimer’s disease [3] and [4], cardiac ischemia/reperfusion injury [5] and [6], tissue inflammation [7], and cancer [8], [9], [10],[11] and [12].
The [O2] in air-saturated aqueous buffer at 37 °C is approx. 200 μM [13]; however, mitochondria in vivo are exposed to a considerably lower [O2] that varies with tissue and physiological state. Under physiological conditions, most human resting cells experience some 5% oxygen tension, however the [O2] gradient occurring between the extracellular environment and mitochondria, where oxygen is consumed by cytochrome c oxidase, results in a significantly lower [O2] exposition of mitochondria. Below this oxygen level, most mammalian tissues are exposed to hypoxic conditions [14]. These may arise in normal development, or as a consequence of pathophysiological conditions where there is a reduced oxygen supply due to a respiratory insufficiency or to a defective vasculature. Such conditions include inflammatory diseases, diabetes, ischemic disorders (cerebral or cardiovascular), and solid tumors. Mitochondria consume the greatest amount (some 85–90%) of oxygen in cells to allow oxidative phosphorylation (OXPHOS), which is the primary metabolic pathway for ATP production. Therefore hypoxia will hamper this metabolic pathway, and if the oxygen level is very low, insufficient ATP availability might result in cell death [15].
When cells are exposed to an atmosphere with reduced oxygen concentration, cells readily “respond” by inducing adaptive reactions for their survival through the AMP-activated protein kinase (AMPK) pathway (see for a recent review [16]) which inter alia increases glycolysis driven by enhanced catalytic efficiency of some enzymes, including phosphofructokinase-1 and pyruvate kinase (of note, this oxidative flux is thermodynamically allowed due to both reduced phosphorylation potential [ATP]/([ADP][Pi]) and the physiological redox state of the cell). However, this is particularly efficient only in the short term, therefore cells respond to prolonged hypoxia also by stimulation of hypoxia-inducible factors (HIFs: HIF-1 being the mostly studied), which are heterodimeric transcription factors composed of α and β subunits, first described by Semenza and Wang [17]. These HIFs in the presence of hypoxic oxygen levels are activated through a complex mechanism in which the oxygen tension is critical (see below). Afterwards HIFs bind to hypoxia-responsive elements, activating the transcription of more than two hundred genes that allow cells to adapt to the hypoxic environment [18] and [19].
Several excellent reviews appeared in the last few years describing the array of changes induced by oxygen deficiency in both isolated cells and animal tissues. In in vivo models, a coordinated regulation of tissue perfusion through vasoactive molecules such as nitric oxide and the action of carotid bodies rapidly respond to changes in oxygen demand [20], [21], [22], [23] and [24]. Within isolated cells, hypoxia induces significant metabolic changes due to both variation of metabolites level and activation/inhibition of enzymes and transporters; the most important intracellular effects induced by different pathways are expertly described elsewhere (for recent reviews, see [25], [26] and [27]). It is reasonable to suppose that the type of cells and both the severity and duration of hypoxia may determine which pathways are activated/depressed and their timing of onset [3], [6], [10], [12], [23] and [28]. These pathways will eventually lead to preferential translation of key proteins required for adaptation and survival to hypoxic stress. Although in the past two decades, the discovery of HIF-1 by Gregg Semenza et al. provided a molecular platform to investigate the mechanism underlying responses to oxygen deprivation, the molecular and cellular biology of hypoxia has still to be completely elucidated. This review summarizes recent experimental data concerned with mitochondrial structure and function adaptation to hypoxia and evaluates it in light of the main structural and functional parameters defining the mitochondrial bioenergetics. Since mitochondria contain an inhibitor protein, IF1, whose action on the F1F0 ATPase has been considered for decades of critical importance in hypoxia/ischemia, particular notice will be dedicated to analyze molecular aspects of IF1 regulation of the enzyme and its possible role in the metabolic changes induced by low oxygen levels in cells.
Mechanism(s) of HIF-1 activation
HIF-1 consists of an oxygen-sensitive HIF-1α subunit that heterodimerizes with the HIF-1β subunit to bind DNA. In high O2 tension, HIF-1α is oxidized (hydroxylated) by prolyl hydroxylases (PHDs) using α-ketoglutarate derived from the tricarboxylic acid (TCA) cycle. The hydroxylated HIF-1α subunit interacts with the von Hippel–Lindau protein, a critical member of an E3 ubiquitin ligase complex that polyubiquitylates HIF. This is then catabolized by proteasomes, such that HIF-1α is continuously synthesized and degraded under normoxic conditions [18]. Under hypoxia, HIF-1α hydroxylation does not occur, thereby stabilizing HIF-1 (Fig. 1). The active HIF-1 complex in turn binds to a core hypoxia response element in a wide array of genes involved in a diversity of biological processes, and directly transactivates glycolytic enzyme genes [29]. Notably, O2 concentration, multiple mitochondrial products, including the TCA cycle intermediates and reactive oxygen species, can coordinate PHD activity, HIF stabilization, hence the cellular responses to O2 depletion [30] and [31]. Incidentally, impaired TCA cycle flux, particularly if it is caused by succinate dehydrogenase dysfunction, results in decreased or loss of energy production from both the electron-transport chain and the Krebs cycle, and also in overproduction of free radicals [32]. This leads to severe early-onset neurodegeneration or, as it occurs in individuals carrying mutations in the non-catalytic subunits of the same enzyme, to tumors such as phaeochromocytoma and paraganglioma. However, impairment of the TCA cycle may be relevant also for the metabolic changes occurring in mitochondria exposed to hypoxia, since accumulation of succinate has been reported to inhibit PHDs [33]. It has to be noticed that some authors believe reactive oxygen species (ROS) to be essential to activate HIF-1 [34], but others challenge this idea [35], therefore the role of mitochondrial ROS in the regulation of HIF-1 under hypoxia is still controversial [36]. Moreover, the contribution of functional mitochondria to HIF-1 regulation has also been questioned by others [37], [38] and [39].
http://ars.els-cdn.com/content/image/1-s2.0-S0005272810000575-gr1.jpg

Major mitochondrial changes in hypoxia
Fig. 1. Major mitochondrial changes in hypoxia. Hypoxia could decrease electron-transport rate determining Δψm reduction, increased ROS generation, and enhanced NO synthase. One (or more) of these factors likely contributes to HIF stabilization, that in turn induces metabolic adaptation of both hypoxic cells and mitophagy. The decreased Δψm could also induce an active binding of IF1, which might change mitochondrial morphology and/or dynamics, and inhibit mitophagy. Solid lines indicate well established hypoxic changes in cells, whilst dotted lines indicate changes not yet stated. Inset, relationships between extracellular O2concentration and oxygen tension.
Oxygen is a major determinant of cell metabolism and gene expression, and as cellular O2 levels decrease, either during isolated hypoxia or ischemia-associated hypoxia, metabolism and gene expression profiles in the cells are significantly altered. Low oxygen reduces OXPHOS and Krebs cycle rates, and participates in the generation of nitric oxide (NO), which also contributes to decrease respiration rate [23] and [40]. However, oxygen is also central in the generation of reactive oxygen species, which can participate in cell signaling processes or can induce irreversible cellular damage and death [41].
As specified above, cells adapt to oxygen reduction by inducing active HIF, whose major effect on cells energy homeostasis is the inactivation of anabolism, activation of anaerobic glycolysis, and inhibition of the mitochondrial aerobic metabolism: the TCA cycle, and OXPHOS. Since OXPHOS supplies the majority of ATP required for cellular processes, low oxygen tension will severely reduce cell energy availability. This occurs through several mechanisms: first, reduced oxygen tension decreases the respiration rate, due first to nonsaturating substrate for cytochrome c oxidase (COX), secondarily, to allosteric modulation of COX[42]. As a consequence, the phosphorylation potential decreases, with enhancement of the glycolysis rate primarily due to allosteric increase of phosphofructokinase activity; glycolysis however is poorly efficient and produces lactate in proportion of 0.5 mol/mol ATP, which eventually drops cellular pH if cells are not well perfused, as it occurs under defective vasculature or ischemic conditions [6]. Besides this “spontaneous” (thermodynamically-driven) shift from aerobic to anaerobic metabolism which is mediated by the kinetic changes of most enzymes, the HIF-1 factor activates transcription of genes encoding glucose transporters and glycolytic enzymes to further increase flux of reducing equivalents from glucose to lactate[43] and [44]. Second, HIF-1 coordinates two different actions on the mitochondrial phase of glucose oxidation: it activates transcription of the PDK1 gene encoding a kinase that phosphorylates and inactivates pyruvate dehydrogenase, thereby shunting away pyruvate from the mitochondria by preventing its oxidative decarboxylation to acetyl-CoA [45] and [46]. Moreover, HIF-1 induces a switch in the composition of cytochrome c oxidase from COX4-1 to COX4-2 isoform, which enhances the specific activity of the enzyme. As a result, both respiration rate and ATP level of hypoxic cells carrying the COX4-2 isoform of cytochrome c oxidase were found significantly increased with respect to the same cells carrying the COX4-1 isoform [47]. Incidentally, HIF-1 can also increase the expression of carbonic anhydrase 9, which catalyses the reversible hydration of CO2 to HCO3− and H+, therefore contributing to pH regulation.
Effects of hypoxia on mitochondrial structure and dynamics
Mitochondria form a highly dynamic tubular network, the morphology of which is regulated by frequent fission and fusion events. The fusion/fission machineries are modulated in response to changes in the metabolic conditions of the cell, therefore one should expect that hypoxia affect mitochondrial dynamics. Oxygen availability to cells decreases glucose oxidation, whereas oxygen shortage consumes glucose faster in an attempt to produce ATP via the less efficient anaerobic glycolysis to lactate (Pasteur effect). Under these conditions, mitochondria are not fueled with substrates (acetyl-CoA and O2), inducing major changes of structure, function, and dynamics (for a recent review see [48]). Concerning structure and dynamics, one of the first correlates that emerge is that impairment of mitochondrial fusion leads to mitochondrial depolarization, loss of mtDNA that may be accompanied by altered respiration rate, and impaired distribution of the mitochondria within cells [49], [50] and [51]. Indeed, exposure of cortical neurons to moderate hypoxic conditions for several hours, significantly altered mitochondrial morphology, decreased mitochondrial size and reduced mitochondrial mean velocity. Since these effects were either prevented by exposing the neurons to inhibitors of nitric oxide synthase or mimicked by NO donors in normoxia, the involvement of an NO-mediated pathway was suggested [52]. Mitochondrial motility was also found inhibited and controlled locally by the [ADP]/[ATP] ratio [53]. Interestingly, the author used an original approach in which mitochondria were visualized using tetramethylrhodamineethylester and their movements were followed by applying single-particle tracking.
Of notice in this chapter is that enzymes controlling mitochondrial morphology regulators provide a platform through which cellular signals are transduced within the cell in order to affect mitochondrial function [54]. Accordingly, one might expect that besides other mitochondrial factors [30] and [55] playing roles in HIF stabilization, also mitochondrial morphology might reasonably be associated with HIF stabilization. In order to better define the mechanisms involved in the morphology changes of mitochondria and in their dynamics when cells experience hypoxic conditions, these pioneering studies should be corroborated by and extended to observations on other types of cells focusing also on single proteins involved in both mitochondrial fusion/fission and motion.
Effects of hypoxia on the respiratory chain complexes
O2 is the terminal acceptor of electrons from cytochrome c oxidase (Complex IV), which has a very high affinity for it, being the oxygen concentration for half-maximal respiratory rate at pH 7.4 approximately 0.7 µM [56]. Measurements of mitochondrial oxidative phosphorylation indicated that it is not dependent on oxygen concentration up to at least 20 µM at pH 7.0 and the oxygen dependence becomes markedly greater as the pH is more alkaline [56]. Similarly, Moncada et al. [57] found that the rate of O2 consumption remained constant until [O2] fell below 15 µM. Accordingly, most reports in the literature consider hypoxic conditions occurring in cells at 5–0.5% O2, a range corresponding to 46–4.6 µM O2 in the cells culture medium (see Fig. 1 inset). Since between the extracellular environment and mitochondria an oxygen pressure gradient is established [58], the O2 concentration experienced by Complex IV falls in the range affecting its kinetics, as reported above.
Under these conditions, a number of changes on the OXPHOS machinery components, mostly mediated by HIF-1 have been found. Thus, Semenza et al. [59] and others thereafter [46] reported that activation of HIF-1α induces pyruvate dehydrogenase kinase, which inhibits pyruvate dehydrogenase, suggesting that respiration is decreased by substrate limitation. Besides, other HIF-1 dependent mechanisms capable to affect respiration rate have been reported. First, the subunit composition of COX is altered in hypoxic cells by increased degradation of the COX4-1 subunit, which optimizes COX activity under aerobic conditions, and increased expression of the COX4-2 subunit, which optimizes COX activity under hypoxic conditions [29]. On the other hand, direct assay of respiration rate in cells exposed to hypoxia resulted in a significant reduction of respiration [60]. According with the evidence of Zhang et al., the respiration rate decrease has to be ascribed to mitochondrial autophagy, due to HIF-1-mediated expression of BNIP3. This interpretation is in line with preliminary results obtained in our laboratory where the assay of the citrate synthase activity of cells exposed to different oxygen tensions was performed. Fig. 2 shows the citrate synthase activity, which is taken as an index of the mitochondrial mass [11], with respect to oxygen tension: [O2] and mitochondrial mass are directly linked.

Citrate synthase activity
http://ars.els-cdn.com/content/image/1-s2.0-S0005272810000575-gr2.jpg
Fig. 2. Citrate synthase activity. Human primary fibroblasts, obtained from skin biopsies of 5 healthy donors, were seeded at a density of 8,000 cells/cm2 in high glucose Dulbecco’s Modified Eagle Medium, DMEM (25 mM glucose, 110 mg/l pyruvate, and 4 mM glutamine) supplemented with 15% Foetal Bovine Serum (FBS). 18 h later, cell culture dishes were washed once with Hank’s Balanced Salt Solution (HBSS) and the medium was replaced with DMEM containing 5 mM glucose, 110 mg/l pyruvate, and 4 mM glutamine supplemented with 15% FBS. Cell culture dishes were then placed into an INVIVO2 humidified hypoxia workstation (Ruskinn Technologies, Bridgend, UK) for 72 h changing the medium at 48 h, and oxygen partial pressure (tension) conditions were: 20%, 4%, 2%, 1% and 0.5%. Cells were subsequently collected within the workstation with trypsin-EDTA (0.25%), washed with PBS and resuspended in a buffer containing 10 mM Tris/HCl, 0.1 M KCl, 5 mM KH2PO4, 1 mM EGTA, 3 mM EDTA, and 2 mM MgCl2 pH 7.4 (all the solutions were preconditioned to the appropriate oxygen tension condition). The citrate synthase activity was assayed essentially by incubating 40 µg of cells with 0.02% Triton X-100, and monitoring the reaction by measuring spectrophotometrically the rate of free coenzyme A released, as described in [90]. Enzymatic activity was expressed as nmol/min/mg of protein. Three independent experiments were carried out and assays were performed in either duplicate or triplicate.
However, the observations of Semenza et al. must be seen in relation with data reported by Moncada et al.[57] and confirmed by others [61] in which it is clearly shown that when cells (various cell lines) experience hypoxic conditions, nitric oxide synthases (NOSs) are activated, therefore NO is released. As already mentioned above, NO is a strong competitor of O2 for cytochrome c oxidase, whose apparent Km results increased, hence reduction of mitochondrial cytochromes and all the other redox centres of the respiratory chain occurs. In addition, very recent data indicate a potential de-activation of Complex I when oxygen is lacking, as it occurs in prolonged hypoxia [62]. According to Hagen et al. [63] the NO-dependent inhibition of cytochrome c oxidase should allow “saved” O2 to redistribute within the cell to be used by other enzymes, including PHDs which inactivate HIF. Therefore, unless NO inhibition of cytochrome c oxidase occurs only when [O2] is very low, inhibition of mitochondrial oxygen consumption creates the paradox of a situation in which the cell may fail to register hypoxia. It has been tempted to solve this paradox, but to date only hypotheses have been proposed [23] and [26]. Interestingly, recent observations on yeast cells exposed to hypoxia revealed abnormal protein carbonylation and protein tyrosine nitration that were ascribed to increased mitochondrially generated superoxide radicals and NO, two species typically produced at low oxygen levels, that combine to form ONOO− [64]. Based on these studies a possible explanation has been proposed for the above paradox.
Finally, it has to be noticed that the mitochondrial respiratory deficiency observed in cardiomyocytes of dogs in which experimental heart failure had been induced lies in the supermolecular assembly rather than in the individual components of the electron-transport chain [65]. This observation is particularly intriguing since loss of respirasomes is thought to facilitate ROS generation in mitochondria [66], therefore supercomplexes disassembly might explain the paradox of reduced [O2] and the enhanced ROS found in hypoxic cells. Specifically, hypoxia could reduce mitochondrial fusion by impairing mitochondrial membrane potential, which in turn could induce supercomplexes disassembly, increasing ROS production[11].
Complex III and ROS production
It has been estimated that, under normoxic physiological conditions, 1–2% of electron flow through the mitochondrial respiratory chain gives rise to ROS [67] and [68]. It is now recognized that the major sites of ROS production are within Complexes I and III, being prevalent the contribution of Complex I [69] (Fig. 3). It might be expected that hypoxia would decrease ROS production, due to the low level of O2 and to the diminished mitochondrial respiration [6] and [46], but ROS level is paradoxically increased. Indeed, about a decade ago, Chandel et al. [70] provided good evidence that mitochondrial reactive oxygen species trigger hypoxia-induced transcription, and a few years later the same group [71] showed that ROS generated at Complex III of the mitochondrial respiratory chain stabilize HIF-1α during hypoxia (Fig. 1 and Fig. 3). Although others have proposed mechanisms indicating a key role of mitochondria in HIF-1α regulation during hypoxia (for reviews see [64] and [72]), the contribution of mitochondria to HIF-1 regulation has been questioned by others [35], [36] and [37]. Results of Gong and Agani [35] for instance show that inhibition of electron-transport Complexes I, III, and IV, as well as inhibition of mitochondrial F0F1 ATPase, prevents HIF-1α expression and that mitochondrial reactive oxygen species are not involved in HIF-1α regulation during hypoxia. Concurrently, Tuttle et al. [73], by means of a non invasive, spectroscopic approach, could find no evidence to suggest that ROS, produced by mitochondria, are needed to stabilize HIF-1α under moderate hypoxia. The same authors found the levels of HIF-1α comparable in both normal and ρ0 cells (i.e. cells lacking mitochondrial DNA). On the contrary, experiments carried out on genetic models consisting of either cells lacking cytochrome c or ρ0 cells both could evidence the essential role of mitochondrial respiration to stabilize HIF-1α [74]. Thus, cytochrome c null cells, being incapable to respire, exposed to moderate hypoxia (1.5% O2) prevented oxidation of ubiquinol and generation of the ubisemiquinone radical, thus eliminating superoxide formation at Complex III [71]. Concurrently, ρ0 cells lacking electron transport, exposed 4 h to moderate hypoxia failed to stabilize HIF-1α, suggesting the essential role of the respiratory chain for the cellular sensing of low O2 levels. In addition, recent evidence obtained on genetic manipulated cells (i.e. cytochrome b deficient cybrids) showed increased ROS levels and stabilized HIF-1α protein during hypoxia [75]. Moreover, RNA interference of the Complex III subunit Rieske iron sulfur protein in the cytochrome b deficient cells, abolished ROS generation at the Qo site of Complex III, preventing HIF-1α stabilization. These observations, substantiated by experiments with MitoQ, an efficient mitochondria-targeted antioxidant, strongly support the involvement of mitochondrial ROS in regulating HIF-1α. Nonetheless, collectively, the available data do not allow to definitely state the precise role of mitochondrial ROS in regulating HIF-1α, but the pathway stabilizing HIF-1α appears undoubtedly mitochondria-dependent [30].

Overview of mitochondrial electron and proton flux in hypoxia
Overview of mitochondrial electron and proton flux in hypoxia
http://ars.els-cdn.com/content/image/1-s2.0-S0005272810000575-gr3.jpg
Fig. 3. Overview of mitochondrial electron and proton flux in hypoxia. Electrons released from reduced cofactors (NADH and FADH2) under normoxia flow through the redox centres of the respiratory chain (r.c.) to molecular oxygen (blue dotted line), to which a proton flux from the mitochondrial matrix to the intermembrane space is coupled (blue arrows). Protons then flow back to the matrix through the F0 sector of the ATP synthase complex, driving ATP synthesis. ATP is carried to the cell cytosol by the adenine nucleotide translocator (blue arrows). Under moderate to severe hypoxia, electrons escape the r.c. redox centres and reduce molecular oxygen to the superoxide anion radical before reaching the cytochrome c (red arrow). Under these conditions, to maintain an appropriate Δψm, ATP produced by cytosolic glycolysis enters the mitochondria where it is hydrolyzed by the F1F0ATPase with extrusion of protons from the mitochondrial matrix (red arrows).
Hypoxia and ATP synthase
The F1F0 ATPase (ATP synthase) is the enzyme responsible of catalysing ADP phosphorylation as the last step of OXPHOS. It is a rotary motor using the proton motive force across the mitochondrial inner membrane to drive the synthesis of ATP [76]. It is a reversible enzyme with ATP synthesis or hydrolysis taking place in the F1 sector at the matrix side of the membrane, chemical catalysis being coupled to H+transport through the transmembrane F0 sector.
Under normoxia the enzyme synthesizes ATP, but when mitochondria experience hypoxic conditions the mitochondrial membrane potential (Δψm) decreases below its endogenous steady-state level (some 140 mV, negative inside the matrix [77]) and the F1F0 ATPase may work in the reversal mode: it hydrolyses ATP (produced by anaerobic glycolysis) and uses the energy released to pump protons from the mitochondrial matrix to the intermembrane space, concurring with the adenine nucleotide translocator (i.e. in hypoxia it exchanges cytosolic ATP4− for matrix ADP3−) to maintain the physiological Δψm ( Fig. 3). Since under conditions of limited oxygen availability the decline in cytoplasmic high energy phosphates is mainly due to hydrolysis by the ATP synthase working in reverse [6] and [78], the enzyme must be strictly regulated in order to avoid ATP dissipation. This is achieved by a natural protein, the H+/Δψm-dependent IF1, that binds to the catalytic F1 sector at low pH and low Δψm (such as it occurs in hypoxia/ischemia) [79]. IF1 binding to the ATP synthase results in a rapid and reversible inhibition of the enzyme [80], which could reach about 50% of maximal activity (for recent reviews see [6] and [81]).
Besides this widely studied effect, IF1 appears to be associated with ROS production and mitochondrial autophagy (mitophagy). This is a mechanism involving the catabolic degradation of macromolecules and organelles via the lysosomal pathway that contributes to housekeeping and regenerate metabolites. Autophagic degradation is involved in the regulation of the ageing process and in several human diseases, such as myocardial ischemia/reperfusion [82], Alzheimer’s Disease, Huntington diseases, and inflammatory diseases (for recent reviews see [83] and [84], and, as mentioned above, it promotes cell survival by reducing ROS and mtDNA damage under hypoxic conditions.
Campanella et al. [81] reported that, in HeLa cells under normoxic conditions, basal autophagic activity varies in relation to the expression levels of IF1. Accordingly, cells overexpressing IF1 result in ROS production similar to controls, conversely cells in which IF1 expression is suppressed show an enhanced ROS production. In parallel, the latter cells show activation of the mitophagy pathway (Fig. 1), therefore suggesting that variations in IF1 expression level may play a significant role in defining two particularly important parameters in the context of the current review: rates of ROS generation and mitophagy. Thus, the hypoxia-induced enhanced expression level of IF1[81] should be associated with a decrease of both ROS production and autophagy, which is in apparent conflict with the hypoxia-induced ROS increase and with the HIF-1-dependent mitochondrial autophagy shown by Zhang et al. [60] as an adaptive metabolic response to hypoxia. However, in the experiments of Zhang et al. the cells were exposed to hypoxia for 48 h, whereas the F1F0-ATPase inhibitor exerts a prompt action on the enzyme and to our knowledge, it has never been reported whether its action persists during prolonged hypoxic expositions. Pertinent with this problem is the very recent observation that IEX-1 (immediate early response gene X-1), a stress-inducible gene that suppresses production of ROS and protects cells from apoptosis [85], targets the mitochondrial F1F0-ATPase inhibitor for degradation, reducing ROS by decreasing Δψm. It has to be noticed that the experiments described were carried out under normal oxygen availability, but it does not seem reasonable to rule out IEX-1 from playing a role under stress conditions as those induced by hypoxia in cells, therefore this issue might deserve an investigation also at low oxygen levels.
In conclusion, data are still emerging regarding the regulation of mitochondrial function by the F1F0 ATPase within hypoxic responses in different cellular and physiological contexts. Given the broad pathophysiological role of hypoxic cellular modulation, an understanding of the subtle tuning among different effectors of the ATP synthase is desirable to eventually target future therapeutics most effectively. Our laboratory is actually involved in carrying out investigations to clarify this context.
Conclusions and perspectives
The mitochondria are important cellular platforms that both propagate and initiate intracellular signals that lead to overall cellular and metabolic responses. During the last decades, a significant amount of relevant data has been obtained on the identification of mechanisms of cellular adaptation to hypoxia. In hypoxic cells there is an enhanced transcription and synthesis of several glycolytic pathway enzymes/transporters and reduction of synthesis of proteins involved in mitochondrial catabolism. Although well defined kinetic parameters of reactions in hypoxia are lacking, it is usually assumed that these transcriptional changes lead to metabolic flux modification. The required biochemical experimentation has been scarcely addressed until now and only in few of the molecular and cellular biology studies the transporter and enzyme kinetic parameters and flux rate have been determined, leaving some uncertainties.
Central to mitochondrial function and ROS generation is an electrochemical proton gradient across the mitochondrial inner membrane that is established by the proton pumping activity of the respiratory chain, and that is strictly linked to the F1F0-ATPase function. Evaluation of the mitochondrial membrane potential in hypoxia has only been studied using semiquantitative methods based on measurements of the fluorescence intensity of probes taken up by cells experiencing normal or hypoxic conditions. However, this approach is intrinsically incorrect due to the different capability that molecular oxygen has to quench fluorescence [86] and [87] and to the uncertain concentration the probe attains within mitochondria, whose mass may be reduced by a half in hypoxia [60]. In addition, the uncertainty about measurement of mitochondrial superoxide radical and H2O2 formation in vivo [88] hampers studies on the role of mitochondrial ROS in hypoxic oxidative damage, redox signaling, and HIF-1 stabilization.
The duration and severity of hypoxic stress differentially activate the responses discussed throughout and lead to substantial phenotypic variations amongst tissues and cell models, which are not consistently and definitely known. Certainly, understanding whether a hierarchy among hypoxia response mechanisms exists and which are the precise timing and conditions of each mechanism to activate, will improve our knowledge of the biochemical mechanisms underlying hypoxia in cells, which eventually may contribute to define therapeutic targets in hypoxia-associated diseases. To this aim it might be worth investigating the hypoxia-induced structural organization of both the respiratory chain enzymes in supramolecular complexes and the assembly of the ATP synthase to form oligomers affecting ROS production [65] and inner mitochondrial membrane structure [89], respectively.
7.9.2 Hypoxia promotes isocitrate dehydrogenase-dependent carboxylation of α-ketoglutarate to citrate to support cell growth and viability
DR Wise, PS Ward, JES Shay, JR Cross, JJ Gruber, UM Sachdeva, et al.
Proc Nat Acad Sci Oct 27, 2011; 108(49):19611–19616
http://dx.doi.org:/10.1073/pnas.1117773108
Citrate is a critical metabolite required to support both mitochondrial bioenergetics and cytosolic macromolecular synthesis. When cells proliferate under normoxic conditions, glucose provides the acetyl-CoA that condenses with oxaloacetate to support citrate production. Tricarboxylic acid (TCA) cycle anaplerosis is maintained primarily by glutamine. Here we report that some hypoxic cells are able to maintain cell proliferation despite a profound reduction in glucose-dependent citrate production. In these hypoxic cells, glutamine becomes a major source of citrate. Glutamine-derived α-ketoglutarate is reductively carboxylated by the NADPH-linked mitochondrial isocitrate dehydrogenase (IDH2) to form isocitrate, which can then be isomerized to citrate. The increased IDH2-dependent carboxylation of glutamine-derived α-ketoglutarate in hypoxia is associated with a concomitant increased synthesis of 2-hydroxyglutarate (2HG) in cells with wild-type IDH1 and IDH2. When either starved of glutamine or rendered IDH2-deficient by RNAi, hypoxic cells are unable to proliferate. The reductive carboxylation of glutamine is part of the metabolic reprogramming associated with hypoxia-inducible factor 1 (HIF1), as constitutive activation of HIF1 recapitulates the preferential reductive metabolism of glutamine-derived α-ketoglutarate even in normoxic conditions. These data support a role for glutamine carboxylation in maintaining citrate synthesis and cell growth under hypoxic conditions.
Citrate plays a critical role at the center of cancer cell metabolism. It provides the cell with a source of carbon for fatty acid and cholesterol synthesis (1). The breakdown of citrate by ATP-citrate lyase is a primary source of acetyl-CoA for protein acetylation (2). Metabolism of cytosolic citrate by aconitase and IDH1 can also provide the cell with a source of NADPH for redox regulation and anabolic synthesis. Mammalian cells depend on the catabolism of glucose and glutamine to fuel proliferation (3). In cancer cells cultured at atmospheric oxygen tension (21% O2), glucose and glutamine have both been shown to contribute to the cellular citrate pool, with glutamine providing the major source of the four-carbon molecule oxaloacetate and glucose providing the major source of the two-carbon molecule acetyl-CoA (4, 5). The condensation of oxaloacetate and acetyl-CoA via citrate synthase generates the 6 carbon citrate molecule. However, both the conversion of glucose-derived pyruvate to acetyl-CoA by pyruvate dehydrogenase (PDH) and the conversion of glutamine to oxaloacetate through the TCA cycle depend on NAD+, which can be compromised under hypoxic conditions. This raises the question of how cells that can proliferate in hypoxia continue to synthesize the citrate required for macromolecular synthesis.
This question is particularly important given that many cancers and stem/progenitor cells can continue proliferating in the setting of limited oxygen availability (6, 7). Louis Pasteur first highlighted the impact of hypoxia on nutrient metabolism based on his observation that hypoxic yeast cells preferred to convert glucose into lactic acid rather than burning it in an oxidative fashion. The molecular basis for this shift in mammalian cells has been linked to the activity of the transcription factor HIF1 (8–10). Stabilization of the labile HIF1α subunit occurs in hypoxia. It can also occur in normoxia through several mechanisms including loss of the von Hippel-Lindau tumor suppressor (VHL), a common occurrence in renal carcinoma (11). Although hypoxia and/or HIF1α stabilization is a common feature of multiple cancers, to date the source of citrate in the setting of hypoxia or HIF activation has not been determined.
Here, we study the sources of hypoxic citrate synthesis in a glioblastoma cell line that proliferates in profound hypoxia (0.5% O2). Glucose uptake and conversion to lactic acid increased in hypoxia. However, glucose conversion into citrate dramatically declined. Glutamine consumption remained constant in hypoxia, and hypoxic cells were addicted to the use of glutamine in hypoxia as a source of α-ketoglutarate. Glutamine provided the major carbon source for citrate synthesis during hypoxia. However, the TCA cycle-dependent conversion of glutamine into citric acid was significantly suppressed. In contrast, there was a relative increase in glutamine-dependent citrate production in hypoxia that resulted from carboxylation of α-ketoglutarate. This reductive synthesis required the presence of mitochondrial isocitrate dehydrogenase 2 (IDH2). In confirmation of the reverse flux through IDH2, the increased reductive metabolism of glutamine-derived α-ketoglutarate in hypoxia was associated with increased synthesis of 2HG. Finally, constitutive HIF1α-expressing cells also demonstrated significant reductive-carboxylation-dependent synthesis of citrate in normoxia and a relative defect in the oxidative conversion of glutamine into citrate. Collectively, the data demonstrate that mitochondrial glutamine metabolism can be rerouted through IDH2-dependent citrate synthesis in support of hypoxic cell growth.
Some Cancer Cells Can Proliferate at 0.5% O2 Despite a Sharp Decline in Glucose-Dependent Citrate Synthesis.
At 21% O2, cancer cells have been shown to synthesize citrate by condensing glucose-derived acetyl-CoA with glutamine-derived oxaloacetate through the activity of the canonical TCA cycle enzyme citrate synthase (4). In contrast, less is known regarding the synthesis of citrate by cells that can continue proliferating in hypoxia. The glioblastoma cell line SF188 is able to proliferate at 0.5% O2 (Fig. 1A), a level of hypoxia that is sufficient to stabilize HIF1α (Fig. 1B) and predicted to limit respiration (12, 13). Consistent with previous observations in hypoxic cells, we found that SF188 cells demonstrated increased lactate production when incubated in hypoxia (Fig. 1C), and the ratio of lactate produced to glucose consumed increased demonstrating an increase in the rate of anaerobic glycolysis. When glucose-derived carbon in the form of pyruvate is converted to lactate, it is diverted away from subsequent metabolism that can contribute to citrate production. However, we observed that SF188 cells incubated in hypoxia maintain their intracellular citrate to ∼75% of the level maintained under normoxia (Fig. 1D). This prompted an investigation of how proliferating cells maintain citrate production under hypoxia.

SF188 glioblastoma cells proliferate at 0.5% O2 despite a profound reduction in glucose-dependent citrate synthesis.
http://www.pnas.org/content/108/49/19611/F1.medium.gif
Fig. 1. SF188 glioblastoma cells proliferate at 0.5% O2 despite a profound reduction in glucose-dependent citrate synthesis. (A) SF188 cells were plated in complete medium equilibrated with 21% O2 (Normoxia) or 0.5% O2 (Hypoxia), total viable cells were counted 24 h and 48 h later (Day 1 and Day 2), and population doublings were calculated. Data are the mean ± SEM of four independent experiments. (B) Western blot demonstrates stabilized HIF1α protein in cells cultured in hypoxia compared with normoxia. (C) Cells were grown in normoxia or hypoxia for 24 h, after which culture medium was collected. Medium glucose and lactate levels were measured and compared with the levels in fresh medium. (D) Cells were cultured for 24 h as in C. Intracellular metabolism was then quenched with 80% MeOH prechilled to −80 °C that was spiked with a 13C-labeled citrate as an internal standard. Metabolites were then extracted, and intracellular citrate levels were analyzed with GC-MS and normalized to cell number. Data for C and D are the mean ± SEM of three independent experiments. (E) Model depicting the pathway for cit+2 production from [U-13C]glucose. Glucose uniformly 13C-labeled will generate pyruvate+3. Pyruvate+3 can be oxidatively decarboxylated by PDH to produce acetyl-CoA+2, which can condense with unlabeled oxaloacetate to produce cit+2. (F) Cells were cultured for 24 h as in C and D, followed by an additional 4 h of culture in glucose-deficient medium supplemented with 10 mM [U-13C]glucose. Intracellular metabolites were then extracted, and 13C-enrichment in cellular citrate was analyzed by GC-MS and normalized to the total citrate pool size. Data are the mean ± SD of three independent cultures from a representative of two independent experiments. *P < 0.05, ***P < 0.001.
Increased glucose uptake and glycolytic metabolism are critical elements of the metabolic response to hypoxia. To evaluate the contributions made by glucose to the citrate pool under normoxia or hypoxia, SF188 cells incubated in normoxia or hypoxia were cultured in medium containing 10 mM [U-13C]glucose. Following a 4-h labeling period, cellular metabolites were extracted and analyzed for isotopic enrichment by gas chromatography-mass spectrometry (GC-MS). In normoxia, the major 13C-enriched citrate species found was citrate enriched with two 13C atoms (cit+2), which can arise from the NAD+-dependent decarboxylation of pyruvate+3 to acetyl-CoA+2 by PDH, followed by the condensation of acetyl-CoA+2 with unenriched oxaloacetate (Fig. 1 E and F). Compared with the accumulation of cit+2, we observed minimal accumulation of cit+3 and cit+5 under normoxia. Cit+3 arises from pyruvate carboxylase (PC)-dependent conversion of pyruvate+3 to oxaloacetate+3, followed by the condensation of oxaloacetate+3 with unenriched acetyl-CoA. Cit+5 arises when PC-generated oxaloacetate+3 condenses with PDH-generated acetyl-CoA+2. The lack of cit+3 and cit+5 accumulation is consistent with PC activity not playing a major role in citrate production in normoxic SF188 cells, as reported (4).
In hypoxic cells, the major citrate species observed was unenriched. Cit+2, cit+3, and cit+5 all constituted minor fractions of the total citrate pool, consistent with glucose carbon not being incorporated into citrate through either PDH or PC-mediated metabolism under hypoxic conditions (Fig. 1F). These data demonstrate that in contrast to normoxic cells, where a large percentage of citrate production depends on glucose-derived carbon, hypoxic cells significantly reduce their rate of citrate production from glucose.
Glutamine Carbon Metabolism Is Required for Viability in Hypoxia.
In addition to glucose, we have previously reported that glutamine can contribute to citrate production during cell growth under normoxic conditions (4). Surprisingly, under hypoxic conditions, we observed that SF188 cells retained their high rate of glutamine consumption (Fig. 2A). Moreover, hypoxic cells cultured in glutamine-deficient medium displayed a significant loss of viability (Fig. 2B). In normoxia, the requirement for glutamine to maintain viability of SF188 cells can be satisfied by α-ketoglutarate, the downstream metabolite of glutamine that is devoid of nitrogenous groups (14). α-ketoglutarate cannot fulfill glutamine’s roles as a nitrogen source for nonessential amino acid synthesis or as an amide donor for nucleotide or hexosamine synthesis, but can be metabolized through the oxidative TCA cycle to regenerate oxaloacetate, and subsequently condense with glucose-derived acetyl-CoA to produce citrate. To test whether the restoration of carbon from glutamine metabolism in the form of α-ketoglutarate could rescue the viability defect of glutamine-starved SF188 cells even under hypoxia, SF188 cells incubated in hypoxia were cultured in glutamine-deficient medium supplemented with a cell-penetrant form of α-ketoglutarate (dimethyl α-ketoglutarate). The addition of dimethyl α-ketoglutarate rescued the defect in cell viability observed upon glutamine withdrawal (Fig. 2B). These data demonstrate that, even under hypoxic conditions, when the ability of glutamine to replenish oxaloacetate through oxidative TCA cycle metabolism is diminished, SF188 cells retain their requirement for glutamine as the carbon backbone for α-ketoglutarate. This result raised the possibility that glutamine could be the carbon source for citrate production through an alternative, nonoxidative, pathway in hypoxia.

Glutamine carbon is required for hypoxic cell viability
Glutamine carbon is required for hypoxic cell viability
http://www.pnas.org/content/108/49/19611/F2.medium.gif
Fig. 2. Glutamine carbon is required for hypoxic cell viability and contributes to increased citrate production through reductive carboxylation relative to oxidative metabolism in hypoxia. (A) SF188 cells were cultured for 24 h in complete medium equilibrated with either 21% O2 (Normoxia) or 0.5% O2(Hypoxia). Culture medium was then removed from cells and analyzed for glutamine levels which were compared with the glutamine levels in fresh medium. Data are the mean ± SEM of three independent experiments. (B) The requirement for glutamine to maintain hypoxic cell viability can be satisfied by α-ketoglutarate. Cells were cultured in complete medium equilibrated with 0.5% O2 for 24 h, followed by an additional 48 h at 0.5% O2 in either complete medium (+Gln), glutamine-deficient medium (−Gln), or glutamine-deficient medium supplemented with 7 mM dimethyl α-ketoglutarate (−Gln +αKG). All medium was preconditioned in 0.5% O2. Cell viability was determined by trypan blue dye exclusion. Data are the mean and range from two independent experiments. (C) Model depicting the pathways for cit+4 and cit+5 production from [U-13C]glutamine (glutamine+5). Glutamine+5 is catabolized to α-ketoglutarate+5, which can then contribute to citrate production by two divergent pathways. Oxidative metabolism produces oxaloacetate+4, which can condense with unlabeled acetyl-CoA to produce cit+4. Alternatively, reductive carboxylation produces isocitrate+5, which can isomerize to cit+5. (D) Glutamine contributes to citrate production through increased reductive carboxylation relative to oxidative metabolism in hypoxic proliferating cancer cells. Cells were cultured for 24 h as in A, followed by 4 h of culture in glutamine-deficient medium supplemented with 4 mM [U-13C]glutamine. 13C enrichment in cellular citrate was quantitated with GC-MS. Data are the mean ± SD of three independent cultures from a representative of three independent experiments. **P < 0.01.
Cells Proliferating in Hypoxia Maintain Levels of Additional Metabolites Through Reductive Carboxylation.
Previous work has documented that, in normoxic conditions, SF188 cells use glutamine as the primary anaplerotic substrate, maintaining the pool sizes of TCA cycle intermediates through oxidative metabolism (4). Surprisingly, we found that, when incubated in hypoxia, SF188 cells largely maintained their levels of aspartate (in equilibrium with oxaloacetate), malate, and fumarate (Fig. 3A). To distinguish how glutamine carbon contributes to these metabolites in normoxia and hypoxia, SF188 cells incubated in normoxia or hypoxia were cultured in medium containing 4 mM [U-13C]glutamine. After a 4-h labeling period, metabolites were extracted and the intracellular pools of aspartate, malate, and fumarate were analyzed by GC-MS.
In normoxia, the majority of the enriched intracellular asparatate, malate, and fumarate were the +4 species, which arise through oxidative metabolism of glutamine-derived α-ketoglutarate (Fig. 3 B and C). The +3 species, which can be derived from the citrate generated by the reductive carboxylation of glutamine-derived α-ketoglutarate, constituted a significantly lower percentage of the total aspartate, malate, and fumarate pools. By contrast, in hypoxia, the +3 species constituted a larger percentage of the total aspartate, malate, and fumarate pools than they did in normoxia. These data demonstrate that, in addition to citrate, hypoxic cells preferentially synthesize oxaloacetate, malate, and fumarate through the pathway of reductive carboxylation rather than the oxidative TCA cycle.
IDH2 Is Critical in Hypoxia for Reductive Metabolism of Glutamine and for Cell Proliferation.
We hypothesized that the relative increase in reductive carboxylation we observed in hypoxia could arise from the suppression of α-ketoglutarate oxidation through the TCA cycle. Consistent with this, we found that α-ketoglutarate levels increased in SF188 cells following 24 h in hypoxia (Fig. 4A). Surprisingly, we also found that levels of the closely related metabolite 2-hydroxyglutarate (2HG) increased in hypoxia, concomitant with the increase in α-ketoglutarate under these conditions. 2HG can arise from the noncarboxylating reduction of α-ketoglutarate (Fig. 4B). Recent work has found that specific cancer-associated mutations in the active sites of either IDH1 or IDH2 lead to a 10- to 100-fold enhancement in this activity facilitating 2HG production (15–17), but SF188 cells lack IDH1/2 mutations. However, 2HG levels are also substantially elevated in the inborn error of metabolism 2HG aciduria, and the majority of patients with this disease lack IDH1/2 mutations. As 2HG has been demonstrated to arise in these patients from mitochondrial α-ketoglutarate (18), we hypothesized that both the increased reductive carboxylation of glutamine-derived α-ketoglutarate to citrate and the increased 2HG accumulation we observed in hypoxia could arise from increased reductive metabolism by wild-type IDH2 in the mitochondria.

Reductive carboxylation of glutamine-derived α-ketoglutarate to citrate in hypoxic cancer cells is dependent on mitochondrial IDH2
Reductive carboxylation of glutamine-derived α-ketoglutarate to citrate in hypoxic cancer cells is dependent on mitochondrial IDH2
http://www.pnas.org/content/108/49/19611/F4.medium.gif
Fig. 4. Reductive carboxylation of glutamine-derived α-ketoglutarate to citrate in hypoxic cancer cells is dependent on mitochondrial IDH2. (A) α-ketoglutarate and 2HG increase in hypoxia. SF188 cells were cultured in complete medium equilibrated with either 21% O2 (Normoxia) or 0.5% O2 (Hypoxia) for 24 h. Intracellular metabolites were then extracted, cell extracts spiked with a 13C-labeled citrate as an internal standard, and intracellular α-ketoglutarate and 2HG levels were analyzed with GC-MS. Data shown are the mean ± SEM of three independent experiments. (B) Model for reductive metabolism from glutamine-derived α-ketoglutarate. Glutamine+5 is catabolized to α-ketoglutarate+5. Carboxylation of α-ketoglutarate+5 followed by reduction of the carboxylated intermediate (reductive carboxylation) will produce isocitrate+5, which can then isomerize to cit+5. In contrast, reductive activity on α-ketoglutarate+5 that is uncoupled from carboxylation will produce 2HG+5. (C) IDH2 is required for reductive metabolism of glutamine-derived α-ketoglutarate in hypoxia. SF188 cells transfected with a siRNA against IDH2 (siIDH2) or nontargeting negative control (siCTRL) were cultured for 2 d in complete medium equilibrated with 0.5% O2. (Upper) Cells were then cultured at 0.5% O2 for an additional 4 h in glutamine-deficient medium supplemented with 4 mM [U-13C]glutamine. 13C enrichment in intracellular citrate and 2HG was determined and normalized to the relevant metabolite total pool size. (Lower) Cells transfected and cultured in parallel at 0.5% O2 were counted by hemacytometer (excluding nonviable cells with trypan blue staining) or harvested for protein to assess IDH2 expression by Western blot. Data shown for GC-MS and cell counts are the mean ± SD of three independent cultures from a representative experiment. **P < 0.01, ***P < 0.001.
In an experiment to test this hypothesis, SF188 cells were transfected with either siRNA directed against mitochondrial IDH2 (siIDH2) or nontargeting control, incubated in hypoxia for 2 d, and then cultured for another 4 h in hypoxia in media containing 4 mM [U-13C]glutamine. After the labeling period, metabolites were extracted and analyzed by GC-MS (Fig. 4C). Hypoxic SF188 cells transfected with siIDH2 displayed a decreased contribution of cit+5 to the total citrate pool, supporting an important role for IDH2 in the reductive carboxylation of glutamine-derived α-ketoglutarate in hypoxic conditions. The contribution of cit+4 to the total citrate pool did not decrease with siIDH2 treatment, consistent with IDH2 knockdown specifically affecting the pathway of reductive carboxylation and not other fundamental TCA cycle-regulating processes. In confirmation of reverse flux occurring through IDH2, the contribution of 2HG+5 to the total 2HG pool decreased in siIDH2-treated cells. Supporting the importance of citrate production by IDH2-mediated reductive carboxylation for hypoxic cell proliferation, siIDH2-transfected SF188 cells displayed a defect in cellular accumulation in hypoxia. Decreased expression of IDH2 protein following siIDH2 transfection was confirmed by Western blot. Collectively, these data point to the importance of mitochondrial IDH2 for the increase in reductive carboxylation flux of glutamine-derived α-ketoglutarate to maintain citrate levels in hypoxia, and to the importance of this reductive pathway for hypoxic cell proliferation.
Reprogramming of Metabolism by HIF1 in the Absence of Hypoxia Is Sufficient to Induce Increased Citrate Synthesis by Reductive Carboxylation Relative to Oxidative Metabolism.
The relative increase in the reductive metabolism of glutamine-derived α-ketoglutarate at 0.5% O2 may be explained by the decreased ability to carry out oxidative NAD+-dependent reactions as respiration is inhibited (12, 13). However, a shift to preferential reductive glutamine metabolism could also result from the active reprogramming of cellular metabolism by HIF1 (8–10), which inhibits the generation of mitochondrial acetyl-CoA necessary for the synthesis of citrate by oxidative glucose and glutamine metabolism (Fig. 5A). To better understand the role of HIF1 in reductive glutamine metabolism, we used VHL-deficient RCC4 cells, which display constitutive expression of HIF1α under normoxia (Fig. 5B). RCC4 cells expressing either a nontargeting control shRNA (shCTRL) or an shRNA directed at HIF1α (shHIF1α) were incubated in normoxia and cultured in medium with 4 mM [U-13C]glutamine. Following a 4-h labeling period, metabolites were extracted and the cellular citrate pool was analyzed by GC-MS. In shCTRL cells, which have constitutive HIF1α expression despite incubation in normoxia, the majority of the total citrate pool was constituted by the cit+5 species, with low levels of all other species including cit+4 (Fig. 5C). By contrast, in HIF1α-deficient cells the contribution of cit+5 to the total citrate pool was greatly decreased, whereas the contribution of cit+4 to the total citrate pool increased and was the most abundant citrate species. These data demonstrate that the relative enhancement of the reductive carboxylation pathway for citrate synthesis can be recapitulated by constitutive HIF1 activation in normoxia.

Reprogramming of metabolism by HIF1 in the absence of hypoxia
http://www.pnas.org/content/108/49/19611/F5.medium.gif
Reprogramming of metabolism by HIF1 in the absence of hypoxia is sufficient to induce reductive carboxylation of glutamine-derived α-ketoglutarate.
Fig. 5. Reprogramming of metabolism by HIF1 in the absence of hypoxia is sufficient to induce reductive carboxylation of glutamine-derived α-ketoglutarate. (A) Model depicting how HIF1 signaling’s inhibition of pyruvate dehydrogenase (PDH) activity and promotion of lactate dehydrogenase-A (LDH-A) activity can block the generation of mitochondrial acetyl-CoA from glucose-derived pyruvate, thereby favoring citrate synthesis from reductive carboxylation of glutamine-derived α-ketoglutarate. (B) Western blot demonstrating HIF1α protein in RCC4 VHL−/− cells in normoxia with a nontargeting shRNA (shCTRL), and the decrease in HIF1α protein in RCC4 VHL−/− cells stably expressing HIF1α shRNA (shHIF1α). (C) HIF1-induced reprogramming of glutamine metabolism. Cells from B at 21% O2 were cultured for 4 h in glutamine-deficient medium supplemented with 4 mM [U-13C]glutamine. Intracellular metabolites were then extracted, and 13C enrichment in cellular citrate was determined by GC-MS. Data shown are the mean ± SD of three independent cultures from a representative of three independent experiments. ***P < 0.001.
Compared with glucose metabolism, much less is known regarding how glutamine metabolism is altered under hypoxia. It has also remained unclear how hypoxic cells can maintain the citrate production necessary for macromolecular biosynthesis. In this report, we demonstrate that in contrast to cells at 21% O2, where citrate is predominantly synthesized through oxidative metabolism of both glucose and glutamine, reductive carboxylation of glutamine carbon becomes the major pathway of citrate synthesis in cells that can effectively proliferate at 0.5% O2. Moreover, we show that in these hypoxic cells, reductive carboxylation of glutamine-derived α-ketoglutarate is dependent on mitochondrial IDH2. Although others have previously suggested the existence of reductive carboxylation in cancer cells (19, 20), these studies failed to demonstrate the intracellular localization or specific IDH isoform responsible for the reductive carboxylation flux. Recently, we identified IDH2 as an isoform that contributes to reductive carboxylation in cancer cells incubated at 21% O2 (16), but remaining unclear were the physiological importance and regulation of this pathway relative to oxidative metabolism, as well as the conditions where this reductive pathway might be advantageous for proliferating cells.
Here we report that IDH2-mediated reductive carboxylation of glutamine-derived α-ketoglutarate to citrate is an important feature of cells proliferating in hypoxia. Moreover, the reliance on reductive glutamine metabolism can be recapitulated in normoxia by constitutive HIF1 activation in cells with loss of VHL. The mitochondrial NADPH/NADP+ ratio required to fuel the reductive reaction through IDH2 can arise from the increased NADH/NAD+ ratio existing in the mitochondria under hypoxic conditions (21, 22), with the transfer of electrons from NADH to NADP+ to generate NADPH occurring through the activity of the mitochondrial transhydrogenase (23). Our data do not exclude a complementary role for cytosolic IDH1 in impacting reductive glutamine metabolism, potentially through its oxidative function in an IDH2/IDH1 shuttle that transfers high energy electrons in the form of NADPH from mitochondria to cytosol (16, 24).
In further support of the increased mitochondrial reductive glutamine metabolism that we observe in hypoxia, we report here that incubation in hypoxia can lead to elevated 2HG levels in cells lacking IDH1/2 mutations. 2HG production from glutamine-derived α-ketoglutarate significantly decreased with knockdown of IDH2, supporting the conclusion that 2HG is produced in hypoxia by enhanced reverse flux of α-ketoglutarate through IDH2 in a truncated, noncarboxylating reductive reaction. However, other mechanisms may also contribute to 2HG elevation in hypoxia. These include diminished oxidative activity and/or enhanced reductive activity of the 2HG dehydrogenase, a mitochondrial enzyme that normally functions to oxidize 2HG back to α-ketoglutarate (25). The level of 2HG elevation we observe in hypoxic cells is associated with a concomitant increase in α-ketoglutarate, and is modest relative to that observed in cancers with IDH1/2 gain-of-function mutations. Nonetheless, 2HG elevation resulting from hypoxia in cells with wild-type IDH1/2 may hold promise as a cellular or serum biomarker for tissues undergoing chronic hypoxia and/or excessive glutamine metabolism.
The IDH2-dependent reductive carboxylation pathway that we propose in this report allows for continued citrate production from glutamine carbon when hypoxia and/or HIF1 activation prevents glucose carbon from contributing to citrate synthesis. Moreover, as opposed to continued oxidative TCA cycle functioning in hypoxia which can increase reactive oxygen species (ROS), reductive carboxylation of α-ketoglutarate in the mitochondria may serve as an electron sink that decreases the generation of ROS. HIF1 activity is not limited to the setting of hypoxia, as a common feature of several cancers is the normoxic stabilization of HIF1α through loss of the VHL tumor suppressor or other mechanisms. We demonstrate here that altered glutamine metabolism through a mitochondrial reductive pathway is a central aspect of hypoxic proliferating cell metabolism and HIF1-induced metabolic reprogramming. These findings are relevant for the understanding of numerous constitutive HIF1-expressing malignancies, as well as for populations, such as stem progenitor cells, which frequently proliferate in hypoxic conditions.
7.9.3 Hypoxia-Inducible Factors in Physiology and Medicine
Gregg L. Semenza
Cell. 2012 Feb 3; 148(3): 399–408.
http://dx.doi.org/10.1016%2Fj.cell.2012.01.021
Oxygen homeostasis represents an organizing principle for understanding metazoan evolution, development, physiology, and pathobiology. The hypoxia-inducible factors (HIFs) are transcriptional activators that function as master regulators of oxygen homeostasis in all metazoan species. Rapid progress is being made in elucidating homeostatic roles of HIFs in many physiological systems, determining pathological consequences of HIF dysregulation in chronic diseases, and investigating potential targeting of HIFs for therapeutic purposes. Oxygen homeostasis represents an organizing principle for understanding metazoan evolution, development, physiology, and pathobiology. The hypoxia-inducible factors (HIFs) are transcriptional activators that function as master regulators of oxygen homeostasis in all metazoan species. Rapid progress is being made in elucidating homeostatic roles of HIFs in many physiological systems, determining pathological consequences of HIF dysregulation in chronic diseases, and investigating potential targeting of HIFs for therapeutic purposes.
Oxygen is central to biology because of its utilization in the process of respiration. O2 serves as the final electron acceptor in oxidative phosphorylation, which carries with it the risk of generating reactive oxygen species (ROS) that react with cellular macromolecules and alter their biochemical or physical properties, resulting in cell dysfunction or death. As a consequence, metazoan organisms have evolved elaborate cellular metabolic and systemic physiological systems that are designed to maintain oxygen homeostasis. This review will focus on the role of hypoxia-inducible factors (HIFs) as master regulators of oxygen homeostasis and, in particular, on recent advances in understanding their roles in physiology and medicine. Due to space limitations and the remarkably pleiotropic effects of HIFs, the description of such roles will be illustrative rather than comprehensive.
O2 and Evolution, Part 1
Accumulation of O2 in Earth’s atmosphere starting ~2.5 billion years ago led to evolution of the extraordinarily efficient system of oxidative phosphorylation that transfers chemical energy stored in carbon bonds of organic molecules to the high-energy phosphate bond in ATP, which is used to power physicochemical reactions in living cells. Energy produced by mitochondrial respiration is sufficient to power the development and maintenance of multicellular organisms, which could not be sustained by energy produced by glycolysis alone (Lane and Martin, 2010). The modest dimensions of primitive metazoan species were such that O2 could diffuse from the atmosphere to all of the organism’s thousand cells, as is the case for the worm Caenorhabditis elegans. To escape the constraints placed on organismal growth by diffusion, systems designed to conduct air to cells deep within the body evolved and were sufficient for O2delivery to organisms with hundreds of thousands of cells, such as the fly Drosophila melanogaster. The final leap in body scale occurred in vertebrates and was associated with the evolution of complex respiratory, circulatory, and nervous systems designed to efficiently capture and distribute O2 to hundreds of millions of millions of cells in the case of the adult Homo sapiens.
Hypoxia-Inducible Factors
Hypoxia-inducible factor 1 (HIF-1) is expressed by all extant metazoan species analyzed (Loenarz et al., 2011). HIF-1 consists of HIF-1α and HIF-1β subunits, which each contain basic helix-loop-helix-PAS (bHLH-PAS) domains (Wang et al., 1995) that mediate heterodimerization and DNA binding (Jiang et al., 1996a). HIF-1β heterodimerizes with other bHLH-PAS proteins and is present in excess, such that HIF-1α protein levels determine HIF-1 transcriptional activity (Semenza et al., 1996).
Under well-oxygenated conditions, HIF-1α is bound by the von Hippel-Lindau (VHL) protein, which recruits an ubiquitin ligase that targets HIF-1α for proteasomal degradation (Kaelin and Ratcliffe, 2008). VHL binding is dependent upon hydroxylation of a specific proline residue in HIF-1α by the prolyl hydroxylase PHD2, which uses O2 as a substrate such that its activity is inhibited under hypoxic conditions (Epstein et al., 2001). In the reaction, one oxygen atom is inserted into the prolyl residue and the other atom is inserted into the co-substrate α-ketoglutarate, splitting it into CO2 and succinate (Kaelin and Ratcliffe, 2008). Factor inhibiting HIF-1 (FIH-1) represses HIF-1α transactivation function (Mahon et al., 2001) by hydroxylating an asparaginyl residue, using O2 and α-ketoglutarate as substrates, thereby blocking the association of HIF-1α with the p300 coactivator protein (Lando et al., 2002). Dimethyloxalylglycine (DMOG), a competitive antagonist of α-ketoglutarate, inhibits the hydroxylases and induces HIF-1-dependent transcription (Epstein et al., 2001). HIF-1 activity is also induced by iron chelators (such as desferrioxamine) and cobalt chloride, which inhibit hydroxylases by displacing Fe(II) from the catalytic center (Epstein et al., 2001).
Studies in cultured cells (Jiang et al., 1996b) and isolated, perfused, and ventilated lung preparations (Yu et al., 1998) revealed an exponential increase in HIF-1α levels at O2 concentrations less than 6% (~40 mm Hg), which is not explained by known biochemical properties of the hydroxylases. In most adult tissues, O2concentrations are in the range of 3-5% and any decrease occurs along the steep portion of the dose-response curve, allowing a graded response to hypoxia. Analyses of cultured human cells have revealed that expression of hundreds of genes was increased in response to hypoxia in a HIF-1-dependent manner (as determined by RNA interference) with direct binding of HIF-1 to the gene (as determined by chromatin immunoprecipitation [ChIP] assays); in addition, the expression of hundreds of genes was decreased in response to hypoxia in a HIF-1-dependent manner but binding of HIF-1 to these genes was not detected (Mole et al., 2009), indicating that HIF-dependent repression occurs via indirect mechanisms, which include HIF-1-dependent expression of transcriptional repressors (Yun et al., 2002) and microRNAs (Kulshreshtha et al., 2007). ChIP-seq studies have revealed that only 40% of HIF-1 binding sites are located within 2.5 kb of the transcription start site (Schödel et al., 2011).
In vertebrates, HIF-2α is a HIF-1α paralog that is also regulated by prolyl and asparaginyl hydroxylation and dimerizes with HIF-1β, but is expressed in a cell-restricted manner and plays important roles in erythropoiesis, vascularization, and pulmonary development, as described below. In D. melanogaster, the gene encoding the HIF-1α ortholog is designated similar and its paralog is designated trachealess because inactivating mutations result in defective development of the tracheal tubes (Wilk et al., 1996). In contrast, C. elegans has only a single HIF-1α homolog (Epstein et al., 2001). Thus, in both invertebrates and vertebrates, evolution of specialized systems for O2 delivery was associated with the appearance of a HIF-1α paralog.
O2 and Metabolism
The regulation of metabolism is a principal and primordial function of HIF-1. Under hypoxic conditions, HIF-1 mediates a transition from oxidative to glycolytic metabolism through its regulation of: PDK1, encoding pyruvate dehydrogenase (PDH) kinase 1, which phosphorylates and inactivates PDH, thereby inhibiting the conversion of pyruvate to acetyl coenzyme A for entry into the tricarboxylic acid cycle (Kim et al., 2006; Papandreou et al., 2006); LDHA, encoding lactate dehydrogenase A, which converts pyruvate to lactate (Semenza et al. 1996); and BNIP3 (Zhang et al. 2008) and BNIP3L (Bellot et al., 2009), which mediate selective mitochondrial autophagy (Figure 1). HIF-1 also mediates a subunit switch in cytochrome coxidase that improves the efficiency of electron transfer under hypoxic conditions (Fukuda et al., 2007). An analogous subunit switch is also observed in Saccharomyces cerevisiae, although it is mediated by a completely different mechanism (yeast lack HIF-1), suggesting that it may represent a fundamental response of eukaryotic cells to hypoxia.

Regulation of Glucose Metabolism nihms-350382-f0001
Regulation of Glucose Metabolism
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3437543/bin/nihms-350382-f0001.gif
Figure 1 Regulation of Glucose Metabolism
It is conventional wisdom that cells switch to glycolysis when O2 becomes limiting for mitochondrial ATP production. Yet, HIF-1α-null mouse embryo fibroblasts, which do not down-regulate respiration under hypoxic conditions, have higher ATP levels at 1% O2 than wild-type cells at 20% O2, demonstrating that under these conditions O2 is not limiting for ATP production (Zhang et al., 2008). However, the HIF-1α-null cells die under prolonged hypoxic conditions due to ROS toxicity (Kim et al. 2006; Zhang et al., 2008). These studies have led to a paradigm shift with regard to our understanding of the regulation of cellular metabolism (Semenza, 2011): the purpose of this switch is to prevent excess mitochondrial generation of ROS that would otherwise occur due to the reduced efficiency of electron transfer under hypoxic conditions (Chandel et al., 1998). This may be particularly important in stem cells, in which avoidance of DNA damage is critical (Suda et al., 2011).
Role of HIFs in Development
Much of mammalian embryogenesis occurs at O2 concentrations of 1-5% and O2 functions as a morphogen (through HIFs) in many developmental systems (Dunwoodie, 2009). Mice that are homozygous for a null allele at the locus encoding HIF-1α die by embryonic day 10.5 with cardiac malformations, vascular defects, and impaired erythropoiesis, indicating that all three components of the circulatory system are dependent upon HIF-1 for normal development (Iyer et al., 1998; Yoon et al., 2011). Depending on the genetic background, mice lacking HIF-2α: die by embryonic day 12.5 with vascular defects (Peng et al., 2000) or bradycardia due to deficient catecholamine production (Tian et al., 1998); die as neonates due to impaired lung maturation (Compernolle et al., 2002); or die several months after birth due to ROS-mediated multi-organ failure (Scortegagna et al., 2003). Thus, while vertebrate evolution was associated with concomitant appearance of the circulatory system and HIF-2α, both HIF-1 and HIF-2 have important roles in circulatory system development. Conditional knockout of HIF-1α in specific cell types has demonstrated important roles in chondrogenesis (Schipani et al., 2001), adipogenesis (Yun et al., 2002), B-lymphocyte development (Kojima et al., 2002), osteogenesis (Wang et al., 2007), hematopoiesis (Takubo et al., 2010), T-lymphocyte differentiation (Dang et al., 2011), and innate immunity (Zinkernagel et al., 2007). While knockout mouse experiments point to the adverse effects of HIF-1 loss-of-function on development, it is also possible that increased HIF-1 activity, induced by hypoxia in embryonic tissues as a result of abnormalities in placental blood flow, may also dysregulate development and result in congenital malformations. For example, HIF-1α has been shown to interact with, and stimulate the transcriptional activity of, Notch, which plays a key role in many developmental pathways (Gustafsson et al., 2005).
Translational Prospects
Drug discovery programs have been initiated at many pharmaceutical and biotech companies to develop prolyl hydroxylase inhibitors (PHIs) that, as described above for DMOG, induce HIF activity for treatment of disorders in which HIF mediates protective physiological responses. Local and/or short term induction of HIF activity by PHIs, gene therapy, or other means are likely to be useful novel therapies for many of the diseases described above. In the case of ischemic cardiovascular disease, local therapy is needed to provide homing signals for the recruitment of BMDACs. Chronic systemic use of PHIs must be approached with great caution: individuals with genetic mutations that constitutively activate the HIF pathway (described below) have increased incidence of cardiovascular disease and mortality (Yoon et al., 2011). On the other hand, the profound inhibition of HIF activity and vascular responses to ischemia that are associated with aging suggest that systemic replacement therapy might be contemplated as a preventive measure for subjects in whom impaired HIF responses to hypoxia can be documented. In C. elegans, VHL loss-of-function increases lifespan in a HIF-1-dependent manner (Mehta et al., 2009), providing further evidence for a mutually antagonistic relationship between HIF-1 and aging.
Cancer
Cancers contain hypoxic regions as a result of high rates of cell proliferation coupled with the formation of vasculature that is structurally and functionally abnormal. Increased HIF-1α and/or HIF-2α levels in diagnostic tumor biopsies are associated with increased risk of mortality in cancers of the bladder, brain, breast, colon, cervix, endometrium, head/neck, lung, ovary, pancreas, prostate, rectum, and stomach; these results are complemented by experimental studies, which demonstrate that genetic manipulations that increase HIF-1α expression result in increased tumor growth, whereas loss of HIF activity results in decreased tumor growth (Semenza, 2010). HIFs are also activated by genetic alterations, most notably, VHL loss of function in clear cell renal carcinoma (Majmunder et al., 2010). HIFs activate transcription of genes that play key roles in critical aspects of cancer biology, including stem cell maintenance (Wang et al., 2011), cell immortalization, epithelial-mesenchymal transition (Mak et al., 2010), genetic instability (Huang et al., 2007), vascularization (Liao and Johnson, 2007), glucose metabolism (Luo et al., 2011), pH regulation (Swietach et al., 2007), immune evasion (Lukashev et al., 2007), invasion and metastasis (Chan and Giaccia, 2007), and radiation resistance (Moeller et al., 2007). Given the extensive validation of HIF-1 as a potential therapeutic target, drugs that inhibit HIF-1 have been identified and shown to have anti-cancer effects in xenograft models (Table 1; Semenza, 2010).
Table 1 Drugs that Inhibit HIF-1
| Process Inhibited |
Drug Class |
Prototype |
| HIF-1 α synthesis |
Cardiac glycosidemTOR inhibitorMicrotubule targeting agent
Topoisomerase I inhibitor |
DigoxinRapamycin2-Methoxyestradiol
Topotecan |
|
| HIF-1 α protein stability |
HDAC inhibitorHSP90 inhibitorCalcineurin inhibitor
Guanylate cyclase activator |
LAQ82417-AAGCyclosporine
YC-1 |
|
| Heterodimerization |
Antimicrobial agent |
Acriflavine |
| DNA binding |
AnthracyclineQuinoxaline antibiotic |
DoxorubicinEchinomycin |
|
| Transactivation |
Proteasome inhibitorAntifungal agent |
BortezomibAmphotericin B |
|
| Signal transduction |
BCR-ABL inhibitorCyclooxygenase inhibitorEGFR inhibitor
HER2 inhibitor |
ImatinibIbuprofenErlotinib, Gefitinib
Trastuzumab |
|
Over 100 women die every day of breast cancer in the U.S. The mean PO2 is 10 mm Hg in breast cancer as compared to > 60 mm Hg in normal breast tissue and cancers with PO2 < 10 mm Hg are associated with increased risk of metastasis and patient mortality (Vaupel et al., 2004). Increased HIF-1α protein levels, as identified by immunohistochemical analysis of tumor biopsies, are associated with increased risk of metastasis and/or patient mortality in unselected breast cancer patients and in lymph node-positive, lymph node-negative, HER2+, or estrogen receptor+ subpopulations (Semenza, 2011). Metastasis is responsible for > 90% of breast cancer mortality. The requirement for HIF-1 in breast cancer metastasis has been demonstrated for both autochthonous tumors in transgenic mice (Liao et al., 2007) and orthotopic transplants in immunodeficient mice (Zhang et al., 2011; Wong et al., 2011). Primary tumors direct the recruitment of bone marrow-derived cells to the lungs and other sites of metastasis (Kaplan et al., 2005). In breast cancer, hypoxia induces the expression of lysyl oxidase (LOX), a secreted protein that remodels collagen at sites of metastatic niche formation (Erler et al., 2009). In addition to LOX, breast cancers also express LOX-like proteins 2 and 4. LOX, LOXL2, and LOXL4 are all HIF-1-regulated genes and HIF-1 inhibition blocks metastatic niche formation regardless of which LOX/LOXL protein is expressed, whereas available LOX inhibitors are not effective against all LOXL proteins (Wong et al., 2011), again illustrating the role of HIF-1 as a master regulator that controls the expression of multiple genes involved in a single (patho)physiological process.
Translational Prospects
Small molecule inhibitors of HIF activity that have anti-cancer effects in mouse models have been identified (Table 1). Inhibition of HIF impairs both vascular and metabolic adaptations to hypoxia, which may decrease O2 delivery and increase O2 utilization. These drugs are likely to be useful (as components of multidrug regimens) in the treatment of a subset of cancer patients in whom high HIF activity is driving progression. As with all novel cancer therapeutics, successful translation will require the development of methods for identifying the appropriate patient cohort. Effects of combination drug therapy also need to be considered. VEGF receptor tyrosine kinase inhibitors, which induce tumor hypoxia by blocking vascularization, have been reported to increase metastasis in mouse models (Ebos et al., 2009), which may be mediated by HIF-1; if so, combined use of HIF-1 inhibitors with these drugs may prevent unintended counter-therapeutic effects.
HIF inhibitors may also be useful in the treatment of other diseases in which dysregulated HIF activity is pathogenic. Proof of principle has been established in mouse models of ocular neovascularization, a major cause of blindness in the developed world, in which systemic or intraocular injection of the HIF-1 inhibitor digoxin is therapeutic (Yoshida et al., 2010). Systemic administration of HIF inhibitors for cancer therapy would be contraindicated in patients who also have ischemic cardiovascular disease, in which HIF activity is protective. The analysis of SNPs at the HIF1A locus described above suggests that the population may include HIF hypo-responders, who are at increased risk of severe ischemic cardiovascular disease. It is also possible that HIF hyper-responders, such as individuals with hereditary erythrocytosis, are at increased risk of particularly aggressive cancer.
O2 and Evolution, Part 2
When lowlanders sojourn to high altitude, hypobaric hypoxia induces erythropoiesis, which is a relatively ineffective response because the problem is not insufficient red cells, but rather insufficient ambient O2. Chronic erythrocytosis increases the risk of heart attack, stroke, and fetal loss during pregnancy. Many high-altitude Tibetans maintain the same hemoglobin concentration as lowlanders and yet, despite severe hypoxemia, they also maintain aerobic metabolism. The basis for this remarkable evolutionary adaptation appears to have involved the selection of genetic variants at multiple loci encoding components of the oxygen sensing system, particularly HIF-2α (Beall et al., 2010; Simonson et al., 2010; Yi et al., 2010). Given that hereditary erythrocytosis is associated with modest HIF-2α gain-of-function, the Tibetan genotype associated with absence of an erythrocytotic response to hypoxia may encode reduced HIF-2α activity along with other alterations that increase metabolic efficiency. Delineating the molecular mechanisms underlying these metabolic adaptations may lead to novel therapies for ischemic disorders, illustrating the importance of oxygen homeostasis as a nexus where evolution, biology, and medicine converge.
7.9.4 Hypoxia-inducible factor 1. Regulator of mitochondrial metabolism and mediator of ischemic preconditioning
Semenza GL1.
Biochim Biophys Acta. 2011 Jul; 1813(7):1263-8.
http://dx.doi.org/10.1016%2Fj.bbamcr.2010.08.006
Hypoxia-inducible factor 1 (HIF-1) mediates adaptive responses to reduced oxygen availability by regulating gene expression. A critical cell-autonomous adaptive response to chronic hypoxia controlled by HIF-1 is reduced mitochondrial mass and/or metabolism. Exposure of HIF-1-deficient fibroblasts to chronic hypoxia results in cell death due to excessive levels of reactive oxygen species (ROS). HIF-1 reduces ROS production under hypoxic conditions by multiple mechanisms including: a subunit switch in cytochrome c oxidase from the COX4-1 to COX4-2 regulatory subunit that increases the efficiency of complex IV; induction of pyruvate dehydrogenase kinase 1, which shunts pyruvate away from the mitochondria; induction of BNIP3, which triggers mitochondrial selective autophagy; and induction of microRNA-210, which blocks assembly of Fe/S clusters that are required for oxidative phosphorylation. HIF-1 is also required for ischemic preconditioning and this effect may be due in part to its induction of CD73, the enzyme that produces adenosine. HIF-1-dependent regulation of mitochondrial metabolism may also contribute to the protective effects of ischemic preconditioning.
The story of life on Earth is a tale of oxygen production and utilization. Approximately 3 billion years ago, primitive single-celled organisms evolved the capacity for photosynthesis, a biochemical process in which photons of solar energy are captured by chlorophyll and used to power the reaction of CO2 and H2O to form glucose and O2. The subsequent rise in the atmospheric O2 concentration over the next billion years set the stage for the ascendance of organisms with the capacity for respiration, a process that consumes glucose and O2 and generates CO2, H2O, and energy in the form of ATP. Some of these single-celled organisms eventually took up residence within the cytoplasm of other cells and devoted all of their effort to energy production as mitochondria. Compared to the conversion of glucose to lactate by glycolysis, the complete oxidation of glucose by respiration provided such a large increase in energy production that it made possible the evolution of multicellular organisms. Among metazoan organisms, the progressive increase in body size during evolution was accompanied by progressively more complex anatomic structures that function to ensure the adequate delivery of O2 to all cells, ultimately resulting in the sophisticated circulatory and respiratory systems of vertebrates.
All metazoan cells can sense and respond to reduced O2 availability (hypoxia). Adaptive responses to hypoxia can be cell autonomous, such as the alterations in mitochondrial metabolism that are described below, or non-cell-autonomous, such as changes in tissue vascularization (reviewed in ref. 1). Primary responses to hypoxia need to be distinguished from secondary responses to sequelae of hypoxia, such as the adaptive responses to ATP depletion that are mediated by AMP kinase (reviewed in ref 2). In contrast, recent data suggest that O2 and redox homeostasis are inextricably linked and that changes in oxygenation are inevitably associated with changes in the levels of reactive oxygen species (ROS), as will be discussed below.
HIF-1 Regulates Oxygen Homeostasis in All Metazoan Species
A key regulator of the developmental and physiological networks required for the maintenance of O2homeostasis is hypoxia-inducible factor 1 (HIF-1). HIF-1 is a heterodimeric transcription factor that is composed of an O2-regulated HIF-1α subunit and a constitutively expressed HIF-1β subunit [3,4]. HIF-1 regulates the expression of hundreds of genes through several major mechanisms. First, HIF-1 binds directly to hypoxia response elements, which are cis-acting DNA sequences located within target genes [5]. The binding of HIF-1 results in the recruitment of co-activator proteins that activate gene transcription (Fig. 1A). Only rarely does HIF-1 binding result in transcriptional repression [6]. Instead, HIF-1 represses gene expression by indirect mechanisms, which are described below. Second, among the genes activated by HIF-1 are many that encode transcription factors [7], which when synthesized can bind to and regulate (either positively or negatively) secondary batteries of target genes (Fig. 1B). Third, another group of HIF-1 target genes encode members of the Jumonji domain family of histone demethylases [8,9], which regulate gene expression by modifying chromatin structure (Fig. 1C). Fourth, HIF-1 can activate the transcription of genes encoding microRNAs [10], which bind to specific mRNA molecules and either block their translation or mediate their degradation (Fig. 1D). Fifth, the isolated HIF-1α subunit can bind to other transcription factors [11,12] and inhibit (Fig. 1E) or potentiate (Fig. 1F) their activity.

Mechanisms by which HIF-1 regulates gene expression. nihms232046f1
Mechanisms by which HIF-1 regulates gene expression.
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3010308/bin/nihms232046f1.gif
Fig. 1 Mechanisms by which HIF-1 regulates gene expression. (A) Top: HIF-1 binds directly to target genes at a cis-acting hypoxia response element (HRE) and recruits coactivator proteins such as p300 to increase gene transcription.
HIF-1α and HIF-1β are present in all metazoan species, including the simple roundworm Caenorhabitis elegans [13], which consists of ~103 cells and has no specialized systems for O2 delivery. The fruit flyDrosophila melanogaster evolved tracheal tubes, which conduct air into the interior of the body from which it diffuses to surrounding cells. In vertebrates, the development of the circulatory and respiratory systems was accompanied by the appearance of HIF-2α, which is also O2-regulated and heterodimerizes with HIF-1β [14] but is only expressed in a restricted number of cell types [15], whereas HIF-1α and HIF-1β are expressed in all human and mouse tissues [16]. In Drosophila, the ubiquitiously expressed HIF-1α ortholog is designatedSimilar [17] and the paralogous gene that is expressed specifically in tracheal tubes is designated Trachealess[18].
HIF-1 Activity is Regulated by Oxygen
In the presence of O2, HIF-1α and HIF-2α are subjected to hydroxylation by prolyl-4-hydroxylase domain proteins (PHDs) that use O2 and α-ketoglutarate as substrates and generate CO2 and succinate as by-products [19]. Prolyl hydroxylation is required for binding of the von Hipple-Lindau protein, which recruits a ubiquitin-protein ligase that targets HIF-1α and HIF-2α for proteasomal degradation (Fig. 2). Under hypoxic conditions, the rate of hydroxylation declines and the non-hydroxylated proteins accumulate. HIF-1α transactivation domain function is also O2-regulated [20,21]. Factor inhibiting HIF-1 (FIH-1) represses transactivation domain function [22] by hydroxylating asparagine residue 803 in HIF-1α, thereby blocking the binding of the co-activators p300 and CBP [23].

Negative regulation of HIF-1 activity by oxygen nihms232046f2
Negative regulation of HIF-1 activity by oxygen
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3010308/bin/nihms232046f2.gif
Fig. 2 Negative regulation of HIF-1 activity by oxygen. Top: In the presence of O2: prolyl hydroxylation of HIF-1a leads to binding of the von Hippel-Lindau protein (VHL), which recruits a ubiquitin protein-ligase that targets HIF-1a for proteasomal degradation;
When cells are acutely exposed to hypoxic conditions, the generation of ROS at complex III of the mitochondrial electron transport chain (ETC) increases and is required for the induction of HIF-1α protein levels [24]. More than a decade after these observations were first made, the precise mechanism by which hypoxia increases ROS generation and by which ROS induces HIF-1α accumulation remain unknown. However, the prolyl and asparaginyl hydroxylases contain Fe2+ in their active site and oxidation to Fe3+would block their catalytic activity. Since O2 is a substrate for the hydroxylation reaction, anoxia also results in a loss of enzyme activity. However, the concentration at which O2 becomes limiting for prolyl or asparaginyl hydroxylase activity in vivo is not known.
HIF-1 Regulates the Balance Between Oxidative and Glycolytic Metabolism
All metazoan organisms depend on mitochondrial respiration as the primary mechanism for generating sufficient amounts of ATP to maintain cellular and systemic homeostasis. Respiration, in turn, is dependent on an adequate supply of O2 to serve as the final electron acceptor in the ETC. In this process, electrons are transferred from complex I (or complex II) to complex III, then to complex IV, and finally to O2, which is reduced to water. This orderly transfer of electrons generates a proton gradient across the inner mitochondrial membrane that is used to drive the synthesis of ATP. At each step of this process, some electrons combine with O2 prematurely, resulting in the production of superoxide anion, which is reduced to hydrogen peroxide through the activity of mitochondrial superoxide dismutase. The efficiency of electron transport appears to be optimized to the physiological range of O2 concentrations, such that ATP is produced without the production of excess superoxide, hydrogen peroxide, and other ROS at levels that would result in the increased oxidation of cellular macromolecules and subsequent cellular dysfunction or death. In contrast, when O2levels are acutely increased or decreased, an imbalance between O2 and electron flow occurs, which results in increased ROS production.
MEFs require HIF-1 activity to make two critical metabolic adaptations to chronic hypoxia. First, HIF-1 activates the gene encoding pyruvate dehydrogenase (PDH) kinase 1 (PDK1), which phosphorylates and inactivates the catalytic subunit of PDH, the enzyme that converts pyruvate to acetyl coenzyme A (AcCoA) for entry into the mitochondrial tricarboxylic acid (TCA) cycle [25]. Second, HIF-1 activates the gene encoding BNIP3, a member of the Bcl-2 family of mitochondrial proteins, which triggers selective mitochondrial autophagy [26]. Interference with the induction of either of these proteins in hypoxic cells results in increased ROS production and increased cell death. Overexpression of either PDK1 or BNIP3 rescues HIF-1α-null MEFs. By shunting pyruvate away from the mitochondria, PDK1 decreases flux through the ETC and thereby counteracts the reduced efficiency of electron transport under hypoxic conditions, which would otherwise increase ROS production. PDK1 functions cooperatively with the product of another HIF-1 target gene, LDHA [27], which converts pyruvate to lactate, thereby further reducing available substrate for the PDH reaction.
PDK1 effectively reduces flux through the TCA cycle and thereby reduces flux through the ETC in cells that primarily utilize glucose as a substrate for oxidative phosphorylation. However, PDK1 is predicted to have little effect on ROS generation in cells that utilize fatty acid oxidation as their source of AcCoA. Hence another strategy to reduce ROS generation under hypoxic conditions is selective mitochondrial autophagy [26]. MEFs reduce their mitochondrial mass and O2 consumption by >50% after only two days at 1% O2. BNIP3 competes with Beclin-1 for binding to Bcl-2, thereby freeing Beclin-1 to activate autophagy. Using short hairpin RNAs to knockdown expression of BNIP3, Beclin-1, or Atg5 (another component of the autophagy machinery) phenocopied HIF-1α-null cells by preventing hypoxia-induced reductions in mitochondrial mass and O2 consumption as a result of failure to induce autophagy [26]. HIF-1-regulated expression of BNIP3L also contributes to hypoxia-induced autophagy [28]. Remarkably, mice heterozygous for the HIF-1α KO allele have a significantly increased ratio of mitochondrial:nuclear DNA in their lungs (even though this is the organ that is exposed to the highest O2 concentrations), indicating that HIF-1 regulates mitochondrial mass under physiological conditions in vivo [26]. In contrast to the selective mitochondrial autophagy that is induced in response to hypoxia as described above, autophagy (of unspecified cellular components) induced by anoxia does not require HIF-1, BNIP3, or BNIP3L, but is instead regulated by AMP kinase [29].
The multiplicity of HIF-1-mediated mechanisms identified so far by which cells regulate mitochondrial metabolism in response to changes in cellular O2 concentration (Fig. 3) suggests that this is a critical adaptive response to hypoxia. The fundamental nature of this physiological response is underscored by the fact that yeast also switch COX4 subunits in an O2-dependent manner but do so by an entirely different molecular mechanism [33], since yeast do not have a HIF-1α homologue. Thus, it appears that by convergent evolution both unicellular and multicellular eukaryotes possess mechanisms by which they modulate mitochondrial metabolism to maintain redox homeostasis despite changes in O2 availability. Indeed, it is the balance between energy, oxygen, and redox homeostasis that represents the key to life with oxygen.

Regulation of mitochondrial metabolism by HIF-1 nihms232046f3
Regulation of mitochondrial metabolism by HIF-1α
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3010308/bin/nihms232046f3.gif
Fig. 3 Regulation of mitochondrial metabolism by HIF-1α. Acute hypoxia leads to increased mitochondrial generation of reactive oxygen species (ROS). Decreased O2 and increased ROS levels lead to decreased HIF-1α hydroxylation (see Fig. 2) and increased HIF-1-dependent …
7.9.5 Regulation of cancer cell metabolism by hypoxia-inducible factor 1
Semenza GL1.
Semin Cancer Biol. 2009 Feb; 19(1):12-6.
The Warburg Effect: The Re-discovery of the Importance of Aerobic Glycolysis in Tumor Cells
http://dx.doi.org:/10.1016/j.semcancer.2008.11.009
The induction of hypoxia-inducible factor 1 (HIF-1) activity, either as a result of intratumoral hypoxia or loss-of-function mutations in the VHL gene, leads to a dramatic reprogramming of cancer cell metabolism involving increased glucose transport into the cell, increased conversion of glucose to pyruvate, and a concomitant decrease in mitochondrial metabolism and mitochondrial mass. Blocking these adaptive metabolic responses to hypoxia leads to cell death due to toxic levels of reactive oxygen species. Targeting HIF-1 or metabolic enzymes encoded by HIF-1 target genes may represent a novel therapeutic approach to cancer.
http://ars.els-cdn.com/content/image/1-s2.0-S1044579X08001065-gr1.sml
http://ars.els-cdn.com/content/image/1-s2.0-S1044579X08001065-gr2.sml
7.9.6 Coming up for air. HIF-1 and mitochondrial oxygen consumption
Simon MC1.
Cell Metab. 2006 Mar;3(3):150-1.
http://dx.doi.org/10.1016/j.cmet.2006.02.007
Hypoxic cells induce glycolytic enzymes; this HIF-1-mediated metabolic adaptation increases glucose flux to pyruvate and produces glycolytic ATP. Two papers in this issue of Cell Metabolism (Kim et al., 2006; Papandreou et al., 2006) demonstrate that HIF-1 also influences mitochondrial function, suppressing both the TCA cycle and respiration by inducing pyruvate dehydrogenase kinase 1 (PDK1). PDK1 regulation in hypoxic cells promotes cell survival.
Comment on
Oxygen deprivation (hypoxia) occurs in tissues when O2 supply via the cardiovascular system fails to meet the demand of O2-consuming cells. Hypoxia occurs naturally in physiological settings (e.g., embryonic development and exercising muscle), as well as in pathophysiological conditions (e.g., myocardial infarction, inflammation, and solid tumor formation). For over a century, it has been appreciated that O2-deprived cells exhibit increased conversion of glucose to lactate (the “Pasteur effect”). Activation of the Pasteur effect during hypoxia in mammalian cells is facilitated by HIF-1, which mediates the upregulation of glycolytic enzymes that support an increase in glycolytic ATP production as mitochondria become starved for O2, the substrate for oxidative phosphorylation (Seagroves et al., 2001). Thus, mitochondrial respiration passively decreases due to O2 depletion in hypoxic tissues. However, reports by Kim et al. (2006) and Papandreou et al. (2006) in this issue of Cell Metabolism demonstrate that this critical metabolic adaptation is more complex and includes an active suppression of mitochondrial pyruvate catabolism and O2consumption by HIF-1.
Mitochondrial oxidative phosphorylation is regulated by multiple mechanisms, including substrate availability. Major substrates include O2 (the terminal electron acceptor) and pyruvate (the primary carbon source). Pyruvate, as the end product of glycolysis, is converted to acetyl-CoA by the pyruvate dehydrogenase enzymatic complex and enters the tricarboxylic acid (TCA) cycle. Pyruvate conversion into acetyl-CoA is irreversible; this therefore represents an important regulatory point in cellular energy metabolism. Pyruvate dehydrogenase kinase (PDK) inhibits pyruvate dehydrogenase activity by phosphorylating its E1 subunit (Sugden and Holness, 2003). In the manuscripts by Kim et al. (2006) and Papandreou et al. (2006), the authors find that PDK1 is a HIF-1 target gene that actively regulates mitochondrial respiration by limiting pyruvate entry into the TCA cycle. By excluding pyruvate from mitochondrial metabolism, hypoxic cells accumulate pyruvate, which is then converted into lactate via lactate dehydrogenase (LDH), another HIF-1-regulated enzyme. Lactate in turn is released into the extracellular space, regenerating NAD+ for continued glycolysis by O2-starved cells (see Figure 1). This HIF-1-dependent block to mitochondrial O2 consumption promotes cell survival, especially when O2 deprivation is severe and prolonged.

multiple-hypoxia-induced-cellular-metabolic-changes-are-regulated-by-hif-1
http://ars.els-cdn.com/content/image/1-s2.0-S1550413106000672-gr1.jpg
Figure 1. Multiple hypoxia-induced cellular metabolic changes are regulated by HIF-1
By stimulating the expression of glucose transporters and glycolytic enzymes, HIF-1 promotes glycolysis to generate increased levels of pyruvate. In addition, HIF-1 promotes pyruvate reduction to lactate by activating lactate dehydrogenase (LDH). Pyruvate reduction to lactate regenerates NAD+, which permits continued glycolysis and ATP production by hypoxic cells. Furthermore, HIF-1 induces pyruvate dehydrogenase kinase 1 (PDK1), which inhibits pyruvate dehydrogenase and blocks conversion of pyruvate to acetyl CoA, resulting in decreased flux through the tricarboxylic acid (TCA) cycle. Decreased TCA cycle activity results in attenuation of oxidative phosphorylation and excessive mitochondrial reactive oxygen species (ROS) production. Because hypoxic cells already exhibit increased ROS, which have been shown to promote HIF-1 accumulation, the induction of PDK1 prevents the persistence of potentially harmful ROS levels.
Papandreou et al. demonstrate that hypoxic regulation of PDK has important implications for antitumor therapies. Recent interest has focused on cytotoxins that target hypoxic cells in tumor microenvironments, such as the drug tirapazamine (TPZ). Because intracellular O2 concentrations are decreased by mitochondrial O2 consumption, HIF-1 could protect tumor cells from TPZ-mediated cell death by maintaining intracellular O2 levels. Indeed, Papandreou et al. show that HIF-1-deficient cells grown at 2% O2 exhibit increased sensitivity to TPZ relative to wild-type cells, presumably due to higher rates of mitochondrial O2 consumption. HIF-1 inhibition in hypoxic tumor cells should have multiple therapeutic benefits, but the use of HIF-1 inhibitors in conjunction with other treatments has to be carefully evaluated for the most effective combination and sequence of drug delivery. One result of HIF-1 inhibition would be a relative decrease in intracellular O2 levels, making hypoxic cytotoxins such as TPZ more potent antitumor agents. Because PDK expression has been detected in multiple human tumor samples and appears to be induced by hypoxia (Koukourakis et al., 2005), small molecule inhibitors of HIF-1 combined with TPZ represent an attractive therapeutic approach for future clinical studies.
Hypoxic regulation of PDK1 has other important implications for cell survival during O2 depletion. Because the TCA cycle is coupled to electron transport, Kim et al. suggest that induction of the pyruvate dehydrogenase complex by PDK1 attenuates not only mitochondrial respiration but also the production of mitochondrial reactive oxygen species (ROS) in hypoxic cells. ROS are a byproduct of electron transfer to O2, and cells cultured at 1 to 5% O2 generate increased mitochondrial ROS relative to those cultured at 21% O2 (Chandel et al., 1998 and Guzy et al., 2005). In fact, hypoxia-induced mitochondrial ROS have also been shown to be necessary for the stabilization of HIF-1 in hypoxic cells (Brunelle et al., 2005, Guzy et al., 2005 and Mansfield et al., 2005). However, the persistence of ROS could ultimately be lethal to tissues during chronic O2 deprivation, and PDK1 induction by HIF-1 should promote cell viability during long-term hypoxia. Kim et al. present evidence that HIF-1-deficient cells exhibit increased apoptosis after 72 hr of culture at 0.5% O2 compared to wild-type cells and that cell survival is rescued by enforced expression of exogenous PDK1. Furthermore, PDK1 reduces ROS production by the HIF-1 null cells. These findings support a novel prosurvival dimension of cellular hypoxic adaptation where PDK1 inhibits the TCA cycle, mitochondrial respiration, and chronic ROS production.
The HIF-1-mediated block to mitochondrial O2 consumption via PDK1 regulation also has implications for O2-sensing pathways by hypoxic cells. One school of thought suggests that perturbing mitochondrial O2consumption increases intracellular O2 concentrations and suppresses HIF-1 induction by promoting the activity of HIF prolyl hydroxylases, the O2-dependent enzymes that regulate HIF-1 stability (Hagen et al., 2003 and Doege et al., 2005). This model suggests that mitochondria function as “O2 sinks.” Although Papandreou et al. demonstrate that increased mitochondrial respiration due to PDK1 depletion results in decreased intracellular O2 levels (based on pimonidazole staining), these changes failed to reduce HIF-1 levels in hypoxic cells. Another model for hypoxic activation of HIF-1 describes a critical role for mitochondrial ROS in prolyl hydroxylase inhibition and HIF-1 stabilization in O2-starved cells (Brunelle et al., 2005, Guzy et al., 2005 and Mansfield et al., 2005) (see Figure 1). The mitochondrial “O2 sink” hypothesis can account for some observations in the literature but fails to explain the inhibition of HIF-1 stabilization by ROS scavengers (Chandel et al., 1998, Brunelle et al., 2005, Guzy et al., 2005 and Sanjuán-Pla et al., 2005). While the relationship between HIF-1 stability, mitochondrial metabolism, ROS, and intracellular O2 redistribution will continue to be debated for some time, these most recent findings shed new light on findings by Louis Pasteur over a century ago.
Selected reading
Brunelle et al., 2005
J.K. Brunelle, E.L. Bell, N.M. Quesada, K. Vercauteren, V. Tiranti, M. Zeviani, R.C. Scarpulla, N.S. Chandel
Cell Metab., 1 (2005), pp. 409–414
Article PDF (324 K) View Record in Scopus Citing articles (357)
Chandel et al., 1998
N.S. Chandel, E. Maltepe, E. Goldwasser, C.E. Mathieu, M.C. Simon, P.T. Schumacker
Proc. Natl. Acad. Sci. USA, 95 (1998), pp. 11715–11720
View Record in Scopus Full Text via CrossRef Citing articles (973)
Doege et al., 2005Doege, S. Heine, I. Jensen, W. Jelkmann, E. Metzen
Blood, 106 (2005), pp. 2311–2317
View Record in Scopus Full Text via CrossRef Citing articles (84)
Guzy et al., 2005
R.D. Guzy, B. Hoyos, E. Robin, H. Chen, L. Liu, K.D. Mansfield, M.C. Simon, U. Hammerling, P.T. Schumacker
Cell Metab., 1 (2005), pp. 401–408
Article PDF (510 K) View Record in Scopus Citing articles (593)
Hagen et al., 2003
Hagen, C.T. Taylor, F. Lam, S. Moncada
Science, 302 (2003), pp. 1975–1978
View Record in Scopus Full Text via CrossRef Citing articles (450)
7.9.7 HIF-1 mediates adaptation to hypoxia by actively downregulating mitochondrial oxygen consumption
Papandreou I1, Cairns RA, Fontana L, Lim AL, Denko NC.
Cell Metab. 2006 Mar; 3(3):187-97.
http://dx.doi.org/10.1016/j.cmet.2006.01.012
The HIF-1 transcription factor drives hypoxic gene expression changes that are thought to be adaptive for cells exposed to a reduced-oxygen environment. For example, HIF-1 induces the expression of glycolytic genes. It is presumed that increased glycolysis is necessary to produce energy when low oxygen will not support oxidative phosphorylation at the mitochondria. However, we find that while HIF-1 stimulates glycolysis, it also actively represses mitochondrial function and oxygen consumption by inducing pyruvate dehydrogenase kinase 1 (PDK1). PDK1 phosphorylates and inhibits pyruvate dehydrogenase from using pyruvate to fuel the mitochondrial TCA cycle. This causes a drop in mitochondrial oxygen consumption and results in a relative increase in intracellular oxygen tension. We show by genetic means that HIF-1-dependent block to oxygen utilization results in increased oxygen availability, decreased cell death when total oxygen is limiting, and reduced cell death in response to the hypoxic cytotoxin tirapazamine.
Comment in
Tissue hypoxia results when supply of oxygen from the bloodstream does not meet demand from the cells in the tissue. Such a supply-demand mismatch can occur in physiologic conditions such as the exercising muscle, in the pathologic condition such as the ischemic heart, or in the tumor microenvironment (Hockel and Vaupel, 2001 and Semenza, 2004). In either the physiologic circumstance or pathologic conditions, there is a molecular response from the cell in which a program of gene expression changes is initiated by the hypoxia-inducible factor-1 (HIF-1) transcription factor. This program of gene expression changes is thought to help the cells adapt to the stressful environment. For example, HIF-1-dependent expression of erythropoietin and angiogenic compounds results in increased blood vessel formation for delivery of a richer supply of oxygenated blood to the hypoxic tissue. Additionally, HIF-1 induction of glycolytic enzymes allows for production of energy when the mitochondria are starved of oxygen as a substrate for oxidative phosphorylation. We now find that this metabolic adaptation is more complex, with HIF-1 not only regulating the supply of oxygen from the bloodstream, but also actively regulating the oxygen demand of the tissue by reducing the activity of the major cellular consumer of oxygen, the mitochondria.
Perhaps the best-studied example of chronic hypoxia is the hypoxia associated with the tumor microenvironment (Brown and Giaccia, 1998). The tumor suffers from poor oxygen supply through a chaotic jumble of blood vessels that are unable to adequately perfuse the tumor cells. The oxygen tension within the tumor is also a function of the demand within the tissue, with oxygen consumption influencing the extent of tumor hypoxia (Gulledge and Dewhirst, 1996 and Papandreou et al., 2005b). The net result is that a large fraction of the tumor cells are hypoxic. Oxygen tensions within the tumor range from near normal at the capillary wall, to near zero in the perinecrotic regions. This perfusion-limited hypoxia is a potent microenvironmental stress during tumor evolution (Graeber et al., 1996 and Hockel and Vaupel, 2001) and an important variable capable of predicting for poor patient outcome. (Brizel et al., 1996, Cairns and Hill, 2004, Hockel et al., 1996 and Nordsmark and Overgaard, 2004).
The HIF-1 transcription factor was first identified based on its ability to activate the erythropoetin gene in response to hypoxia (Wang and Semenza, 1993). Since then, it is has been shown to be activated by hypoxia in many cells and tissues, where it can induce hypoxia-responsive target genes such as VEGF and Glut1 (Airley et al., 2001 and Kimura et al., 2004). The connection between HIF-regulation and human cancer was directly linked when it was discovered that the VHL tumor suppressor gene was part of the molecular complex responsible for the oxic degradation of HIF-1α (Maxwell et al., 1999). In normoxia, a family of prolyl hydroxylase enzymes uses molecular oxygen as a substrate and modifies HIF-1α and HIF2α by hydroxylation of prolines 564 and 402 (Bruick and McKnight, 2001 and Epstein et al., 2001). VHL then recognizes the modified HIF-α proteins, acts as an E3-type of ubiquitin ligase, and along with elongins B and C is responsible for the polyubiquitination of HIF-αs and their proteosomal degradation (Bruick and McKnight, 2001, Chan et al., 2002, Ivan et al., 2001 and Jaakkola et al., 2001). Mutations in VHL lead to constitutive HIF-1 gene expression, and predispose humans to cancer. The ability to recognize modified HIF-αs is at least partly responsible for VHL activity as a tumor suppressor, as introduction of nondegradable HIF-2α is capable of overcoming the growth–inhibitory activity of wild-type (wt) VHL in renal cancer cells (Kondo et al., 2003).
Mitochondrial function can be regulated by PDK1 expression. Mitochondrial oxidative phosphorylation (OXPHOS) is regulated by several mechanisms, including substrate availability (Brown, 1992). The major substrates for OXPHOS are oxygen, which is the terminal electron acceptor, and pyruvate, which is the primary carbon source. Pyruvate is the end product of glycolysis and is converted to acetyl-CoA through the activity of the pyruvate dehydrogenase complex of enzymes. The acetyl-CoA then directly enters the TCA cycle at citrate synthase where it is combined with oxaloacetate to generate citrate. In metazoans, the conversion of pyruvate to acetyl-CoA is irreversible and therefore represents a critical regulatory point in cellular energy metabolism. Pyruvate dehydrogenase is regulated by three known mechanisms: it is inhibited by acetyl-CoA and NADH, it is stimulated by reduced energy in the cell, and it is inhibited by regulatory phosphorylation of its E1 subunit by pyruvate dehydrogenase kinase (PDK) (Holness and Sugden, 2003 and Sugden and Holness, 2003). There are four members of the PDK family in vertebrates, each with specific tissue distributions (Roche et al., 2001). PDK expression has been observed in human tumor biopsies (Koukourakis et al., 2005), and we have reported that PDK3 is hypoxia-inducible in some cell types (Denko et al., 2003). In this manuscript, we find that PDK1 is also a hypoxia-responsive protein that actively regulates the function of the mitochondria under hypoxic conditions by reducing pyruvate entry into the TCA cycle. By excluding pyruvate from mitochondrial consumption, PDK1 induction may increase the conversion of pyruvate to lactate, which is in turn shunted to the extracellular space, regenerating NAD for continued glycolysis.
Identification of HIF-dependent mitochondrial proteins through genomic and bioinformatics approaches
In order to help elucidate the role of HIF-1α in regulating metabolism, we undertook a genomic search for genes that were regulated by HIF-1 in tumor cells exposed to hypoxia in vitro. We used genetically matched human RCC4 cells that had lost VHL during tumorigenesis and displayed constitutive HIF-1 activity, and a cell line engineered to re-express VHL to establish hypoxia-dependent HIF activation. These cells were treated with 18 hr of stringent hypoxia (<0.01% oxygen), and microarray analysis performed. Using a strict 2.5-fold elevation as our cutoff, we identified 173 genes that were regulated by hypoxia and/or VHL status (Table S1 in the Supplemental Data available with this article online). We used the pattern of expression in these experiments to identify putative HIF-regulated genes—ones that were constitutively elevated in the parent RCC4s independent of hypoxia, downregulated in the RCC4VHL cells under normoxia, and elevated in response to hypoxia. Of the 173 hypoxia and VHL-regulated genes, 74 fit the putative HIF-1 target pattern. The open reading frames of these genes were run through a pair of bioinformatics engines in order to predict subcellular localization, and 10 proteins scored as mitochondrial on at least one engine. The genes, fold induction, and mitochondrial scores are listed in Table 1.
HIF-1 downregulates mitochondrial oxygen consumption
Having identified several putative HIF-1 responsive gene products that had the potential to regulate mitochondrial function, we then directly measured mitochondrial oxygen consumption in cells exposed to long-term hypoxia. While other groups have studied mitochondrial function under acute hypoxia (Chandel et al., 1997), this is one of the first descriptions of mitochondrial function after long-term hypoxia where there have been extensive hypoxia-induced gene expression changes. Figure 1A is an example of the primary oxygen trace from a Clark electrode showing a drop in oxygen concentration in cell suspensions of primary fibroblasts taken from normoxic and hypoxic cultures. The slope of the curve is a direct measure of the total cellular oxygen consumption rate. Exposure of either primary human or immortalized mouse fibroblasts to 24 hr of hypoxia resulted in a reduction of this rate by approximately 50% (Figures 1A and 1B). In these experiments, the oxygen consumption can be stimulated with the mitochondrial uncoupling agent CCCP (carbonyl cyanide 3-chloro phenylhydrazone) and was completely inhibited by 2 mM potassium cyanide. We determined that the change in total cellular oxygen consumption was due to changes in mitochondrial activity by the use of the cell-permeable poison of mitochondrial complex 3, Antimycin A. Figure 1C shows that the difference in the normoxic and hypoxic oxygen consumption in murine fibroblasts is entirely due to the Antimycin-sensitive mitochondrial consumption. The kinetics with which mitochondrial function slows in hypoxic tumor cells also suggests that it is due to gene expression changes because it takes over 6 hr to achieve maximal reduction, and the reversal of this repression requires at least another 6 hr of reoxygenation (Figure 1D). These effects are not likely due to proliferation or toxicity of the treatments as these conditions are not growth inhibitory or toxic to the cells (Papandreou et al., 2005a).
Since we had predicted from the gene expression data that the mitochondrial oxygen consumption changes were due to HIF-1-mediated expression changes, we tested several genetically matched systems to determine what role HIF-1 played in the process (Figure 2). We first tested the cell lines that had been used for microarray analysis and found that the parental RCC4 cells had reduced mitochondrial oxygen consumption when compared to the VHL-reintroduced cells. Oxygen consumption in the parental cells was insensitive to hypoxia, while it was reduced by hypoxia in the wild-type VHL-transfected cell lines. Interestingly, stable introduction of a tumor-derived mutant VHL (Y98H) that cannot degrade HIF was also unable to restore oxygen consumption. These results indicate that increased expression of HIF-1 is sufficient to reduce oxygen consumption (Figure 2A). We also investigated whether HIF-1 induction was required for the observed reduction in oxygen consumption in hypoxia using two genetically matched systems. We measured normoxic and hypoxic oxygen consumption in murine fibroblasts derived from wild-type or HIF-1α null embryos (Figure 2B) and from human RKO tumor cells and RKO cells constitutively expressing ShRNAs directed against the HIF-1α gene (Figures 2C and 4C). Neither of the HIF-deficient cell systems was able to reduce oxygen consumption in response to hypoxia. These data from the HIF-overexpressing RCC cells and the HIF-deficient cells indicate that HIF-1 is both necessary and sufficient for reducing mitochondrial oxygen consumption in hypoxia.
HIF-dependent mitochondrial changes are functional, not structural
Because addition of CCCP could increase oxygen consumption even in the hypoxia-treated cells, we hypothesized that the hypoxic inhibition was a regulated activity, not a structural change in the mitochondria in response to hypoxic stress. We confirmed this interpretation by examining several additional mitochondrial characteristics in hypoxic cells such as mitochondrial morphology, quantity, and membrane potential. We examined morphology by visual inspection of both the transiently transfected mitochondrially localized DsRed protein and the endogenous mitochondrial protein cytochrome C. Both markers were indistinguishable in the parental RCC4 and the RCC4VHL cells (Figure 3A). Likewise, we measured the mitochondrial membrane potential with the functional dye rhodamine 123 and found that it was identical in the matched RCC4 cells and the matched HIF wt and knockout (KO) cells when cultured in normoxia or hypoxia (Figure 3B). Finally, we determined that the quantity of mitochondria per cell was not altered in response to HIF or hypoxia by showing that the amount of the mitochondrial marker protein HSP60 was identical in the RCC4 and HIF cell lines (Figure 3C)
PDK1 is a HIF-1 inducible target protein
After examination of the list of putative HIF-regulated mitochondrial target genes, we hypothesized that PDK1 could mediate the functional changes that we observed in hypoxia. We therefore investigated PDK1 protein expression in response to HIF and hypoxia in the genetically matched cell systems. Figure 4A shows that in the RCC4 cells PDK1 and the HIF-target gene BNip3 (Greijer et al., 2005 and Papandreou et al., 2005a) were both induced by hypoxia in a VHL-dependent manner, with the expression of PDK1 inversely matching the oxygen consumption measured in Figure 1 above. Likewise, the HIF wt MEFs show oxygen-dependent induction of PDK1 and BNip3, while the HIF KO MEFs did not show any expression of either of these proteins under any oxygen conditions (Figure 4B). Finally, the parental RKO cells were able to induce PDK1 and the HIF target gene BNip3L in response to hypoxia, while the HIF-depleted ShRNA RKO cells could not induce either protein (Figure 4C). Therefore, in all three cell types, the HIF-1-dependent regulation of oxygen consumption seen in Figure 2, corresponds to the HIF-1-dependent induction of PDK1 seen in Figure 4.
In order to determine if PDK1 was a direct HIF-1 target gene, we analyzed the genomic sequence flanking the 5′ end of the gene for possible HIF-1 binding sites based on the consensus core HRE element (A/G)CGTG (Caro, 2001). Several such sites exist within the first 400 bases upstream, so we generated reporter constructs by fusing the genomic sequence from −400 to +30 of the start site of transcription to the firefly luciferase gene. In transfection experiments, the chimeric construct showed significant induction by either cotransfection with a constitutively active HIF proline mutant (P402A/P564G) (Chan et al., 2002) or exposure of the transfected cells to 0.5% oxygen (Figure 4D). Most noteworthy, when the reporter gene was transfected into the HIF-1α null cells, it did not show induction when the cells were cultured in hypoxia, but it did show induction when cotransfected with expression HIF-1α plasmid. We then generated deletions down to the first 36 bases upstream of transcription and found that even this short sequence was responsive to HIF-1 (Figure 4D). Analysis of this small fragment showed only one consensus HRE site located in an inverted orientation in the 5′ untranslated region. We synthesized and cloned a mutant promoter fragment in which the core element ACGTG was replaced with AAAAG, and this construct lost over 90% of its hypoxic induction. These experiments suggest that it is this HRE within the proximal 5′ UTR that HIF-1 uses to transactivate the endogenous PDK1 gene in response to hypoxia.
PDK1 is responsible for the HIF-dependent mitochondrial oxygen consumption changes
In order to directly test if PDK1 was the HIF-1 target gene responsible for the hypoxic reduction in mitochondrial oxygen consumption, we generated RKO cell lines with either knockdown or overexpression of PDK1 and measured the oxygen consumption in these derivatives. The PDK1 ShRNA stable knockdown line was generated as a pool of clones cotransfected with pSUPER ShPDK1 and pTK-hygro resistance gene. After selection for growth in hygromycin, the cells were tested by Western blot for the level of PDK1 protein expression. We found that normoxic PDK1 is reduced by 75%, however, there was measurable expression of PDK1 in these cells in response to hypoxia (Figure 5A). When we measured the corresponding oxygen consumption in these cells, we found a change commensurate with the level of PDK1. The knockdown cells show elevated baseline oxygen consumption, and partial reduction in this activity in response to hypoxia. Therefore, reduction of PDK1 expression by genetic means increased mitochondrial oxygen consumption in both normoxic and hypoxic conditions. Interestingly, these cells still induced HIF-1α (Figure 5A) and HIF-1 target genes such as BNip3L in response to hypoxia (data not shown), suggesting that altered PDK1 levels do not alter HIF-1α function.

pdk1-expression-directly-regulates-cellular-oxygen-consumption-rate
PDK1 expression directly regulates cellular oxygen consumption rate
http://ars.els-cdn.com/content/image/1-s2.0-S155041310600060X-gr5.jpg
Figure 5. PDK1 expression directly regulates cellular oxygen consumption rate
- A)Western blot of RKO cell and ShRNAPDK1RKO cell lysates after exposure to 24 hr of normoxia or 0.5% O2. Blots were probed for HIF 1α, PDK1, and tubulin as a loading control.
- B)Oxygen consumption rate in RKO and ShRNAPDK1RKO cells after exposure to 24 hr of normoxia or 0.5% O2.
- C)Western blot of RKOiresGUS cell and RKOiresPDK1 cell lysates after exposure to 24 hr of normoxia or 0.5% O2. Blots were probed for HIF 1α, PDK1, and tubulin as a loading control.
- D)Oxygen consumption rate in RKOiresGUS and RKOiresPDK1 cells after exposure to 24 hr of normoxia or 0.5% O2.
- E)Model describing the interconnected effects of HIF-1 target gene activation on hypoxic cell metabolism. Reduced oxygen conditions causes HIF-1 to coordinately induce the enzymes shown in boxes. HIF-1 activation results in increased glucose transporter expression to increase intracellular glucose flux, induction of glycolytic enzymes increases the conversion of glucose to pyruvate generating energy and NADH, induction of PDK1 decreases mitochondrial utilization of pyruvate and oxygen, and induction of LDH increases the removal of excess pyruvate as lactate and also regenerates NAD+ for increased glycolysis.
For all graphs, the error bars represent the standard error of the mean.
We also determined if overexpression of PDK1 could lead to reduced mitochondrial oxygen consumption. A separate culture of RKO cells was transfected with a PDK1-IRES-puro expression plasmid and selected for resistance to puromycin. The pool of puromycin resistant cells was tested for PDK1 expression by Western blot. These cells showed a modest increase in PDK1 expression under control conditions when compared to the cells transfected with GUS-IRES-puro, with an additional increase in PDK1 protein in response to hypoxia (Figure 5C). The corresponding oxygen consumption measurements showed that the mitochondria is very sensitive to changes in the levels of PDK1, as even this slight increase was able to significantly reduce oxygen consumption in the normoxic PDK1-puro cultures. Further increase in PDK1 levels with hypoxia further reduced oxygen consumption in both cultures (Figure 5D). The model describing the relationship between hypoxia, HIF-1, PDK1, and intermediate metabolism is described inFigure 5E.
Altering oxygen consumption alters intracellular oxygen tension and sensitivity to hypoxia-dependent cell killing
The intracellular concentration of oxygen is a net result of the rate at which oxygen diffuses into the cell and the rate at which it is consumed. We hypothesized that the rate at which oxygen was consumed within the cell would significantly affect its steady-state intracellular concentrations. We tested this hypothesis in vitro using the hypoxic marker drug pimonidazole (Bennewith and Durand, 2004). We plated high density cultures of HIF wild-type and HIF knockout cells and placed these cultures in normoxic, 2% oxygen, and anoxic incubators for overnight treatment. The overnight treatment gives the cells time to adapt to the hypoxic conditions and establish altered oxygen consumption profiles. Pimonidozole was then added for the last 4 hr of the growth of the culture. Pimonidazole binding was detected after fixation of the cells using an FITC labeled anti-pimonidazole antibody and it was quantitated by flow cytometry. The quantity of the bound drug is a direct indication of the oxygen concentration within the cell (Bennewith and Durand, 2004). The histograms in Figure 6A show that the HIF-1 knockout and wild-type cells show similar staining in the cells grown in 0% oxygen. However, the cells treated with 2% oxygen show the consequence of the genetic removal of HIF-1. The HIF-proficient cells showed relatively less pimonidazole binding at 2% when compared to the 0% culture, while the HIF-deficient cells showed identical binding between the cells at 2% and those at 0%. We interpret these results to mean that the HIF-deficient cells have greater oxygen consumption, and this has lowered the intracellular oxygenation from the ambient 2% to close to zero intracellularly. The HIF-proficient cells reduced their oxygen consumption rate so that the rate of diffusion into the cell is greater than the rate of consumption.
Figure 6. HIF-dependent decrease in oxygen consumption raises intracellular oxygen concentration, protects when oxygen is limiting, and decreases sensitivity to tirapazamine in vitro
- A)Pimonidazole was used to determine the intracellular oxygen concentration of cells in culture. HIF wt and HIF KO MEFs were grown at high density and exposed to 2% O2or anoxia for 24 hr in glass dishes. For the last 4 hr of treatment, cells were exposed to 60 μg/ml pimonidazole. Pimonidazole binding was quantitated by flow cytometry after binding of an FITC conjugated anti-pimo mAb. Results are representative of two independent experiments.
- B)HIF1α reduces oxygen consumption and protects cells when total oxygen is limited. HIF wt and HIF KO cells were plated at high density and sealed in aluminum jigs at <0.02% oxygen. At the indicated times, cells were harvested, and dead cells were quantitated by trypan blue exclusion. Note both cell lines are equally sensitive to anoxia-induced apoptosis, so the death of the HIF null cells indicates that the increased oxygen consumption removed any residual oxygen in the jig and resulted in anoxia-induced death.
- C)PDK1 is responsible for HIF-1’s adaptive response when oxygen is limiting. A similar jig experiment was performed to measure survival in the parental RKO, the RKO ShRNAHIF1α, and the RKOShPDK1 cells. Cell death by trypan blue uptake was measured 48 hr after the jigs were sealed.
- D)HIF status alters sensitivity to TPZ in vitro. HIF wt and HIF KO MEFs were grown at high density in glass dishes and exposed to 21%, 2%, and <0.01% O2conditions for 18 hr in the presence of varying concentrations of Tirapazamine. After exposure, cells were harvested and replated under normoxia to determine clonogenic viability. Survival is calculated relative to the plating efficiency of cells exposed to 0 μM TPZ for each oxygen concentration.
- E)Cell density alters sensitivity to TPZ. HIF wt and HIF KO MEFs were grown at varying cell densities in glass dishes and exposed to 2% O2in the presence of 10 μM TPZ for 18 hr. After the exposure, survival was determined as described in (C).
For all graphs, the error bars represent the standard error of the mean.
HIF-induced PDK1 can reduce the total amount of oxygen consumed per cell. The reduction in the amount of oxygen consumed could be significant if there is a finite amount of oxygen available, as would be the case in the hours following a blood vessel occlusion. The tissue that is fed by the vessel would benefit from being economical with the oxygen that is present. We experimentally modeled such an event using aluminum jigs that could be sealed with defined amounts of cells and oxygen present (Siim et al., 1996). We placed 10 × 106 wild-type or HIF null cells in the sealed jig at 0.02% oxygen, waited for the cells to consume the remaining oxygen, and measured cell viability. We have previously shown that these two cell types are resistant to mild hypoxia and equally sensitive to anoxia-induced apoptosis (Papandreou et al., 2005a). Therefore, any death in this experiment would be the result of the cells consuming the small amount of remaining oxygen and dying in response to anoxia. We found that in sealed jigs, the wild-type cells are more able to adapt to the limited oxygen supply by reducing consumption. The HIF null cells continued to consume oxygen, reached anoxic levels, and started to lose viability within 36 hr (Figure 6B). This is a secondary adaptive effect of HIF1. We confirmed that PDK1 was responsible for this difference by performing a similar experiment using the parental RKO cells, the RKOShRNAHIF1α and the RKOShRNAPDK1 cells. We found similar results in which both the cells with HIF1α knockdown and PDK1 knockdown were sensitive to the long-term effects of being sealed in a jig with a defined amount of oxygen (Figure 6c). Note that the RKOShPDK1 cells are even more sensitive than the RKOShHIF1α cells, presumably because they have higher basal oxygen consumption rates (Figure 5B).
Because HIF-1 can help cells adapt to hypoxia and maintain some intracellular oxygen level, it may also protect tumor cells from killing by the hypoxic cytotoxin tirapazamine (TPZ). TPZ toxicity is very oxygen dependent, especially at oxygen levels between 1%–4% (Koch, 1993). We therefore tested the relative sensitivity of the HIF wt and HIF KO cells to TPZ killing in high density cultures (Figure 6D). We exposed the cells to the indicated concentrations of drug and oxygen concentrations overnight. The cells were then harvested and replated to determine reproductive viability by colony formation. Both cell types were equally resistant to TPZ at 21% oxygen, while both cell types are equally sensitive to TPZ in anoxic conditions where intracellular oxygen levels are equivalent (Figure 6A). The identical sensitivity of both cell types in anoxia indicates that both cell types are equally competent in repairing the TPZ-induced DNA damage that is presumed to be responsible for its toxicity. However, in 2% oxygen cultures, the HIF null cells displayed a significantly greater sensitivity to the drug than the wild-type cells. This suggests that the increased oxygen consumption rate in the HIF-deficient cells is sufficient to lower the intracellular oxygen concentration relative to that in the HIF-proficient cells. The lower oxygen level is significant enough to dramatically sensitize these cells to killing by TPZ.
If the increased sensitivity to TPZ in the HIF ko cells is determined by intracellular oxygen consumption differences, then this effect should also be cell-density dependent. We showed that this is indeed the case in Figure 6E where oxygen and TPZ concentrations were held constant, and increased cell density lead to increased TPZ toxicity. The effect was much more pronounced in the HIF KO cells, although the HIF wt cells showed some increased toxicity in the highest density cultures, consistent with the fact they were still consuming some oxygen, even with HIF present (Figure 1). The in vitro TPZ survival data is therefore consistent with our hypothesis that control of oxygen consumption can regulate intracellular oxygen concentration, and suggests that increased oxygen consumption could sensitize cells to hypoxia-dependent therapy.
Discussion
The findings presented here show that HIF-1 is actively responsible for regulating energy production in hypoxic cells by an additional, previously unrecognized mechanism. It has been shown that HIF-1 induces the enzymes responsible for glycolysis when it was presumed that low oxygen did not support efficient oxidative phosphorylation (Iyer et al., 1998 and Seagroves et al., 2001). The use of glucose to generate ATP is capable of satisfying the energy requirements of a cell if glucose is in excess (Papandreou et al., 2005a). We now find that at the same time that glycolysis is increasing, mitochondrial respiration is decreasing. However, the decreased respiration is not because there is not enough oxygen present to act as a substrate for oxidative phosphorylation, but because the flow of pyruvate into the TCA cycle has been reduced by the activity of pyruvate dehydrogenase kinase. Other reports have suggested that oxygen utilization is shifted in cells exposed to hypoxia, but these reports have focused on other regulators such as nitric oxide synthase (Hagen et al., 2003). NO can reduce oxygen consumption through direct inhibition of cytochrome oxidase, but this effect seems to be more significant at physiologic oxygen concentrations, not at severe levels seen in the tumor (Palacios-Callender et al., 2004).
7.9.8 HIF-1. upstream and downstream of cancer metabolism
Semenza GL1.
Curr Opin Genet Dev. 2010 Feb; 20(1):51-6
http://dx.doi.org/10.1016%2Fj.gde.2009.10.009
Hypoxia-inducible factor 1 (HIF-1) plays a key role in the reprogramming of cancer metabolism by activating transcription of genes encoding glucose transporters and glycolytic enzymes, which take up glucose and convert it to lactate; pyruvate dehydrogenase kinase 1, which shunts pyruvate away from the mitochondria; and BNIP3, which triggers selective mitochondrial autophagy. The shift from oxidative to glycolytic metabolism allows maintenance of redox homeostasis and cell survival under conditions of prolonged hypoxia. Many metabolic abnormalities in cancer cells increase HIF-1 activity. As a result, a feed-forward mechanism can be activated that drives HIF-1 activation and may promote tumor progression. Hypoxia-inducible factor 1 (HIF-1) plays a key role in the reprogramming of cancer metabolism by activating transcription of genes encoding glucose transporters and glycolytic enzymes, which take up glucose and convert it to lactate; pyruvate dehydrogenase kinase 1, which shunts pyruvate away from the mitochondria; and BNIP3, which triggers selective mitochondrial autophagy. The shift from oxidative to glycolytic metabolism allows maintenance of redox homeostasis and cell survival under conditions of prolonged hypoxia. Many metabolic abnormalities in cancer cells increase HIF-1 activity. As a result, a feed-forward mechanism can be activated that drives HIF-1 activation and may promote tumor progression.
Metastatic cancer is characterized by reprogramming of cellular metabolism leading to increased uptake of glucose for use as both an anabolic and catabolic substrate. Increased glucose uptake is such a reliable feature that it is utilized clinically to detect metastases by positron emission tomography using 18F-fluorodeoxyglucose (FDG-PET) with a sensitivity of ~90% [1]. As with all aspects of cancer biology, the details of metabolic reprogramming differ widely among individual tumors. However, the role of specific signaling pathways and transcription factors in this process is now understood in considerable detail. This review will focus on the involvement of hypoxia-inducible factor 1 (HIF-1) in both mediating metabolic reprogramming and responding to metabolic alterations. The placement of HIF-1 both upstream and downstream of cancer metabolism results in a feed-forward mechanism that may play a major role in the development of the invasive, metastatic, and lethal cancer phenotype.
O2 concentrations are significantly reduced in many human cancers compared to the surrounding normal tissue. The median PO2 in breast cancers is ~10 mm Hg, as compared to ~65 mm Hg in normal breast tissue [2]. Reduced O2 availability induces HIF-1, which regulates the transcription of hundreds of genes [3*,4*] that encode proteins involved in every aspect of cancer biology, including: cell immortalization and stem cell maintenance; genetic instability; glucose and energy metabolism; vascularization; autocrine growth factor signaling; invasion and metastasis; immune evasion; and resistance to chemotherapy and radiation therapy [5].
HIF-1 is a transcription factor that consists of an O2-regulated HIF-1α and a constitutively expressed HIF-1β subunit [6]. In well-oxygenated cells, HIF-1α is hydroxylated on proline residue 402 (Pro-402) and/or Pro-564 by prolyl hydroxylase domain protein 2 (PHD2), which uses O2 and α-ketoglutarate as substrates in a reaction that generates CO2 and succinate as byproducts [7]. Prolyl-hydroxylated HIF-1α is bound by the von Hippel-Lindau tumor suppressor protein (VHL), which recruits an E3-ubiquitin ligase that targets HIF-1α for proteasomal degradation (Figure 1A). Asparagine 803 in the transactivation domain is hydroxylated in well-oxygenated cells by factor inhibiting HIF-1 (FIH-1), which blocks the binding of the coactivators p300 and CBP [7]. Under hypoxic conditions, the prolyl and asparaginyl hydroxylation reactions are inhibited by substrate (O2) deprivation and/or the mitochondrial generation of reactive oxygen species (ROS), which may oxidize Fe(II) present in the catalytic center of the hydroxylases [8].

HIF-1 and metabolism nihms156580f1
HIF-1 and metabolism
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2822127/bin/nihms156580f1.gif
Figure 1 HIF-1 and metabolism. (A) Regulation of HIF-1α protein synthesis and stability and HIF-1-dependent metabolic reprogramming. The rate of translation of HIF-1α mRNA into protein in cancer cells is dependent upon the activity of the mammalian …
The finding that acute changes in PO2 increase mitochondrial ROS production suggests that cellular respiration is optimized at physiological PO2 to limit ROS generation and that any deviation in PO2 — up or down — results in increased ROS generation. If hypoxia persists, induction of HIF-1 leads to adaptive mechanisms to reduce ROS and re-establish homeostasis, as described below. Prolyl and asparaginyl hydroxylation provide a molecular mechanism by which changes in cellular oxygenation can be transduced to the nucleus as changes in HIF-1 activity. This review will focus on recent advances in our understanding of the role of HIF-1 in controlling glucose and energy metabolism, but it should be appreciated that any increase in HIF-1 activity that leads to changes in cell metabolism will also affect many other critical aspects of cancer biology [5] that will not be addressed here.
HIF-1 target genes involved in glucose and energy metabolism
HIF-1 activates the transcription of SLC2A1 and SLC2A3, which encode the glucose transporters GLUT1 and GLUT3, respectively, as well as HK1 and HK2, which encode hexokinase, the first enzyme of the Embden-Meyerhoff (glycolytic) pathway [9]. Once taken up by GLUT and phosphorylated by HK, FDG cannot be metabolized further; thus, FDG-PET signal is determined by FDG delivery to tissue (i.e. perfusion) and GLUT/HK expression/activity. Unlike FDG, glucose is further metabolized to pyruvate by the action of the glycolytic enzymes, which are all encoded by HIF-1 target genes (Figure 1A). Glycolytic intermediates are also utilized for nucleotide and lipid synthesis [10]. Lactate dehydrogenase A (LDHA), which converts pyruvate to lactate, and monocarboxylate transporter 4 (MCT4), which transports lactate out of the cell (Figure 1B), are also regulated by HIF-1 [9,11]. Remarkably, lactate produced by hypoxic cancer cells can be taken up by non-hypoxic cells and used as a respiratory substrate [12**].
Pyruvate represents a critical metabolic control point, as it can be converted to acetyl coenzyme A (AcCoA) by pyruvate dehydrogenase (PDH) for entry into the tricarboxylic acid (TCA) cycle or it can be converted to lactate by LDHA (Figure 1B). Pyruvate dehydrogenase kinase (PDK), which phosphorylates and inactivates the catalytic domain of PDH, is encoded by four genes and PDK1 is activated by HIF-1 [13,14]. (Further studies are required to determine whether PDK2, PDK3, or PDK4 is regulated by HIF-1.) As a result of PDK1 activation, pyruvate is actively shunted away from the mitochondria, which reduces flux through the TCA cycle, thereby reducing delivery of NADH and FADH2 to the electron transport chain. This is a critical adaptive response to hypoxia, because in HIF-1α–null mouse embryo fibroblasts (MEFs), PDK1 expression is not induced by hypoxia and the cells die due to excess ROS production, which can be ameliorated by forced expression of PDK1 [13]. MYC, which is activated in ~40% of human cancers, cooperates with HIF-1 to activate transcription of PDK1, thereby amplifying the hypoxic response [15]. Pharmacological inhibition of HIF-1 or PDK1 activity increases O2 consumption by cancer cells and increases the efficacy of a hypoxia-specific cytotoxin [16].
Hypoxia also induces mitochondrial autophagy in many human cancer cell lines through HIF-1-dependent expression of BNIP3 and a related BH3 domain protein, BNIP3L [19**]. Autocrine signaling through the platelet-derived growth factor receptor in cancer cells increases HIF-1 activity and thereby increases autophagy and cell survival under hypoxic conditions [21]. Autophagy may also occur in a HIF-1-independent manner in response to other physiological stimuli that are associated with hypoxic conditions, such as a decrease in the cellular ATP:AMP ratio, which activates AMP kinase signaling [22].
In clear cell renal carcinoma, VHL loss of function (LoF) results in constitutive HIF-1 activation, which is associated with impaired mitochondrial biogenesis that results from HIF-1-dependent expression of MXI1, which blocks MYC-dependent expression of PGC-1β, a coactivator that is required for mitochondrial biogenesis [23]. Inhibition of wild type MYC activity in renal cell carcinoma contrasts with the synergistic effect of HIF-1 and oncogenic MYC in activating PDK1 transcription [24].
Genetic and metabolic activators of HIF-1
Hypoxia plays a critical role in cancer progression [2,5] but not all cancer cells are hypoxic and a growing number of O2-independent mechanisms have been identified by which HIF-1 is induced [5]. Several mechanisms that are particularly relevant to cancer metabolism are described below.
Activation of mTOR
Alterations in mitochondrial metabolism
NAD+ levels
It is of interest that the NAD+-dependent deacetylase sirtuin 1 (SIRT1) was found to bind to, deacetylate, and increase transcriptional activation by HIF-2α but not HIF-1α [42**]. Another NAD+-dependent enzyme is poly(ADP-ribose) polymerase 1 (PARP1), which was recently shown to bind to HIF-1α and promote transactivation through a mechanism that required the enzymatic activity of PARP1 [43]. Thus, transactivation mediated by both HIF-1α and HIF-2α can be modulated according to NAD+ levels.
Nitric oxide
Increased expression of nitric oxide (NO) synthase isoforms and increased levels of NO have been shown to increase HIF-1α protein stability in human oral squamous cell carcinoma [44]. In prostate cancer, nuclear co-localization of endothelial NO synthase, estrogen receptor β, HIF-1α, and HIF-2α was associated with aggressive disease and the proteins were found to form chromatin complexes on the promoter of TERT gene encoding telomerase [45**]. The NOS2 gene encoding inducible NO synthase is HIF-1 regulated [5], suggesting another possible feed-forward mechanism.
7.9.9 In Vivo HIF-Mediated Reductive Carboxylation
Gameiro PA1, Yang J, Metelo AM, Pérez-Carro R, et al.
Cell Metab. 2013 Mar 5; 17(3):372-85.
http://dx.doi.org/10.1016%2Fj.cmet.2013.02.002
Hypoxic and VHL-deficient cells use glutamine to generate citrate and lipids through reductive carboxylation (RC) of α-ketoglutarate. To gain insights into the role of HIF and the molecular mechanisms underlying RC, we took advantage of a panel of disease-associated VHL mutants and showed that HIF expression is necessary and sufficient for the induction of RC in human renal cell carcinoma (RCC) cells. HIF expression drastically reduced intracellular citrate levels. Feeding VHL-deficient RCC cells with acetate or citrate or knocking down PDK-1 and ACLY restored citrate levels and suppressed RC. These data suggest that HIF-induced low intracellular citrate levels promote the reductive flux by mass action to maintain lipogenesis. Using [1–13C] glutamine, we demonstrated in vivo RC activity in VHL-deficient tumors growing as xenografts in mice. Lastly, HIF rendered VHL-deficient cells sensitive to glutamine deprivation in vitro, and systemic administration of glutaminase inhibitors suppressed the growth of RCC cells as mice xenografts.
Cancer cells undergo fundamental changes in their metabolism to support rapid growth, adapt to limited nutrient resources, and compete for these supplies with surrounding normal cells. One of the metabolic hallmarks of cancer is the activation of glycolysis and lactate production even in the presence of adequate oxygen. This is termed the Warburg effect, and efforts in cancer biology have revealed some of the molecular mechanisms responsible for this phenotype (Cairns et al., 2011). More recently, 13C isotopic studies have elucidated the complementary switch of glutamine metabolism that supports efficient carbon utilization for anabolism and growth (DeBerardinis and Cheng, 2010). Acetyl-CoA is a central biosynthetic precursor for lipid synthesis, being generated from glucose-derived citrate in well-oxygenated cells (Hatzivassiliou et al., 2005). Warburg-like cells, and those exposed to hypoxia, divert glucose to lactate, raising the question of how the tricarboxylic acid (TCA) cycle is supplied with acetyl-CoA to support lipogenesis. We and others demonstrated, using 13C isotopic tracers, that cells under hypoxic conditions or defective mitochondria primarily utilize glutamine to generate citrate and lipids through reductive carboxylation (RC) of α-ketoglutarate by isocitrate dehydrogenase 1 (IDH1) or 2 (IDH2) (Filipp et al., 2012; Metallo et al., 2012;Mullen et al., 2012; Wise et al., 2011).
The transcription factors hypoxia inducible factors 1α and 2α (HIF-1α, HIF-2α) have been established as master regulators of the hypoxic program and tumor phenotype (Gordan and Simon, 2007; Semenza, 2010). In addition to tumor-associated hypoxia, HIF can be directly activated by cancer-associated mutations. The von Hippel-Lindau (VHL) tumor suppressor is inactivated in the majority of sporadic clear-cell renal carcinomas (RCC), with VHL-deficient RCC cells exhibiting constitutive HIF-1α and/or HIF-2α activity irrespective of oxygen availability (Kim and Kaelin, 2003). Previously, we showed that VHL-deficient cells also relied on RC for lipid synthesis even under normoxia. Moreover, metabolic profiling of two isogenic clones that differ in pVHL expression (WT8 and PRC3) suggested that reintroduction of wild-type VHL can restore glucose utilization for lipogenesis (Metallo et al., 2012). The VHL tumor suppressor protein (pVHL) has been reported to have several functions other than the well-studied targeting of HIF. Specifically, it has been reported that pVHL regulates the large subunit of RNA polymerase (Pol) II (Mikhaylova et al., 2008), p53 (Roe et al., 2006), and the Wnt signaling regulator Jade-1. VHL has also been implicated in regulation of NF-κB signaling, tubulin polymerization, cilia biogenesis, and proper assembly of extracellular fibronectin (Chitalia et al., 2008; Kim and Kaelin, 2003; Ohh et al., 1998; Thoma et al., 2007; Yang et al., 2007). Hypoxia inactivates the α-ketoglutarate-dependent HIF prolyl hydroxylases, leading to stabilization of HIF. In addition to this well-established function, oxygen tension regulates a larger family of α-ketoglutarate-dependent cellular oxygenases, leading to posttranslational modification of several substrates, among which are chromatin modifiers (Melvin and Rocha, 2012). It is therefore conceivable that the effect of hypoxia on RC that was reported previously may be mediated by signaling mechanisms independent of the disruption of the pVHL-HIF interaction. Here we (1) demonstrate that HIF is necessary and sufficient for RC, (2) provide insights into the molecular mechanisms that link HIF to RC, (3) detected RC activity in vivo in human VHL-deficient RCC cells growing as tumors in nude mice, (4) provide evidence that the reductive phenotype ofVHL-deficient cells renders them sensitive to glutamine restriction in vitro, and (5) show that inhibition of glutaminase suppresses growth of VHL-deficient cells in nude mice. These observations lay the ground for metabolism-based therapeutic strategies for targeting HIF-driven tumors (such as RCC) and possibly the hypoxic compartment of solid tumors in general.
Functional Interaction between pVHL and HIF Is Necessary to Inhibit RC
Figure 1 HIF Inactivation Is Necessary for Downregulation of Reductive Carboxylation by pVHL
We observed a concurrent regulation in glucose metabolism in the different VHL mutants. Reintroduction of wild-type or type 2C pVHL mutant, which can meditate HIF-α destruction, stimulated glucose oxidation via pyruvate dehydrogenase (PDH), as determined by the degree of 13C-labeled TCA cycle metabolites (M2 enrichment) (Figures 1D and 1E). In contrast, reintroduction of an HIF nonbinding Type 2B pVHL mutant failed to stimulate glucose oxidation, resembling the phenotype observed in VHL-deficient cells (Figures 1D and 1E). Additional evidence for the overall glucose utilization was obtained from the enrichment of M3 isotopomers using [U13-C6]glucose (Figure S1A), which shows a lower contribution of glucose-derived carbons to the TCA cycle in VHL-deficient RCC cells (via pyruvate carboxylase and/or continued TCA cycling).
To test the effect of HIF activation on the overall glutamine incorporation in the TCA cycle, we labeled an isogenic pair of VHL-deficient and VHL-reconstituted UMRC2 cells with [U-13C5]glutamine, which generates M4 fumarate, M4 malate, M4 aspartate, and M4 citrate isotopomers through glutamine oxidation. As seen in Figure S1B, VHL-deficient/VHL-positive UMRC2 cells exhibit similar enrichment of M4 fumarate, M4 malate, and M4 asparate (but not citrate) showing that VHL-deficient cells upregulate reductive carboxylation without compromising oxidative metabolism from glutamine. … Labeled carbon derived from [5-13C1]glutamine can be incorporated into fatty acids exclusively through RC, and the labeled carbon cannot be transferred to palmitate through the oxidative TCA cycle (Figure 1B, red carbons). Tracer incorporation from [5-13C1]glutamine occurs in the one carbon (C1) of acetyl-CoA, which results in labeling of palmitate at M1, M2, M3, M4, M5, M6, M7, and M8 mass isotopomers. In contrast, lipogenic acetyl-CoA molecules originating from [U-13C6]glucose are fully labeled, and the labeled palmitate is represented by M2, M4, M6, M8, M10, M12, M14, and M16 mass isotopomers.
Figure 2 HIF Inactivation Is Necessary for Downregulation of Reductive Lipogenesis by pVHL
To determine the specific contribution from glucose oxidation or glutamine reduction to lipogenic acetyl-CoA, we performed isotopomer spectral analysis (ISA) of palmitate labeling patterns. ISA indicates that wild-type pVHL or pVHL L188V mutant-reconstituted UMRC2 cells relied mainly on glucose oxidation to produce lipogenic acetyl-CoA, while UMRC2 cells reconstituted with a pVHL mutant defective in HIF inactivation (Y112N or Y98N) primarily employed RC. Upon disruption of the pVHL-HIF interaction, glutamine becomes the preferred substrate for lipogenesis, supplying 70%–80% of the lipogenic acetyl-CoA (Figure 2C). This is not a cell-line-specific phenomenon, but it applies to VHL-deficient human RCC cells in general; the same changes are observed in 786-O cells reconstituted with wild-type pVHL or mutant pVHL or infected with vector only as control (Figure S2).
HIF Is Sufficient to Induce RC (reductive carboxylation) from Glutamine in RCC Cells
As shown in Figure 3C, reintroduction of wild-type VHLinto 786-O cells suppressed RC, whereas the expression of the constitutively active HIF-2α mutant was sufficient to stimulate this reaction, restoring the M1 enrichment of TCA cycle metabolites observed in VHL-deficient 786-O cells. Expression of HIF-2α P-A also led to a concomitant decrease in glucose oxidation, corroborating the metabolic alterations observed in glutamine metabolism (Figures 3D and 3E).
Figure 3 Expression of HIF-2α Is Sufficient to Induce Reductive Carboxylation and Lipogenesis from Glutamine in RCC Cells
Expression of HIF-2α P-A in 786-O cells phenocopied the loss-of-VHL with regards to glutamine reduction for lipogenesis (Figure 3G), suggesting that HIF-2α can induce the glutamine-to-lipid pathway in RCC cells per se. Although reintroduction of wild-type VHL restored glucose oxidation in UMRC2 and UMRC3 cells (Figures S3B–S3I), HIF-2α P-A expression did not measurably affect the contribution of each substrate to the TCA cycle or lipid synthesis in these RCC cells (data not shown). UMRC2 and UMRC3 cells endogenously express both HIF-1α and HIF-2α, whereas 786-O cells exclusively express HIF-2α. There is compelling evidence suggesting, at least in RCC cells, that HIF-α isoforms have overlapping—but also distinct—functions and their roles in regulating bioenergetic processes remain an area of active investigation. Overall, HIF-1α has an antiproliferative effect, and its expression in vitro leads to rapid death of RCC cells while HIF-2α promotes tumor growth (Keith et al., 2011; Raval et al., 2005).
Metabolic Flux Analysis Shows Net Reversion of the IDH Flux upon HIF Activation
To determine absolute fluxes in RCC cells, we employed 13C metabolic flux analysis (MFA) as previously described (Metallo et al., 2012). Herein, we performed MFA using a combined model of [U-13C6]glucose and [1-13C1]glutamine tracer data sets from the 786-O derived isogenic clones PRC3 (VHL−/ −)/WT8 (VHL+) cells, which show a robust metabolic regulation by reintroduction of pVHL. To this end, we first determined specific glucose/glutamine consumption and lactate/glutamate secretion rates. As expected, PRC3 exhibited increased glucose consumption and lactate production when compared to WT8 counterparts (Figure 4A). While PRC3 exhibited both higher glutamine consumption and glutamate production rates than WT8 (Figure 4A), the net carbon influx was higher in PRC3 cells (Figure 4B). Importantly, the fitted data show that the flux of citrate to α-ketoglutarate was negative in PRC3 cells (Figure 4C). This indicates that the net (forward plus reverse) flux of isocitrate dehydrogenase and aconitase (IDH + ACO) is toward citrate production. The exchange flux was also higher in PRC3 than WT8 cells, whereas the PDH flux was lower in PRC3 cells. In agreement with the tracer data, these MFA results strongly suggest that the reverse IDH + ACO fluxes surpass the forward flux in VHL-deficient cells. The estimated ATP citrate lyase (ACLY) flux was also lower in PRC3 than in WT8 cells. Furthermore, the malate dehydrogenase (MDH) flux was negative, reflecting a net conversion of oxaloacetate into malate in VHL-deficient cells (Figure 4C). This indicates an increased flux through the reductive pathway downstream of IDH, ACO, and ACLY. Additionally, some TCA cycle flux estimates downstream of α-ketoglutarate were not significantly different between PRC and WT8 (Table S1). This shows that VHL-deficient cells maintain glutamine oxidation while upregulating reductive carboxylation (Figure S1B). This finding is in agreement with the higher glutamine uptake observed in VHL-deficient cells. Table S1 shows the metabolic network and complete MFA results. …
Addition of citrate in the medium, in contrast to acetate, led to an increase in the citrate-to-α-ketoglutarate ratio (Figure 5L) and absolute citrate levels (Figure S4H) not only in VHL-deficient but alsoVHL-reconstituted cells. The ability of exogenous citrate, but not acetate, to also affect RC in VHL-reconstituted cells may be explained by compartmentalization differences or by allosteric inhibition of citrate synthase (Lehninger, 2005); that is, the ability of acetate to raise the intracellular levels of citrate may be limited in (VHL-reconstituted) cells that exhibit high endogenous levels of citrate. Whatever the mechanism, the results imply that increasing the pools of intracellular citrate has a direct biochemical effect in cells with regards to their reliance on RC. Finally, we assayed the transcript and protein levels of enzymes involved in the reductive utilization of glutamine and did not observe significant differences between VHL-deficient andVHL-reconstituted UMRC2 cells (Figures S4I and S4J), suggesting that HIF does not promote RC by direct transactivation of these enzymes. The IDH1/IDH2 equilibrium is defined as follows:
[α−ketoglutrate][NADPH][CO2]/[Isocitrate][NADP+]=K(IDH)
Figure 5 Regulation of HIF-Mediated Reductive Carboxylation by Citrate Levels
We sought to investigate whether HIF could affect the driving force of the IDH reaction by also enhancing NADPH production. We did not observe a significant alteration of the NADP+/NADPH ratio between VHL-deficient and VHL-positive cells in the cell lysate (Figure S4I). Yet, we determined the ratio of the free dinucleotides using the measured ratios of suitable oxidized (α-ketoglutarate) and reduced (isocitrate/citrate) metabolites that are linked to the NADP-dependent IDH enzymes. The determined ratios (Figure S4J) are in close agreement with the values initially reported by the Krebs lab (Veech et al., 1969) and showed that HIF-expressing UMRC2 cells exhibit a higher NADP+/NADPH ratio. Collectively, these data strongly suggest that HIF-regulated citrate levels modulate the reductive flux to maintain adequate lipogenesis.
Reductive Carboxylation from Glutamine Is Detectable In Vivo
Figure 6 Evidence for Reductive Carboxylation Activity In Vivo
Loss of VHL Renders RCC Cells Sensitive to Glutamine Deprivation
We hypothesized that VHL deficiency results in cell addiction to glutamine for proliferation. We treated the isogenic clones PRC3 (VHL-deficient cells) and WT8 (VHL-reconstituted cells) with the glutaminase inhibitor 968 (Wang et al., 2010a). VHL-deficient PRC3 cells were more sensitive to treatment with 968, compared to the VHL-reconstituted WT8 cells (Figure 7A). To confirm that this is not only a cell-line-specific phenomenon, we also cultured UMRC2 cells in the presence of 968 or diluent control and showed selective sensitivity of VHL-deficient cells (Figure 7B).
Figure 7 VHL-Deficient Cells and Tumors Are Sensitive to Glutamine Deprivation
(A–E) Cell proliferation is normalized to the corresponding cell type grown in 1 mM glutamine-containing medium. Effect of treatment with glutaminase (GLS) inhibitor 968 in PRC3/WT8 (A) and UMRC2 cells (B). Rescue of GLS inhibition with dimethyl alpha-ketoglutarate (DM-Akg; 4 mM) or acetate (4 mM) in PRC3/WT8 clonal cells (C) and polyclonal 786-O cells (D). Effect of GLS inhibitor BPTES in UMRC2 cells (E). Student’s t test compares VHL-reconstituted cells to control cells in (A), (B), and (E) and DM-Akg or acetate-rescued cells to correspondent control cells treated with 968 only in (C) and (D) (asterisk in parenthesis indicates comparison between VHL-reconstituted to control cells). Error bars represent SEM.
(F) GLS inhibitor BPTES suppresses growth of human UMRC3 RCC cells as xenografts in nu/nu mice. When the tumors reached 100mm3, injections with BPTES or vehicle control were carried out daily for 14 days (n = 12). BPTES treatment decreases tumor size and mass (see insert). Student’s t test compares control to BPTES-treated mice (F). Error bars represent SEM.
(G) Diagram showing the regulation of reductive carboxylation by HIF.
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4003458/bin/nihms449661f7.jpg
In summary, our findings show that HIF is necessary and sufficient to promote RC from glutamine. By inhibiting glucose oxidation in the TCA cycle and reducing citrate levels, HIF shifts the IDH reaction toward RC to support citrate production and lipogenesis (Figure 7G). The reductive flux is active in vivo, fuels tumor growth, and can potentially be targeted pharmacologically. Understanding the significance of reductive glutamine metabolism in tumors may lead to metabolism-based therapeutic strategies.
Along with others, we reported that hypoxia and loss of VHL engage cells in reductive carboxylation (RC) from glutamine to support citrate and lipid synthesis (Filipp et al., 2012; Metallo et al., 2012; Wise et al., 2011). Wise et al. (2011) suggested that inactivation of HIF in VHL-deficient cells leads to reduction of RC. These observations raise the hypothesis that HIF, which is induced by hypoxia and is constitutively active inVHL-deficient cells, mediates RC. In our current work, we provide mechanistic insights that link HIF to RC. First, we demonstrate that polyclonal reconstitution of VHL in several human VHL-deficient RCC cell lines inhibits RC and restores glucose oxidation. Second, the VHL mutational analysis demonstrates that the ability of pVHL to mitigate reductive lipogenesis is mediated by HIF and is not the outcome of previously reported, HIF-independent pVHL function(s). Third, to prove our hypothesis we showed that constitutive expression of a VHL-independent HIF mutant is sufficient to phenocopy the reductive phenotype observed in VHL-deficient cells. In addition, we showed that RC is not a mere in vitro phenomenon, but it can be detected in vivo in human tumors growing as mouse xenografts. Lastly, treatment of VHL-deficient human xenografts with glutaminase inhibitors led to suppression of their growth as tumors.
7.9.10 Evaluation of HIF-1 inhibitors as anticancer agents
Semenza GL1.
Drug Discov Today. 2007 Oct; 12(19-20):853-9
http://dx.doi.org/10.1016/j.drudis.2007.08.006
Hypoxia-inducible factor 1 (HIF-1) regulates the transcription of many genes involved in key aspects of cancer biology, including immortalization, maintenance of stem cell pools, cellular dedifferentiation, genetic instability, vascularization, metabolic reprogramming, autocrine growth factor signaling, invasion/metastasis, and treatment failure. In animal models, HIF-1 overexpression is associated with increased tumor growth, vascularization, and metastasis, whereas HIF-1 loss-of-function has the opposite effect, thus validating HIF-1 as a target. In further support of this conclusion, immunohistochemical detection of HIF-1α overexpression in biopsy sections is a prognostic factor in many cancers. A growing number of novel anticancer agents have been shown to inhibit HIF-1 through a variety of molecular mechanisms. Determining which combination of drugs to administer to any given patient remains a major obstacle to improving cancer treatment outcomes.
Like this:
Like Loading...
Read Full Post »



 Published: May 28, 2008
Published: May 28, 2008 



































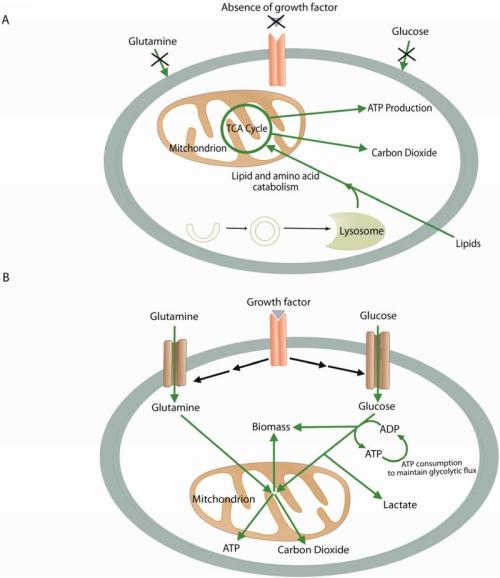








Warburg Effect revisited
It is very interesting the series of commentaries following Warburg Effect revisited. However, it comes as no surprise that almost all of them have small or greater emphasis in the molecular biology (changes in gene expression) events of the metabolic regulation involved.
I would like to comment on some aspects: 1- Warburg did the initial experiments following Pasteur line of reasoning that aimed at carbon flow through the cell (yeast in his case) instead of describing anything inside the cell. It is worth to recall that for the sake of his study, Pasteur considered anything inside the cell under the domain of divine forces. He, at least in defence of his work, entirely made outside the cell, considered that inside the cells was beyond human capability of understanding – He has followed vitalism as his line of reasoning in defence of his work – Interestingly, the same scientist that has ruled out spontaneous generation when Pasteurization was started. Therefore, Pasteur measured everything outside the cell (mainly sugar, ethanol – the equivalent of our lactic acid end product of anaerobic metabolism) and found that as soon as yeasts were placed in the presence of oxygen, sugar was consumed at low speed in comparison with the speed measured in anaerobiosis and ethanol was also produced at reduced speed. This is an indication of a fast biological regulatory mechanism that obviously, do not require changes in gene expression. As previously said, Warburg work translated for republishing in the Journal Biological Chemistry mentioned “grana” for mitochondria calling attention on an “inside-the-cell” component. It seems that, there is not a unique, single site of metabolism, where the Pasteur Effect – Warburg Effect seems to be elicited by the shift from anaerobiosis to aerobiosis or vice versa.
In order to find a core for the mechanism the best approach seems to take into account one of the most important contributions of one of the greatest north-American biochemists, Briton Chance. He has made it with his polarographic method of following continuously the oxygen consumption of the cell´s mitochondria.
Mitochondria burn organic carbon molecules under a very stringent control mechanism of oxidative-phosphorylation ATP production. Measured in the form of changes in the speed of oxygen consumption over time as Respiratory Control Ratio (RCR). When no ATP is required by the cell, oxygen consumption goes at low speed (basal or state II or IV). When ADP is offered to the mitochondria as an indication that ATP synthesis is necessary, oxygen consumption is activated in state III respiration. Low respiration means low burning activity of organic (carbon) molecules what in this case, means indirectly low glucose consumption. While high respiration is the converse – greater glucose consumption.
Aerobic metabolism of glucose to carbonic acid and water provides a change in free energy enough for 38 molecules of ATP (the real production is +/- 32 ATP in aerobic condition) while glucose to lactic acid metabolism in anaerobiosis leads to 2 ATP production after discounting the other 2 required at initial stages of glucose metabolism.
The low ATP yield in anaerobiosis explains the fast glucose metabolism in anaerobiosis while the control by RCR in mitochondria explains the reduction in glucose metabolism under aerobiosis as long as the ATP requirements of the cell remains the same – This is what it is assumed to happen in quiescent cells. Not necessarily in fast growing cells as cancer cells are. However, this will not be discussed here. In my first experiments in the early seventies, with M. Rouxii a dimorphic mold-yeast biological system the environmental change (aerobic – anaerobic) led to morphogenetic change presented as morphogenetic expression of the Pasteur Effect. In this case, the enzyme that replaces mitochondria in ATP production (Pyruvate Kinase) converting phosphoenolpyruvate into pyruvate together with ADP into ATP, shows changes that can be interpreted as change in gene expression together with new self-assembly of enzyme subunits. (Dimer AA – yeast in anaerobic growth or sporangiospores- converted into dimer AB in aerobic mold). In Leloir opinions at that time, PK I (AA) was only highly glycosylated, while PK II (AB) was less glycosylated without changes in gene expression.
In case you read comments posted, you will see that the reference to aerobic glycolysis, continues to be made together with, new deranged forms of reasoning as is indicated by referring to: Mitochondrial role in ion homeostasis…
Homeostasis is a regulation of something, ions, molecules, pH etc. that is kept outside the cell, therefore any role for mitochondria on it is only made indirectly, by its ATP production.
However, mitochondria has a role together with other cell components in the regulation of for instance, intracellular Ca levels (Something that is not a homeostatic regulation). This is a very important point for the following reason: Homeostasis is maintained as a composite result of several differentiated cellular, tissue and organ functions. Differentiated function is something clearly missing in cancer cells. The best form to refer to the mitochondrial function regarding ions is to indicate a mitochondrial role in ion fluxes.
In short, to indicate how an environmental event or better saying condition could favour genetic changes instead of being caused by genetic changes is to follow the same line of reasoning that is followed in understanding the role of cardioplegia. To stop heart beating is adequate for heart surgery it is also adequate for heart cells by sparing the ATP use during surgery and therefore, offering better recovery condition to the heart afterwards.
In the case, here considered, even assuming that the genome is not made more unstable during hypoxic condition it is quite possible to understand that sharing ATP with both differentiated cell function and replication may led quality control of DNA in short supply of much needed ATP and this led to maintenance of mutations as well as less organized genome.
Thank you. I enjoy reading your comments. They are very instructive. I don’t really think that I comprehend the use of the term “epigenetics” and longer. In fact, it was never clear to me when I first heard it used some years ago.
The term may have been closely wedded to the classic hypothesis of a unidirectional DNA–> RNA–> protein model that really has lost explanatory validity for the regulated cell in its environment. The chromatin has an influence, and protein-protein interactions are everywhere. As you point out, these are adjusting to a fast changing substrate milieu, and the genome is not involved. But in addition, the proteins may well have a role in suppression or activation of signaling pathways, and thereby, may well have an effect on gene expression. I don’t have any idea about how it would work, but mutations would appear to follow the metabolic condition of the cell over time. It would appear to be – genomic modification.
In aerobic glucose metabolism, the oxidation of citric acid requires ADP and Mg²+, which will increase the speed of the reaction: Iso-citric acid + NADP (NAD) — isocitrate dehydrogenase (IDH) = alpha-ketoglutaric acid. In the Krebs cycle (the citric cycle), IDH1 and IDH2 are NADP+-dependent enzymes that normally catalyze the inter-conversion of D-isocitrate and alpha-ketoglutarate (α-KG). The IDH1 and IDH2 genes are mutated in > 75% of different malignant diseases. Two distinct alterations are caused by tumor-derived mutations in IDH1 or IDH2: the loss of normal catalytic activity in the production of α-ketoglutarate (α-KG) and the gain of catalytic activity to produce 2-hydroxyglutarate (2-HG), [22].
This product is a competitive inhibitor of multiple α-KG-dependent dioxygenases, including demethylases, prolyl-4-hydroxylase and the TET enzymes family (Ten-Eleven Translocation-2), resulting in genome-wide alternations in histones and DNA methylation. [23]
IDH1 and IDH2 mutations have been observed in myeloid malignancies, including de novo and secondary AML (15%–30%), and in pre-leukemic clone malignancies, including myelodysplastic syndrome and myeloproliferative neoplasm (85% of the chronic phase and 20% of transformed cases in acute leukemia), [24].
Normally, cells in the body communicate via intra-cytoplasmic channels and maintain the energetic potential across cell membranes, which is 1-2.5 µmol of ATP in the form of ATP-ADP/ATP-ADP-IMP. These normal energetic values occur during normal cell division. If the intra-cellular and extra-cellular levels of Mg2+ are high, the extra-cellular charges of the cells will not be uniformly distributed.
This change in distribution induces a high net positive charge for the cell and induces a loss of contact inhibition via the electromagnetic induction of oscillation [28, 29, 30]. Thereafter, malignant cells become invasive and metastasize.
,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,
-22. Hartmann C, Meyer J, Balss J. Capper D, et al. Type and frequency of IDH1 and IDH2 mutations are related to astrocytic and oligodendroglial differentiation and age: a study of 1,010 diffuse gliomas. Acta Neuropathol 2009; 118: 464-474.
23. Raymakers R.A, Langemeijer S.M., Kuiper R.P, Berends M, et al. Acquired mutations in TET2 are common in myelodysplastic syndromes. Nat. Genet 2009; 41; 838–849.
24 Wagner K, Damm F, Gohring G., Gorlich K et al. Impact of IDH1 R132 mutations and an IDH1 single nucleotide polymorphism in cytogenetically normal acute myeloid leukemia: SNP rs11554137 is an adverse prognostic factor. J. Clin. Oncol.2010; 28: 2356–2364.
Plant Molecular Biology 1989; 1: 271–303.
29. Chien MM, Zahradka CE, Newel MC, Fred JW. Fas induced in B cells apoptosis require an increase in free cytosolic magnesium as in early event. J Biol Chem.1999; 274: 7059-7066.
30. Milionis H J, Bourantas C L, Siamopoulos C K, Elisaf MS. Acid bases and electrolytes abnormalities in Acute Leukemia. Am J Hematol 1999; (62): 201-207.
31. Thomas N Seyfried; Laura M Shelton.Cancer as a Metabolic Disease. Nutr Metab 2010; 7: 7
– Aurelian Udristioiu, M.D,
– Lab Director, EuSpLM,
– City Targu Jiu, Romania
AACC, National Academy of Biochemical Chemistry (NACB) Member, Washington D.C, USA.