Reporter: Aviva Lev-Ari, PhD, RN
http://www.biomarkerworldcongress.com/
DOWNLOADS
http://www.biomarkerworldcongress.com/bmc_content.aspx?id=95326
Conference Programs
May 6 – 7, 2013
Track 1: Translational Biomarkers in Drug Development
Track 2: Clinical Assay Development
Track 3: Cancer Tissue Diagnostics
May 6 – 8, 2013
Track 4: Executive Summit: Companion Diagnostics
May 7 – 8, 2013
Track 5: Biomarkers for Patient Selection
Track 6: Cancer Drug Resistance
Track 7: Exosomes and Microvesicles as Biomarkers
and Diagnostics
SPEAKERS
SPEAKERS
Jason M. Aliotta, M.D., Assistant Professor, Medicine, Warren Alpert Medical School, Brown University
John L. Allinson, FIBMS, Vice President, Biomarker Laboratory Services, ICON Development Solutions
Maria E. Arcila, M.D., Department of Pathology, Memorial Sloan-Kettering Cancer Center
Khusru Asadullah, M.D., Vice President and Head, Global Biomarkers, Bayer Pharma
Jiri Aubrecht, Pharm.D., Ph.D., Senior Director, Safety Biomarker Group Lead, Drug Safety Research & Development, Pfizer
M.J. Finley Austin, Ph.D., Personalized Healthcare & Biomarker Strategy Director, AstraZeneca
Nazneen Aziz, Ph.D., Director, Molecular Medicine, Transformation Program Office, College of American Pathologists
Geoffrey Stuart Baird, M.D., Ph.D., Assistant Professor, Laboratory Medicine, University of Washington
Robert A. Beckman, M.D., External Faculty, Center for Evolution and Cancer, Helen Diller Family Cancer Center, UCSF; Executive Director, Clinical Development Oncology, Daiichi Sankyo Pharma Development
Darrell R. Borger, Ph.D., Co-Director, Translational Research Laboratory, Massachusetts General Hospital Cancer Center
Mark Broenstrup, Ph.D., Director, Biomarker and Diagnostics, R&D Diabetes Division, Sanofi
Michael Burczynski, Ph.D., Executive Director, Biomarker Technologies, Discovery Medicine and Clinical Pharmacology, Bristol-Myers Squibb
Claudio Carini, M.D., Global Clinical Immunology and Biomarkers Lead, Bioenhancement Development Unit, Pfizer
Luigi Catanzariti, Ph.D., Executive Director and Global Program Director, Diagnostics, Novartis
David Chen, Ph.D., Senior Director, Correlative Sciences, Oncology Clinical Development, Novartis Pharmaceuticals
Carol Cheung, M.D., Ph.D., Department of Pathology, University Health Network
Atish Choudhury, M.D., Instructor in Medicine, Medical Oncology, Dana-Farber Cancer Institute
Seth Crosby, M.D., Director, Partnerships & Alliances, Washington University School of Medicine
Mark E. Curran, Ph.D., Vice President, Immunology Biomarkers, Janssen Research & Development
Stephen P. Day, Ph.D., Director, Medical Affairs, Hologic
Viswanath Devanarayan, Ph.D., Global Head, Exploratory Statistics, AbbVie, Inc.
Luis Alberto Diaz, M.D., Associate Professor of Oncology, Johns Hopkins Sidney Kimmel Comprehensive Cancer Center
Max Diem, Ph.D., Professor, Chemistry and Chemical Biology, Northeastern University
Nicholas C. Dracopoli, Ph.D., Vice President, Janssen R&D, Johnson & Johnson
Crislyn D’Souza-Schorey, Ph.D., Professor, Biological Sciences, University of Notre Dame
Dominik Duelli, Ph.D., Assistant Professor, Cellular and Molecular Pharmacology, Rosalind Franklin University of Medicine & Science, Chicago Medical School
Daniel Edelman, Ph.D., Facility Head, Clinical Molecular Profiling Core, National Cancer Institute, NIH
Reyna Favis, Ph.D., Scientific Director, Clinical Research & Development, Janssen Pharmaceutical Companies of Johnson & Johnson
Andrea Ferreira-Gonzalez, Ph.D., Professor and Chair, Division of Molecular Diagnostics; Director, Molecular Diagnostics Laboratory, Department of Pathology, Virginia Commonwealth University
Andrew Fish, Executive Director, AdvaMedDx
Herbert A. Fritsche, Ph.D., Senior Vice President and CSO, Health Discovery Corporation
Felix Frueh, Entrepreneur-in-Residence, Third Rock Ventures
Steve Furlong, Ph.D., Safety Science Lead, AstraZeneca Clinical Development
George A. Green IV, Ph.D., Director, Pharmacodiagnostics, Bristol-Myers Squibb
Patrick Groody, Ph.D., Divisional Vice President, Research & Development, Abbott
Steven Gutman, M.D., MBA, Strategic Advisor, Myraqa
Abdel Halim, Pharm.D., Ph.D., DABCC-MDx, DABCC-TOX, DABCC-CC, FACB, Director, Clinical Biomarkers, Daiichi Sankyo Pharma Development
Sam Hanash, M.D., Ph.D., Director, McCombs Institute for Cancer Early Detection and Treatment, MD Anderson Cancer Center
Charles R. Handorf, M.D., Ph.D., Professor and Chair, Pathology and Laboratory Medicine, University of Tennessee
Amir Handzel, Ph.D., Associate Director, Translational and Clinical Sciences, OSI Pharma (Astellas)
Chunhai “Charlie” Hao, M.D., Ph.D., Associate Professor, Neuropathology Attending, Department of Pathology & Laboratory Medicine, Emory University School of Medicine
Madhuri Hegde, Ph.D., Executive Director, Emory Genetics Laboratory; Associate Professor, Human Genetics, Emory University School of Medicine
Philip Hewitt, Ph.D., Head, Early Non-Clinical Safety (Liver and Kidney), Merck Serono
Stephen M. Hewitt, M.D., Ph.D., Clinical Investigator, Laboratory of Pathology, National Cancer Institute, NIH
Holly Hilton, Ph.D., Head, Disease and Translational Genomics, Hoffmann-La Roche; Adjunct Professor, University of Medicine and Dentistry New Jersey
Fred H. Hochberg, M.D., Associate Professor, Neurology, Massachusetts General Hospital
Shidong Jia, Ph.D., Scientist, Oncology Biomarker Development, Genentech
Chris Jowett, General Manager, Commercial Operations, Abbott Molecular
Jingfang Ju, Ph.D., Co-Director of Translational Research, Pathology, Stony Brook University
Peter M. Kazon, General Counsel, American Clinical Laboratory Association
Eric Lai, Ph.D., Senior Vice President and Head, Pharmacogenomics, Takeda Pharmaceuticals International
Ira M. Lubin, Ph.D., Team Lead, Genetics Laboratory Research and Evaluation Branch, CDC
Johan Luthman, D.D.S., Ph.D., Senior Program Leader, Neuroscience & Ophthalmology Research & Development Franchise Integrator, Merck
Elaine Lyon, Ph.D., Medical Director, Molecular Genetics, ARUP Laboratories
Ron Mazumder, Ph.D., MBA, Global Head, Research and Product Development, Janssen Diagnostics, Janssen Pharmaceutical Companies of Johnson & Johnson
Duncan McHale, Ph.D., Vice President, Global Exploratory Development at UCB Pharma
Alan Mertz, President, American Clinical Laboratory Association
Yoshi Oda, Ph.D., President, Biomarkers and Personalized Medicine Core Function Unit, Eisai
Lorraine O’Driscoll, Ph.D., Associate Professor, Pharmacology; Director, Research, School of Pharmacy and Pharmaceutical Sciences, Trinity College Dublin
Carol S. Palackdharry, M.D., MS, Medical Director, ActiveHealth Management; Clinical Lead, Oncology Condition Analysis, Aetna
Saumya Pant, Ph.D., Research Fellow, Merck
Liron Pantanowitz, M.D., Associate Professor, Pathology and Biomedical Informatics, University of Pittsburgh Medical Center
Scott D. Patterson, Ph.D., Executive Director, Medical Sciences, Amgen
Sonia Pearson-White, Ph.D., Scientific Program Manager, Oncology, The Biomarkers Consortium, Foundation for the National Institutes of Health
Emanuel Petricoin III, Ph.D., Co-Director, The Center for Applied Proteomics and Molecular Medicine, George Mason University
Suso Platero, Ph.D., Director, Oncology Biomarkers, Janssen Pharmaceuticals
Mark Priebe, MT(ASCP)SBB, Managing Director, QualityStar Quality Consortium
Debra Rasmussen, MBA, Senior Director, Regulatory Affairs, Johnson & Johnson
Hallgeir Rui, M.D., Ph.D., Professor, Cancer Biology, Medical Oncology, and Pathology, Thomas Jefferson University
Hakan Sakul, Ph.D., Executive Director and Head, Diagnostics, Worldwide R&D, Clinical Research and Precision Medicine, Pfizer
Kurt A. Schalper, M.D., Ph.D., Associate Research Scientist, Pathology, Yale School of Medicine
Stephen C. Schmechel, M.D., Ph.D., Associate Professor, Pathology, University of Washington School of Medicine
Robert Schupp, Ph.D., Executive Director, Diagnostics Hematology/Oncology, Celgene Corporation
Jason S. Simon, Ph.D., Director, Immuno-Oncology Biomarkers, Discovery Medicine and Clinical Pharmacology, Bristol-Myers Squibb
Sharon Sokolowski, Ph.D., Principal Scientist, Pfizer Global Research & Development
Gyongyi Szabo, M.D., Ph.D., Professor, Gastroenterology, University of Massachusetts Medical School
Douglas D. Taylor, Ph.D., Professor, Obstetrics and Gynecology, University of Louisville School of Medicine
Meghna Das Thakur, Ph.D., Presidential PostDoctoral Fellow, Novartis Institutes for BioMedical Research
Emina Torlakovic, M.D., Ph.D., Associate Professor, Laboratory Medicine and Pathobiology, University of Toronto
Jessie Villanueva, Ph.D., Assistant Professor, Molecular & Cellular Oncogenesis Program, The Wistar Institute
Glen J. Weiss, M.D., Co-Head, Lung Cancer Unit, The Translational Genomics Research Institute (TGen); Director, Clinical Research, Cancer Treatment Centers of America; CMO, CRAB-Clinical Trials Consortium
David Wholley, Director, Biomarkers Consortium, Foundation for the NIH
David T.W. Wong, D.M.D., D.M.Sc., Professor, Associate Dean of Research, UCLA School of Dentistry and Director of Dental Research Institute
Janghee Woo, M.D., Albert Einstein Medical Center
Wen Jin Wu, M.D., Ph.D., Principal Investigator, Division of Monoclonal Antibodies, Office of Biotechnology Products, Center for Drug Evaluation and Research, FDA
Brenda Yanak, Ph.D., Precision Medicine Leader, Clinical Innovation, Pfizer
Tammie Yeh, Ph.D., Molecular Biomarkers, Oncology Lead, Merck
Eunhee S. Yi, M.D., Consultant, Anatomic Pathology, Mayo Clinic; Professor, Pathology, Mayo Clinic College of Medicine
Malcolm York, MPhil, Director and Head, Clinical Pathology and Safety Assessment, GlaxoSmithKline R&D
Theresa Zhang, Ph.D., Associate Director, Exploratory and Translational Sciences, Merck
BIOMARKERS
& DIAGNOSTICS
world congress 2013
MAY 6 – 8, 2013 | LOEWS PHILADELPHIA HOTEL | PHILADELPHIA, PA
Cambridge Healthtech Institute’s Ninth Annual
BiomarkerWorldCongress.com
The Leading Annual Meeting Dedicated to Biomarkers
and Diagnostics Research and Implementation
Dinner Courses:
Fit-for-Purpose Biomarker Assay
Development and Validation
Next-Generation Sequencing as
a Clinical Test
Laboratory-Developed Tests
Conference Programs:
May 6 – 7, 2013
Track 1: Translational
Biomarkers in Drug Development
Track 2: Clinical Assay
Development
Track 3: Cancer Tissue
Diagnostics
May 6 – 8, 2013
Track 4: Executive Summit:
Companion Diagnostics
May 7 – 8, 2013
Track 5: Biomarkers for
Patient Selection
Track 6: Cancer Drug Resistance
Track 7: Exosomes and
Microvesicles as Biomarkers
and Diagnostics
Register by March 29th and SAVE up to $250!
Premier Sponsor
Featured Speakers
Hakan Sakul
Head, Diagnostics
Pfizer
Yoshi Oda
President, Biomarkers & Personalized Medicine
Eisai
David Wholley
Director
Biomarkers Consortium
Khusru Asadullah
VP, Head, Global Biomarkers
Bayer
Nicholas C. Dracopoli
VP, Janssen R&D
Johnson & Johnson
Eric Lai
SVP, Head, Pharmacogenomics
Takeda
2 | Biomarkers & Diagnostics World Congress BiomarkerWorldCongress.com
BIOMARKERS & DIAGNOSTICS
world congress 2013
Track 1: Translational Biomarkers
in Drug Development
Track 2: Clinical
Assay Development
Track 3: Cancer Tissue Diagnostics Track 4: Executive Summit:
Companion Diagnostics*
Sunday, May 5
5:00-6:00 Conference Pre-Registration
Monday, May 6
8:30-10:00 Biomarkers in Translational Medicine From Research Biomarkers to
Clinical Assays
Whole-Slide Imaging and
Digital Pathology
Commercialization of
Companion Diagnostics
10:00-10:30 Networking Coffee Break
10:30-11:50 Biomarkers in Translational Medicine From Research Biomarkers to
Clinical Assays
Whole-Slide Imaging and
Digital Pathology
Commercialization of
Companion Diagnostics
11:50-1:20 Luncheon Presentation
Sponsored by
Lunch on Your Own
1:20-2:40 Biomarker Utility in Clinical Development NGS in Clinical Use Strategies for Rx-Dx Partnerships
2:40-3:40 Refreshment Break in the Exhibit Hall with Poster Viewing
3:40-5:00 Biomarker Utility in Clinical Development NGS in Clinical Use Strategies for Rx-Dx Partnerships
5:00-6:00 Networking Reception in the Exhibit Hall with Poster Viewing
6:00-9:00 Dinner Courses (Separate registration required)
Fit-for-Purpose Biomarker Assay Development and Validation
Next-Generation Sequencing as a Clinical Test
Tuesday, May 7
7:30-8:15 Breakfast Presentation Sponsored by
8:25-10:00 Biomarkers for Safety Assessment Choosing a Platform for
Companion Diagnostics
Advances in IHC: Guiding
Therapy Decisions
Choosing a Platform for
Companion Diagnostics
10:00-11:00 Coffee Break in the Exhibit Hall with Poster Viewing
11:00-12:15 Biomarker Collaborations and Consortia Multiplexed Assays Tissue Biomarkers for Targeted Therapy Panel Discussion: Next-Generation
CDx Platforms
12:15-1:45 Lunch on Your Own and Conference Registration for Tracks 5-7
Track 5: Biomarkers for
Patient Selection
Track 6: Cancer Drug Resistance Track 7: Exosomes and Microvesicles
as Biomarkers and Diagnostics
12:15-1:45 Conference Registration
1:45-2:40 Biomarkers to Diagnostics Exosome Biomarkers in
Drug Development
Biomarkers to Diagnostics
2:40-3:45 Refreshment Break in the Exhibit Hall with Poster Viewing
3:45-5:30 Molecular Profiling of Tumor Heterogeneity to Guide Therapy Exosome Biomarkers in
Drug Development
Timeline for CDx Development
6:00-9:00 Dinner Course (Separate registration required)
Laboratory-Developed Tests
Wednesday, May 8
7:30-8:15 Breakfast Presentation (Sponsorship Opportunity Available) or Morning Coffee
8:25-10:30 Advancing Personalized Medicine 8:05 Secondary Resistance to Targeted
Cancer Therapy
Exosomes as Disease Markers Advancing Personalized Medicine
10:30-11:30 Coffee Break in the Exhibit Hall with Poster Viewing
11:30-12:45 Advancing Personalized Medicine 11:30-1:15 Resistance to
Various Therapies: Cancer Does
Not Discriminate
11:30-1:15 Exosomes as Novel
Cancer Biomarkers
Advancing Personalized Medicine
12:45 Close of Conference 1:15 Close of Conference 1:15 Close of Conference Close of Conference
*Executive pricing registration required
Conference-at-a-Glance
BiomarkerWorldCongress.com Biomarkers & Diagnostics World Congress | 3
BIOMARKERS & DIAGNOSTICS
world congress 2013
Distinguished Faculty
Jason M. Aliotta, M.D., Assistant Professor,
Medicine, Warren Alpert Medical School,
Brown University
John L. Allinson, FIBMS, Vice President,
Biomarker Laboratory Services, ICON
Development Solutions
Maria E. Arcila, M.D., Department of Pathology,
Memorial Sloan-Kettering Cancer Center
Khusru Asadullah, M.D., Vice President and
Head, Global Biomarkers, Bayer Pharma AG
Jiri Aubrecht, Pharm.D., Ph.D., Senior Director,
Safety Biomarker Group Lead, Drug Safety
Research & Development, Pfizer
M.J. Finley Austin, Ph.D., Personalized
Healthcare & Biomarker Strategy
Director, AstraZeneca
Nazneen Aziz, Ph.D., Director, Molecular
Medicine, Transformation Program Office,
College of American Pathologists
Geoffrey Stuart Baird, M.D., Ph.D., Assistant
Professor, Laboratory Medicine, University
of Washington
Robert A. Beckman, M.D., External Faculty,
Center for Evolution and Cancer, Helen Diller
Family Cancer Center, UCSF; Executive
Director, Clinical Development Oncology,
Daiichi Sankyo Pharma Development
Darrell R. Borger, Ph.D., Co-Director,
Translational Research Laboratory,
Massachusetts General Hospital Cancer Center
Mark Broenstrup, Ph.D., Director, Biomarker
and Diagnostics, R&D Diabetes Division, Sanofi
Michael Burczynski, Ph.D., Executive
Director, Biomarker Technologies, Discovery
Medicine and Clinical Pharmacology,
Bristol-Myers Squibb
Claudio Carini, M.D., Global Clinical
Immunology and Biomarkers Lead,
Bioenhancement Development Unit, Pfizer
Luigi Catanzariti, Ph.D., Executive Director and
Global Program Director, Diagnostics, Novartis
David Chen, Ph.D., Senior Director, Correlative
Sciences, Oncology Clinical Development,
Novartis Pharmaceuticals
Carol Cheung, M.D., Ph.D., Department of
Pathology, University Health Network
Atish Choudhury, M.D., Instructor in
Medicine, Medical Oncology, Dana-Farber
Cancer Institute
Seth Crosby, M.D., Director, Partnerships
& Alliances, Washington University School
of Medicine
Mark E. Curran, Ph.D., Vice President,
Immunology Biomarkers, Janssen Research &
Development
Stephen P. Day, Ph.D., Director, Medical
Affairs, Hologic
Viswanath Devanarayan, Ph.D., Global Head,
Exploratory Statistics, AbbVie, Inc
Luis Alberto Diaz, M.D., Associate Professor
of Oncology, Johns Hopkins Sidney Kimmel
Comprehensive Cancer Center
Max Diem, Ph.D., Professor, Chemistry and
Chemical Biology, Northeastern University
Nicholas C. Dracopoli, Ph.D., Vice President,
Janssen R&D, Johnson & Johnson
Crislyn D’Souza-Schorey, Ph.D., Professor,
Biological Sciences, University of Notre Dame
Dominik Duelli, Ph.D., Assistant Professor,
Cellular and Molecular Pharmacology, Rosalind
Franklin University of Medicine & Science,
Chicago Medical School
Daniel Edelman, Ph.D., Facility Head, Clinical
Molecular Profiling Core, National Cancer
Institute, NIH
Reyna Favis, Ph.D., Scientific Director,
Clinical Research & Development, Janssen
Pharmaceutical Companies of Johnson &
Johnson
Andrea Ferreira-Gonzalez, Ph.D., Professor
and Chair, Division of Molecular Diagnostics;
Director, Molecular Diagnostics Laboratory,
Department of Pathology, Virginia
Commonwealth University
Andrew Fish, Executive Director, AdvaMedDx
Herbert A. Fritsche, Ph.D., Senior
Vice President and CSO, Health
Discovery Corporation
Felix Frueh, Entrepreneur-in-Residence, Third
Rock Ventures
Steve Furlong, Ph.D., Safety Science Lead,
AstraZeneca Clinical Development
George A. Green IV, Ph.D., Director,
Pharmacodiagnostics, Bristol-Myers Squibb
Patrick Groody, Ph.D., Divisional Vice President,
Research & Development, Abbott
Steven Gutman, M.D., MBA, Strategic
Advisor, Myraqa
Abdel Halim, Pharm.D., Ph.D., DABCCMDx,
DABCC-TOX, DABCC-CC, FACB,
Director, Clinical Biomarkers, Daiichi Sankyo
Pharma Development
Sam Hanash, M.D., Ph.D., Director, McCombs
Institute for Cancer Early Detection and
Treatment, MD Anderson Cancer Center
Charles R. Handorf, M.D., Ph.D., Professor
and Chair, Pathology and Laboratory Medicine,
University of Tennessee
Amir Handzel, Ph.D., Associate Director,
Translational and Clinical Sciences, OSI
Pharma (Astellas)
Chunhai “Charlie” Hao, M.D., Ph.D., Associate
Professor, Neuropathology Attending,
Department of Pathology & Laboratory
Medicine, Emory University School of Medicine
Madhuri Hegde, Ph.D., Associate Professor,
Human Genetics; Senior Director, Emory
Genetics Laboratory, Emory University
Philip Hewitt, Ph.D., Head, Early Non-Clinical
Safety (Liver and Kidney), Merck Serono
Stephen M. Hewitt, M.D., Ph.D., Clinical
Investigator, Laboratory of Pathology, National
Cancer Institute, NIH
Holly Hilton, Ph.D., Head, Disease and
Translational Genomics, Hoffmann-La Roche;
Adjunct Professor, University of Medicine and
Dentistry New Jersey
Fred H. Hochberg, M.D., Associate Professor,
Neurology, Massachusetts General Hospital
Shidong Jia, Ph.D., Scientist, Oncology
Biomarker Development, Genentech
Chris Jowett, General Manager, Commercial
Operations, Abbott Molecular
Jingfang Ju, Ph.D., Co-Director of Translational
Research, Pathology, Stony Brook University
Peter M. Kazon, General Counsel, American
Clinical Laboratory Association
Eric Lai, Ph.D., Senior Vice President
and Head, Pharmacogenomics, Takeda
Pharmaceuticals International
Ira M. Lubin, Ph.D., Team Lead, Genetics
Laboratory Research and Evaluation
Branch, CDC
Johan Luthman, D.D.S., Ph.D., Senior Program
Leader, Neuroscience & Ophthalmology
Research & Development Franchise
Integrator, Merck
Elaine Lyon, Ph.D., Medical Director, Molecular
Genetics, ARUP Laboratories
Ron Mazumder, Ph.D., MBA, Global Head,
Research and Product Development, Janssen
Diagnostics, Janssen Pharmaceutical Companies
of Johnson & Johnson
Duncan McHale, Ph.D., Vice President, Global
Exploratory Development at UCB Pharma
Alan Mertz, President, American Clinical
Laboratory Association
Yoshi Oda, Ph.D., President, Biomarkers and
Personalized Medicine Core Function Unit, Eisai
Lorraine O’Driscoll, Ph.D., Associate Professor,
Pharmacology; Director, Research, School of
Pharmacy and Pharmaceutical Sciences, Trinity
College Dublin
Carol S. Palackdharry, M.D., MS, Medical
Director, ActiveHealth Management; Clinical
Lead, Oncology Condition Analysis, Aetna
Saumya Pant, Ph.D., Research Fellow, Merck
Liron Pantanowitz, M.D., Associate Professor,
Pathology and Biomedical Informatics,
University of Pittsburgh Medical Center
Scott D. Patterson, Ph.D., Executive Director,
Medical Sciences, Amgen
Sonia Pearson-White, Ph.D., Scientific
Program Manager, Oncology, The Biomarkers
Consortium, Foundation for the National
Institutes of Health
Emanuel Petricoin III, Ph.D., Co-Director, The
Center for Applied Proteomics and Molecular
Medicine, George Mason University
Suso Platero, Ph.D., Director, Oncology
Biomarkers, Janssen Pharmaceuticals
Mark Priebe, MT(ASCP)SBB, Managing
Director, QualityStar Quality Consortium
Debra Rasmussen, MBA, Senior Director,
Regulatory Affairs, Johnson & Johnson
Hallgeir Rui, M.D., Ph.D., Professor, Cancer
Biology, Medical Oncology, and Pathology,
Thomas Jefferson University
Hakan Sakul, Ph.D., Executive Director and
Head, Diagnostics, Worldwide R&D, Clinical
Research and Precision Medicine, Pfizer
Kurt A. Schalper, M.D., Ph.D., Associate
Research Scientist, Pathology, Yale School
of Medicine
Stephen C. Schmechel, M.D., Ph.D., Associate
Professor, Pathology, University of Washington
School of Medicine
Robert Schupp, Ph.D., Executive Director,
Diagnostics Hematology/Oncology,
Celgene Corporation
Jason S. Simon, Ph.D., Director, Immuno-
Oncology Biomarkers, Discovery Medicine and
Clinical Pharmacology, Bristol-Myers Squibb
Sharon Sokolowski, Ph.D., Principal Scientist,
Pfizer Global Research & Development
Gyongyi Szabo, M.D., Ph.D., Professor,
Gastroenterology, University of Massachusetts
Medical School
Douglas D. Taylor, Ph.D., Professor, Obstetrics
and Gynecology, University of Louisville School
of Medicine
Meghna Das Thakur, Ph.D., Presidential
PostDoctoral Fellow, Novartis Institutes for
BioMedical Research
Emina Torlakovic, M.D., Ph.D., Associate
Professor, Laboratory Medicine and
Pathobiology, University of Toronto
Jessie Villanueva, Ph.D., Assistant Professor,
Molecular & Cellular Oncogenesis Program,
The Wistar Institute
Glen J. Weiss, M.D., Co-Head, Lung Cancer
Unit, The Translational Genomics Research
Institute (TGen); Director, Thoracic Oncology,
Virginia G. Piper Cancer Center Clinical Trials
at Scottsdale Healthcare; CMO, CRAB-Clinical
Trials Consortium
David Wholley, Director, Biomarkers
Consortium, Foundation for the NIH
David T.W. Wong, D.M.D., D.M.Sc., Professor,
Associate Dean of Research, UCLA
School of Dentistry and Director of Dental
Research Institute
Janghee Woo, M.D., Albert Einstein
Medical Center
Wen Jin Wu, M.D., Ph.D., Principal Investigator,
Division of Monoclonal Antibodies, Office
of Biotechnology Products, Center for Drug
Evaluation and Research, FDA
Brenda Yanak, Ph.D., Precision Medicine
Leader, Clinical Innovation, Pfizer
Tammie Yeh, Ph.D., Molecular Biomarkers,
Oncology Lead, Merck
Eunhee S. Yi, M.D., Consultant, Anatomic
Pathology, Mayo Clinic; Professor, Pathology,
Mayo Clinic College of Medicine
Malcolm York, MPhil, Director and Head,
Clinical Pathology and Safety Assessment,
GlaxoSmithKline R&D
Theresa Zhang, Ph.D., Associate Director,
Exploratory and Translational Sciences, Merck
Premier Sponsor Corporate
Sponsors
4 | Biomarkers & Diagnostics World Congress BiomarkerWorldCongress.com
Monday, May 6, 6:00-9:00 pm
Fit-for-Purpose Biomarker Assay Development
and Validation
Instructors:
John L. Allinson, FIBMS, Vice President, Biomarker Laboratory Services, ICON
Development Solutions
Viswanath Devanarayan, Ph.D., Global Head, Exploratory Statistics, AbbVie, Inc
This tutorial will provide recommendations on the “fit-for-purpose” best
practices in the development and validation of biomarker assays for
exploratory or advanced biomarker applications. Strategies for different
applications at various phases of biomarker development will be described.
Key elements in the method of development and validation will be illustrated
with examples, including reference to standard material, sample stability and
collection integrity, validation and QC samples, validity of reference standards,
calibration curve fitting methods, method optimization and feasibility studies.
Special challenges in protein biomarker assays will be discussed, including
strategies for moving from biomarker panels in the exploratory phase to the
few markers chosen to support clinical trials, cross-validation of biomarker
assays, etc.
Outline:
1. Introduction: Nomenclature, types of biomarker methods/assays, method
development and validation road-map, fundamental validity, similarity and
differences from PK assays and diagnostic applications
2. Pre-analytical and bioanalytical elements: Target range, standards,
validation and QC samples, stability, matrix effect, specificity and
relative selectivity
3. Calibration curve model selection, evaluation and weighting
4. Method feasibility and optimization with precision profiles
5. Evaluation of some pre-study validation characteristics such as precision,
bias, sensitivity and quantification limits
6. Use of sample controls for in-study performance monitoring and
conformance testing among laboratories
7. Special considerations for multiplex assays, cross-validation of assays, etc.
8. Method comparisons
Next-Generation Sequencing as a Clinical Test:
It Takes a Community
Instructors:
Nazneen Aziz, Ph.D., Director, Molecular Medicine, Transformation Program
Office, College of American Pathologists
Madhuri Hegde, Ph.D., Associate Professor, Human Genetics; Senior Director,
Emory Genetics Laboratory, Emory University
Next-Generation Sequencing (NGS) is used widely in clinical research for
the discovery of disease-associated genes and the clinical community
is beginning to embrace this technology for diagnostic testing. The rapid
evolution of NGS technologies presents significant opportunities and
challenges for researchers and clinicians for improving health outcomes,
particularly with respect to an increased emphasis on personalized and
preventive medicine. Adoption of NGS in the clinical laboratory setting
requires the adoption of many processes and procedures, such as
the analytic and clinical validation of the test, CLIA certification/CAP
accreditation, standards for reference materials, availability for proficiency
testing, and questions regarding reimbursement and informed consent.
The success of NGS as a viable diagnostic modality depends on many
branches of the health care community working together. This session will
be informative and practical for the researcher and laboratorians who are
considering launching NGS as a clinical test.
Tuesday, May 7, 6:00-9:00 pm
Laboratory-Developed Tests
Regulatory Issues Facing Laboratory-Developed Tests
Peter M. Kazon, General Counsel, American Clinical Laboratory Association
The development of molecular diagnostics has been accompanied by a host
of regulatory issues, including coding, billing and FDA issues. This session will
review recent changes affecting these codes as well as the position of Medicare
on how to pay for these tests and tests including an algorithm, also referred
to as MAAAs. It will review the latest developments from the FDA concerning
whether such tests will require FDA pre-market approval or clearance, and what
action FDA is likely to take in the future. It will also review other actions that affect
these tests, such as the new MolDx program being overseen by Palmetto GBA, a
Medicare contractor.
Laboratory-Developed Tests in the Genomic Medicine Era: Validation, Regulation
and Challenges Faced by New Technologies and Clinical Applications
Andrea Ferreira-Gonzalez, Ph.D., Professor and Chair, Division of Molecular
Diagnostics; Director, Molecular Diagnostics Laboratory, Department of
Pathology, Virginia Commonwealth University
Laboratory-developed tests are those tests developed, validated and
performed by clinical laboratories. There are standards and regulations in
place for the validation of these tests before they are introduced into clinical
practice. This presentation will discuss the process of validation under the
current regulatory framework, and regulatory challenges posed by new
technologies such as NGS and its clinical applications.
LDTs in the Context of CLIA: An NIH Experience
Daniel Edelman, Ph.D., Facility Head, Clinical Molecular Profiling Core, National
Cancer Institute, NIH
The mission of the Clinical Molecular Profiling Core (CMPC) of the National
Cancer Institute (NCI) is to provide state of the art genomic testing for specimens
obtained from NCI clinical trials. The greatest impact is effected where test results
have immediate clinical application for personalized cancer care for individual
patients enrolled in these trials. To that end, the CMPC is CLIA certified and
provides a growing set of clinical test modalities. In this talk we’ll discuss the
challenges of meeting CLIA regulations in this new age of genomics at NIH for
high-complexity assays that did not exist as diagnostic tests when the federal
guidelines were written.
LDT Regulation Guidance from the FDA: Where Does It Stand after Three Years?
Stephen P. Day, Ph.D., Director, Medical Affairs, Hologic
The FDA’s announced intent to further regulate laboratory developed tests
(LDTs) enters its third year without the issuance of the anticipated guidance.
This presentation will summarize what the FDA has made publicly available
on the subject up to this time, the positions and recommendations put
forward by medical and industry professional societies, and how it will
potentially affect clinical laboratories offering LDTs and the delivery of quality
medical care.
Next-Generation Sequencing Assays as Laboratory-Developed Tests
Elaine Lyon, Ph.D., Medical Director, Molecular Genetics; Co-Medical Director,
Pharmacogenomics, ARUP Laboratories; Associate Professor, University of Utah
As next-generation sequencing (NGS) technologies improve in accuracy
and cost effectiveness, they will become standard in clinical laboratories.
Multi-gene panels, exome or genome analysis are alternative approaches.
With the complexity of genomic scale sequencing, implementing NGS
assays into clinical laboratories requires expertise in laboratory techniques,
informatics and interpretation. CLIA-certified clinical laboratories are
developing NGS assays as laboratory-developed tests (LDTs). The
presentation will discuss how NGS assays are “procedures” involving input
from health care professionals, and how they fit under the category of high
complexity LDTs.
*Separate registration required
Dinner Courses*
BiomarkerWorldCongress.com Biomarkers & Diagnostics World Congress | 5
Sunday, May 5
5:00-6:00 pm Conference Pre-Registration
Monday, May 6
7:30-8:30 am Conference Registration and Morning Coffee
8:30-8:40 Welcome Remarks from Conference Director
Julia Boguslavsky, Executive Director, Conferences, Cambridge Healthtech Institute
Biomarkers in Translational Medicine
8:40-8:45 Chairperson’s Opening Remarks
8:45-9:10 Translating Biological Data into Predictive Biomarker
Development Strategies
Nicholas C. Dracopoli, Ph.D., Vice President, Janssen R&D, Johnson & Johnson
A decade after completion of the human genome sequence, the translation
of complex genomic data into widely used clinical tests has been slower
than anticipated. Three complex tests (in vitro diagnostic multiplex index
assays – IVDMIA) have been approved as prognostic tests, but there has
still not been a single approval of an IVDMIA to predict response to therapy.
Retrospective analyses of the development of predictive biomarkers for
first-in-class oncology drugs over the last ten years shows that 1) insufficient
patients have been exposed to an efficacious dose to support complex
statistical analyses to correlate high-content data against clinical endpoints,
and 2) biomarkers that correlate to response in Phase II studies are not
always good predictors of overall survival in Phase III trials. We will need to
modify the clinical development paradigm for first-in-class agents to support
the efficient co-development of predictive markers.
9:10-9:35 Application of Next-Generation Sequencing in Phase III
Oncology Trials
David Chen, Ph.D., Senior Director, Oncology Correlative Sciences, Novartis
Analysis of tumor samples by next-generation sequencing (NGS) has
increased dramatically in the last 2 years. Most of the reported results
are genetic landscapes generated on samples collected outside clinical
trials or from early phase trials. Application of this technology in large
global Phase III trials provides an excellent opportunity for treatment
efficacy predictive biomarker explorations. Study design considerations
and analysis strategies for the implementation of complex and resource
demanding NGS analysis in Phase III trials will be discussed.
9:35-10:00 Can Biomarkers Recover Drug Development from the Ditch?
Abdel Halim, Pharm.D., Ph.D., DABCC-MDx, DABCC-TOX, DABCC-CC, FACB,
Director, Clinical Biomarkers, Daiichi Sankyo Pharma Development
Despite all the potential benefits of using biomarkers to advance the
pharmaceutical industry, discrepant results can pose a threat to development
programs by triggering false decisions. This talk will highlight the following
topics: biomarkers and their potential utility in drug development, limitations,
major reasons behind discrepant results and possibility of its mitigation.
10:00-10:30 Networking Coffee Break
10:30-10:55 Advancing Biomarkers for Alzheimer’s Disease—From
Target Engagement to Diagnostics
Johan Luthman, D.D.S., Ph.D., Senior Program Leader, Neuroscience &
Ophthalmology Research & Development Franchise Integrator, Merck
Measuring pathophysiology associated factors, such as Aβ peptide
and tau protein in cerebrospinal fluid, and imaging brain function with
fluorodeoxyglucose PET or functional MRI, or pathology with amyloid
PET or MRI, allows us to detect and follow the progression of very early,
pre-dementia stages of AD. While the use of pathophysiology associated
biomarkers allows pharmacodynamics monitoring of putative disease
modifying therapeutics, further qualification efforts are paving the way for
diagnostic and prognostic readouts.
10:55-11:20 Perspectives of Target and Marker Discovery in Multiple Myeloma
Robert Schupp, Ph.D., Executive Director, Diagnostics Hematology/Oncology,
Celgene Corporation
While major advances in therapeutic treatments have almost quadrupled the
overall survival of patients with multiple myeloma (MM) within the last decade,
the clinical challenges are now to optimize clinical utility of drugs and their
therapeutic sequence. The immunomodulatory drugs (IMiDs) thalidomide,
lenalidomide, and pomalidomide comprise an essential treatment modality in
MM and have shown to needfully bind to a newly identified protein in order
to exert their antimyeloma activity. Since this potential new target protein
may qualify as a biomarker predicting clinical response and the emergence of
resistance, significant challenges remain with regards to validating a robust
assay. This talk will highlight key challenges in the methodology and further
development of measurements leading to establishing a new biomarker in MM.
11:20-11:35 Measure for Measure: Assaying the Sponsored by
Rise and Fall of Protein Biomarkers across the Proteome
Steve Williams, M.D., Ph.D., CMO, SomaLogic
Shakespeare’s proteome: “Oh what may man within him hide, though angel on
the outward side!” – Early disease markers. “Our doubts are traitors and make
us lose the good we oft might win, by fearing to attempt” – Now is proteomics’
time. “Truth is truth to the end of reckoning” – Truth standards and biomarkers.
“The miserable have no other medicine but only hope” – Proteomics applied.
“What’s mine is yours, and what is yours is mine.” – Working with us.
11:35-11:50 Sponsored Presentation
(Opportunity available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
11:50-1:20 pm Luncheon Presentation Sponsored by
Obtaining NAT Sensitivity with ELISA: Results from
Application of Simoa to Blood Screening
David Wilson, Ph.D., Vice President, Product Development, Quanterix
Until recently, nucleic acid testing (NAT) represented the most sensitive
method for early acute HIV infection, when individuals are most
contagious. Using Single Molecule Arrays (Simoa), a digital ELISA
technique, researchers were able to demonstrate a 3000x sensitivity
improvement over conventional ELISA and equivalence with the NAT gold
standard but at a fraction of the cost. This ground-breaking research has
significant implications for blood banking, HIV detection and beyond.
Biomarker Utility in Clinical Development
1:20-1:25 Chairperson’s Remarks
1:25-1:50 Implementing Biomarkers in Clinical Trials
Suso Platero, Ph.D., Director, Oncology Biomarkers, Janssen Pharmaceuticals
Finding biomarkers is relatively easy nowadays. One has only to open
any journal and find dozens of articles showing the discovery of new
biomarkers. The bottleneck in the development of biomarkers is in the
correlation of the appropriate biomarkers to each specific drug. This is done
in the context of clinical trials. Several strategies will be presented of how to
better accomplish this task in an efficient and time sensitive manner.
1:50-2:15 Clinical Innovation in Precision Medicine
Brenda Yanak, Precision Medicine Leader, Clinical Innovation, Pfizer
This presentation will give examples of how Pfizer is innovating in the
clinical development space to aid in the advancement of precision medicine.
2:15-2:40 Discovering Oncology Biomarkers and Translating into
Clinical Trials
Theresa Zhang, Ph.D., Associate Director, Exploratory and Translational
Sciences, Merck
This talk will present a platform for discovering oncology response
biomarkers using a large panel of tumor cell lines, validating them in
selected in vivo models, and refining and estimating biomarker prevalence
in a large human tumor reference dataset. The predictive signature
will then be converted into an analytically validated assay that will be
performed in a CLIA- or CAP-certified laboratory in order to enroll patients
for clinical trials. The process will be illustrated by examples.
Track 1: Translational Biomarkers in Drug Development
6 | Biomarkers & Diagnostics World Congress BiomarkerWorldCongress.com
2:40-3:40 Refreshment Break in the Exhibit Hall with Poster Viewing
3:40-4:05 Biomarker Discovery for Immuno-Oncology Agents
Jason S. Simon, Ph.D., Director, Immuno-Oncology Biomarkers, Discovery
Medicine and Clinical Pharmacology, Bristol-Myers Squibb
Tumor cells can use escape mechanisms to avoid or suppress the
natural immune response, ultimately resulting in tumor growth; in fact,
avoiding immune destruction is one of the emerging hallmarks of cancer.
Therefore, understanding and dismantling key immune escape mechanisms
(“checkpoints”) is a key focus of immuno-oncology research. In concert with
identifying agents to regulate the immune checkpoint is working to understand
which tumor types and patient characteristics will respond best to this
treatment approach. This talk will review our strategy to identify biomarkers
which help support clinical development and commercialization strategies.
4:05-4:30 Accelerating and Personalizing Clinical Trials with
Biomarkers and Adaptive Design, the I-SPY 2 Example
Sonia Pearson-White, Ph.D., Scientific Program Manager, Oncology, The
Biomarkers Consortium, Foundation for the National Institutes of Health
I-SPY 2 is a unique clinical trial managed as a public/private partnership
by the Foundation for the NIH (FNIH) Biomarkers Consortium. I-SPY
2 employs an innovative adaptive trial design testing multiple drugs in
high-risk breast cancers in the neoadjuvant setting, and will advance the
understanding of which drugs work best with tumor types with different
biomarker profiles, and the drive toward personalized medicine.
4:30 Metabolomic Profiling for NMR Based Clinical Sponsored by
Assay Development
Thomas O’Connell, Ph.D., Senior Director, Assay Research &
Development, LipoScience, Inc.
Metabolomic profiling yields a unique picture of the downstream phenotype
taking into account genetic influences as well as environmental factors such
as diet, lifestyle and the microbiome. In this presentation it will be shown how
NMR technology is used in both the discovery and translation of biomarkers
into the clinical laboratory. Applications include the prediction, diagnosis and
prognosis of disease as well as the guidance of pharmaceutical interventions.
5:00-6:00 Networking Reception in the Exhibit Hall with Poster Viewing
6:00-9:00 Dinner Courses
Fit-for-Purpose Biomarker Assay Development and Validation
Next-Generation Sequencing as a Clinical Test
(Separate registration required. See Page 4 for additional information.)
Tuesday, May 7
7:30-8:15 am Breakfast Presentation Sponsored by
Identifying Non-Invasive Biomarkers of Smoking-
Related Parenchymal Lung Disease (i.e. COPD and
IPF) to Detect Subclinical Lung Disease
Ivan O. Rosas, M.D., Assistant Professor, Medicine Division, Pulmonary &
Critical Care Medicine, Brigham & Women’s Hospital, Harvard Medical School
Biomarkers for Safety Assessment
8:25-8:30 Chairperson’s Opening Remarks
8:30-8:55 Pre-Clinical Safety Biomarkers and Translation to Clinical
Testing: Current Perspectives
Malcolm York, MPhil, Director and Head, Clinical Pathology and Safety
Assessment, GlaxoSmithKline R&D
Significant efforts have been conducted into the analytical validation,
characterization and qualification of safety biomarkers, notably for the
assessment and prediction of cardiac, kidney and liver safety. However,
significant challenges are still evident in the interpretation of toxicity
biomarker signals above defined thresholds, which report apparent injury
(but in the absence of the histopathology) and their relevance for clinical
safety testing. Current perspectives on the measurement and use of toxicity
biomarkers/multiple biomarkers, along with integration with other measures
of biological response, to improve definition of safety margins and riskbenefit
characterization of exploratory new medicines will be discussed.
8:55-9:20 Biomarkers of Acute Kidney Injury: From Pre-Clinical Species
to Human Patients
Jiri Aubrecht, Pharm.D., Ph.D., Senior Director, Safety Biomarker Group Lead,
Drug Safety Research & Development, Pfizer
Acute kidney injury provides a significant challenge to drug development.
Recently, new biomarkers of acute kidney injury have been developed. In this
presentation we will review the recent progress in applying emerging biomarkers
of acute kidney injury across pre-clinical species and human subjects.
9:20-9:45 Preparing for Safety Biomarkers to Support Clinical Trials
Stephen T. Furlong, Ph.D., Safety Science Lead, AstraZeneca Patient Safety
Many new biomarkers are being considered for use in clinical trials to
monitor drug-induced organ toxicity. However, deciding which biomarkers
to use, selecting a vendor to perform the assays, establishing sample
handling protocols, preparing for statistical analysis of the data and
deciding how to use the data all represent significant challenges. This talk
will review these topics, provide examples from specific biomarkers and
provide suggestions for overcoming some of these challenges.
9:45-10:10 Identifying Biomarkers of Kidney and Liver Toxicity by
Integrating Toxicogenomics Datasets with Biological Networks
Philip Hewitt, Ph.D., Head, Early Non-Clinical Safety (Liver and Kidney), Merck Serono
Candidate nephrotoxicity biomarkers were identified by interrogating
profiles from hundreds of publicly available toxicogenomics datasets,
including datasets from the EU PredTox and Japanese TG-GATEs
projects. Application of multiple bioinformatics approaches identified
43 significant candidates. These findings were corroborated by testing
model nephrotoxic compounds using whole genome expression profile
experiments both in vivo and in vitro. This in silico approach greatly
enriched candidates for those likely to be true biomarkers.
10:10-11:00 Coffee Break in the Exhibit Hall with Poster Viewing
Biomarker Collaborations and Consortia
11:00-11:25 From Promise to Progress: An Update on the Biomarkers
Consortium
David Wholley, Director, Biomarkers Consortium, Foundation for the NIH
11:25-11:50 Open Innovation in Biomarker Discovery: Experiences
from Our Grants for Targets and Biomarkers Initiative
Khusru Asadullah, M.D., Vice President and Head, Global Biomarkers, Bayer
HealthCare
To combine expertise Bayer Healthcare has set up a novel open innovation
approach called Grants4Targets. After a review process, grants are
provided to perform focused experiments to further validate the proposed
targets/biomarkers. In addition to financial support, specific know-how on
target validation and drug discovery is provided. Experienced scientists
are nominated as project partners and, depending on the project, tools
or specific models are provided. More than 600 applications have been
received and 77 projects granted so far.
11:50-12:15 pm Biomarker Discovery—The Power of Collaborative
Networks
Duncan McHale, Ph.D., Vice President, Global Exploratory Development, UCB
Pharma
Clinically useful, predictive biomarkers have been very elusive despite
the growth of Big Biology. Individual technology solutions are commonly
touted as being able to identify drug response biomarkers but are rarely
successful. It is likely that to be successful a network of collaborators will
be needed bringing together technology discipline experts with disease
biology experts. A case example is given in rheumatoid arthritis.
12:15-1:45 Enjoy Lunch on Your Own
Track 1: Translational Biomarkers in Drug Development
BiomarkerWorldCongress.com Biomarkers & Diagnostics World Congress | 7
Track 2: Clinical Assay Development
Sunday, May 5
5:00-6:00 pm Conference Pre-Registration
Monday, May 6
7:30-8:30 am Conference Registration and Morning Coffee
8:30-8:40 Welcome Remarks from Conference Director
Julia Boguslavsky, Executive Director, Conferences, Cambridge Healthtech Institute
From Research Biomarkers to Clinical Assays
8:40-8:45 Chairperson’s Opening Remarks
8:45-9:10 Biomarkers and the Quest for Clinical Utility—Obstacles,
Challenges and Opportunities
Steven Gutman, M.D., MBA, Strategic Advisor, Myraqa
Over the past ten years there has been an explosive increase in the number
of biomarker assays available for the study and evaluation of human
disease. To ensure stakeholders are able to use this growing menu of tests
responsibly, there is a compelling need to understand the clinical utility of
these assays. Unfortunately a surprising number of tests are plagued by
inadequate information on clinical utility. This talk will focus on obstacles,
challenges and opportunities for addressing this problem.
9:10-9:35 Clinical Assay Development—The Process and Considerations
Herbert A. Fritsche, Ph.D., Senior Vice President and CSO, Health Discovery Corporation
The process for successful development of a clinical laboratory test begins
with a strict definition of the test concept and its clinical utility; design of an
accurate and robust assay for the analyte; analytical validation followed by
clinical validation; and lastly, translation of the new test from the research
lab to routine clinical use, which includes validation of the new test outside
of the research setting. Success of development is defined as acceptance of
the new test by the medical community as the “standard-of-care.”
9:35-10:00 Bridging Research and “Clinical” Assays in Pharmaceutical
Research & Development
John L. Allinson, FIBMS, Vice President, Biomarker Laboratory Services, ICON
Development Solutions
Many biomarker assays used in drug development are research assays
(i.e., not accredited diagnostic devices). This presentation will look at the
following: basic validation experiments across assays in research and
diagnostics; differences and assay evolution as methods progress through
different uses of results data; the requirements for accreditation of assays to
be used in diagnostics; and a brief look at the development of a companion
diagnostic and its implications from the laboratory perspective.
10:00-10:30 Networking Coffee Break
10:30-10:55 Key Considerations for Choosing and Transitioning a
Research Grade Assay to the Clinical Setting
Tammie C. Yeh, Ph.D., Molecular Biomarkers, Oncology Lead, Merck
Developing a biomarker assay with the clinical perspective in mind is critical to
the success of the biomarker. Identifying/choosing a robust biomarker readout
is as important as developing a robust analytical assay to ensure clinical utility.
It is important to understand the inherent biological variability as well as the
clinical feasibility of a biomarker readout, both of which will depend on tissue
type, tissue processing and the specific clinical setting. Both patient selection
and pharmacodynamic biomarkers will be addressed in this presentation.
10:55-11:20 Clinical Assay Development for Cancer Protein Biomarkers:
What Works and What Does Not Work
Samir Hanash, Ph.D., Program Head, Molecular Diagnostics, Fred Hutchinson
Cancer Research Center
The breadth and depth of proteomics technologies for the discovery of
biomarkers has increased substantially over the past decade, covering
a dynamic range of more than 7 logs in protein abundance. As a result,
numerous cancer biomarker candidates have emerged from discovery
studies. There remains a need for the development of high-throughput
technologies that allow testing the utility of these biomarkers for their
intended clinical application to meet regulatory requirements. Current
opportunities and challenges will be presented.
11:20-11:45 Will Regulation of Laboratory-Developed Tests Stifle Innovation?
Alan Mertz, President, American Clinical Laboratory Association
Laboratory developed tests (LDTs) are regulated by Federal law (CLIA),
state law, and industry standards established by the College of American
Pathologists. For many years, FDA has maintained that LDTs are medical
devices. FDA’s legal authority has been questioned, however, and Congress,
in July 2011, considered legislation that would enhance the CLIA framework
for regulating LDTs. FDA’s work plan for 2013 does not mention new
guidance on LDTs, but remains a possibility. Closing the LDT pathway would
have substantial effects on clinical laboratories, health care providers, and
patients. This presentation will examine the role of LDTs in creating new
tests, diagnosing rare diseases, and including the most up-to-date clinical
information in diagnostic tests.
11:45-1:20 pm Enjoy Lunch on Your Own
NGS in Clinical Use
1:20-1:25 Chairperson’s Remarks
1:25-1:50 College of American Pathologists’ Standards and Proficiency
Testing for Next-Generation Sequencing for the Clinical Laboratory
Nazneen Aziz, Ph.D., Director, Molecular Medicine, Transformation Program
Office, College of American Pathologists
The rapid and ongoing advances in the genetic test market, spurred by the
opportunities of Next-Generation Sequencing (NGS), necessitate many facets
of the health care industry to work cohesively. Adoption of NGS as a clinical test
requires the adoption of many processes and procedures, such as the analytic
and clinical validation of the test, CLIA certification/CAP accreditation, standards
for reference materials, availability for proficiency testing, genetic counseling,
and questions regarding reimbursement, informed consent and incidental
findings. This talk will focus on the laboratory requirements developed at CAP
for CLIA/CAP accreditation and the plans for proficiency testing for NGS.
1:50-2:15 Assuring the Quality of Next-Generation Sequencing in
Clinical Laboratory Practice
Ira M. Lubin, Ph.D., Team Lead, Genetics Laboratory Research and Evaluation
Branch, Division of Laboratory Science and Standards, Laboratory Science,
Policy, and Practice Program Office, Office of Surveillance, Epidemiology, and
Laboratory Services, Centers for Disease Control and Prevention
Integration of next-generation sequencing (NGS) into the clinical laboratory
requires test validation, establishment of quality control procedures, and
the independent assessment of test performance by proficiency testing
or alternate approaches. Existing regulatory requirements and professional
guidance do not adequately address these quality issues for clinical NGS
testing. This talk will describe the outcomes of a national workgroup
organized by the Centers for Disease Control and Prevention tasked to
identify principles and develop guidance to promote good clinical laboratory
practices for NGS and meet regulatory and professional standards.
2:15-2:40 Clinical NGS: Validation, Reporting and Economics
Seth Crosby, M.D., Director, Partnerships & Alliances, Washington University
School of Medicine
As NGS enters the clinic, matters of analytic and clinical validation are just
the start of the medical director’s worries. How should results be quickly
generated and communicated to a physician in a meaningful and actionable
manner? What are the new rules for billing and reimbursement?
2:40-3:40 Refreshment Break in the Exhibit Hall with Poster Viewing
3:40-4:05 Exome Sequencing in a Clinical Setting to Guide Patient Care
Madhuri Hegde, Ph.D., Associate Professor, Human Genetics; Senior Director,
Emory Genetics Laboratory, Emory University
8 | Biomarkers & Diagnostics World Congress BiomarkerWorldCongress.com
Track 2: Clinical Assay Development
Advances in genomic medicine have made it necessary for clinical laboratories
to rapidly implement new technologies to guide patient care. Exome
sequencing is rapidly being implemented across different specialties such as
inherited diseases, cancer and infectious diseases. This talk will focus on the
clinical utility of exome sequencing in patient care with real case examples.
4:05-4:30 Interpreting Clinical Next-Generation Sequencing Data:
Current Challenges and Hope for the Future
Elaine Lyon, Ph.D., Medical Director, Molecular Genetics; Co-Medical Director,
Pharmacogenomics, ARUP Laboratories; Associate Professor, University of Utah
With the complexity of genomic scale sequencing (next-generation
sequencing or NGS) and the massive amounts of data obtained, informatics
is essential. Two challenges in evaluating a variant are 1) is it real and 2)
is it clinically significant. Informatics allow alignment and variant calling
(differences from a reference sequence), and sifting of probable clinically
insignificant variants. More challenging is prioritizing variants that are likely
to be associated with the clinical symptoms. In addition to the symptomguided
analysis approach, NGS data can reveal variants in genes related to
drug metabolism that may affect efficacy or response. This presentation will
discuss approaches to prioritize symptom-related variants as well as the
potential of NGS data for companion diagnostics or therapeutics.
4:30-5:00 Sponsored Presentations
(Opportunities available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
5:00-6:00 Networking Reception in the Exhibit Hall with Poster Viewing
6:00-9:00 Dinner Courses
Fit-for-Purpose Biomarker Assay Development and Validation
Next-Generation Sequencing as a Clinical Test
(Separate registration required. See Page 4 for additional information.)
Tuesday, May 7
7:30-8:15 am Breakfast Presentation Sponsored by
Identifying Non-Invasive Biomarkers of Smoking-
Related Parenchymal Lung Disease (i.e. COPD and
IPF) to Detect Subclinical Lung Disease
Ivan O. Rosas, M.D., Assistant Professor, Medicine Division, Pulmonary &
Critical Care Medicine, Brigham & Women’s Hospital, Harvard Medical School
Choosing a Platform for Companion Diagnostics
8:25-8:30 Chairperson’s Opening Remarks
8:30-8:55 Validating Biomarker Assays as a Prelude to Companion
Diagnostic Development: Emerging Platform-Specific Considerations
Michael Burczynski, Ph.D., Executive Director, Biomarker Technologies, Discovery
Medicine and Clinical Pharmacology, Bristol-Myers Squibb
Timely implementation of companion diagnostics alongside therapeutic
products has amplified the need to validate predictive biomarkers in earlier
phases of drug development. Today, biomarker strategies are more complex
and require more diverse platforms than ever before. Ensuring that analytical
validation strategies for these exploratory predictive biomarker assays
are aligned with the downstream requirements for full-blown companion
diagnostic development is a critical activity that ultimately helps determine
the efficiency with which targeted medicines can be brought to market.
8:55-9:20 Choosing a Platform for Companion Diagnostic Development
Ron Mazumder, Ph.D., MBA, Global Head, Research and Product Development,
Janssen Diagnostics, Janssen Pharmaceutical Companies of Johnson & Johnson
One of the early considerations in developing a companion diagnostic is
choice of platform. Several factors, such as technical performance, regulatory
and reimbursement path, and commercial access will be discussed in this
context. Examples from the literature and case studies will be presented.
9:20-9:45 Thoughts and Considerations for Choosing a Companion
Diagnostic Technology and Platform Delivery System
Patrick Groody, Ph.D., Divisional Vice President, Research & Development, Abbott
Choosing a diagnostic technology and testing platform for the
development of a companion diagnostic test can be a significant challenge.
A wide variety of factors including the development time, capabilities of
potential partners and the ability of laboratories and physicians to access
and perform the test routinely in a clinical setting are key factors in
developing a companion diagnostic program. This talk will focus on variety
of strategies for developing commercial companion diagnostic tests.
9:45-10:00 Sponsored Presentation
(Opportunity available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
10:00-11:00 Coffee Break in the Exhibit Hall with Poster Viewing
Multiplexed Assays
11:00-11:25 Measurement of Telomere Repeats in Human Cancer Cell Lines
and Tissues Using a Monochrome Multiplex Quantitative PCR Assay
Daniel Edelman, Ph.D., Facility Head, Clinical Molecular Profiling Core, National
Cancer Institute, NIH
This talk will describe our efforts for the development and validation of a
QPCR multiplex assay to enable the quantitation of overall telomere length
(TL) in cancerous cell lines and tissues. A TL pattern between cancers
might provide valuable diagnostic or prognostic information to promote
a better understanding of the molecular or pathogenic characteristics of
specific cancer types.
11:25-11:50 Multiplexed Immunoassays on Formalin-Fixed, Paraffin-
Embedded Tissue Homogenates as Cancer Diagnostics
Geoffrey Stuart Baird, M.D., Ph.D., Assistant Professor, Laboratory Medicine,
University of Washington
Multiplex immunoassays (MIs) performed on formalin-fixed, paraffinembedded
(FFPE) tissue homogenates offer several advantages
over immunohistochemistry as cancer diagnostics. In contrast to
immunohistochemistry, MIs offer absolute quantitation and improved
sensitivity and specificity through the use of sandwich assay geometries.
Moreover, MI instrumentation has already been adopted in the clinical
laboratory, and is much less expensive than a mass spectrometer. MIs have
been validated as a clinical diagnostic for pituitary adenoma classification in
FFPE tissue, with current work focused on breast carcinoma.
11:50-12:05 pm Diagnostic Classifiers for the Detection Sponsored by
of Bladder Cancer
Mark Ruddock, Ph.D., Team Leader, Molecular Biology, Randox
Pharma Services
Patients presenting with hematuria require investigations, including
cystoscopy and imaging of their upper urinary tracts, to identify the source
of bleeding. This is a significant health burden, which is set to increase
because of our aging population. Using biochip array technology, we have
identified diagnostic classifiers for detecting bladder cancer.
12:05-12:30 Development of Multiplexed Protein Pathway Activation
Mapping Clinical Assays for Personalized Cancer Therapy
Emanuel Petricoin III, Ph.D., Co-Director, The Center for Applied Proteomics and
Molecular Medicine, George Mason University
Cellular signaling pathways are a protein-based network, and the intended
drug effect is to disrupt aberrant protein phosphorylation-based enzymatic
activity, and epigenetic phenomena. The reverse-phase protein microarray
platform provides detailed information about the state of the cellular
“circuitry” from small samples. Measurements of dozens to hundreds of
specific phosphorylated proteins that represent most of the targets for
targeted therapeutics can be obtained at once from only a few thousand
cells. This information helps select specific therapy(ies) tailored to the
patient’s tumor activated protein “circuitry.”
12:30-1:45 Enjoy Lunch on Your Own
BiomarkerWorldCongress.com Biomarkers & Diagnostics World Congress | 9
Track 3: Cancer Tissue Diagnostics
Sunday, May 5
5:00-6:00 pm Conference Pre-Registration
Monday, May 6
7:30-8:30 am Conference Registration and Morning Coffee
8:30-8:40 Welcome Remarks from Conference Director
Julia Boguslavsky, Executive Director, Conferences, Cambridge Healthtech Institute
Whole-Slide Imaging and Digital Pathology
8:40-8:45 Chairperson’s Opening Remarks
8:45-9:10 Validation of Whole Slide Imaging in Pathology
Liron Pantanowitz, M.D., Associate Professor, Pathology and Biomedical
Informatics, University of Pittsburgh Medical Center
Validation of whole slide imaging (WSI) is important to ensure that digitized
slides are at least equivalent to that of glass slides. The College of American
Pathologists (CAP) Pathology and Laboratory Quality Center convened a
panel to recommend validation requirements for WSI systems to be used
for clinical diagnostic purposes employing a combination of evidence-based
evaluation of the literature, expert consensus and public commentary. The
recommendations are comprehensive and address technical, interpretation
components and administrative issues related to WSI in pathology providing
practical guidance for all types of laboratories who are using or plan to
utilize WSI systems for diagnostic clinical work. This session will educate
participants about WSI in pathology, the regulatory issues surrounding digital
pathology, and review the validation guidelines developed by the CAP.
9:10-9:35 New Applications Utilizing Whole Slide Digital Imaging
for Anatomic Pathology Inter- and Intra-Lab Peer Review and
Benchmarking Quality Assurance
Mark Priebe, MT(ASCP)SBB, Managing Director, QualityStar Quality Consortium
Although application of Whole Slide Digital Imagining (WSI) for primary
diagnosis is limited by the FDA at this time, WSI is a significant enabling
technology for anatomic pathology (AP) quality assurance (QA) initiatives
both inter- and intra-laboratory. This presentation will review current AP/
QA programs and the application of WSI to a novel approach of gaining
longitudinal benchmarking data for quality review. The presentation
will focus on understanding design requirements for development and
implementation, investment requirements, confidentiality considerations
and methods to encourage pathologist participation and acceptance.
9:35-10:00 Label-Free Infrared Spectral Histopathology: Diagnostics and
Prognostics
Max Diem, Ph.D., Professor, Chemistry and Chemical Biology, Northeastern University
Infrared spectral histopathology is a method in which the biochemical
composition of a histopathological sample is used, rather than
morphometric criteria, to diagnose disease. To this end, thousands of
infrared spectra are collected from pixels about 10 μm on edge, and
analyzed to produce spectral images that detect abnormality based on
variations in composition. The accuracy of this method is comparable to
multi-panel immunohistochemistry.
10:00-10:30 Networking Coffee Break
10:30-10:55 Tumor Heterogeneity Assessed by Immunohistochemistry
of Multiplexed Protein Biomarkers
Steve Schmechel, M.D., Ph.D., Associate Professor, Pathology, University of
Washington School of Medicine
Intratumoral heterogeneity of protein expression may be linked to the
biological aggressiveness of tumors and selection of therapies. Analytical
and statistical methods to quantify heterogeneity are needed, particularly
for multiplexed assays. This presentation will discuss novel methods to
measure tumor heterogeneity.
10:55-11:20 Application of WSI in Consensus Review for Clinical Trials
Stephen M. Hewitt, M.D., Ph.D., Clinical Investigator, Laboratory of Pathology,
National Cancer Institute, NIH
Whole Slide Imaging is an enabling technology within pathology, altering all
aspects of current practice. Consensus review processes for clinical trials have
previously been expensive, slow, and complicated by issues of reproducibility.
Whole Slide Imaging and distributed review overcome many of these issues,
and provide new opportunities that have previously not been feasible.
11:20-11:50 Sponsored Presentations
(Opportunities available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
11:50-1:20 pm Enjoy Lunch on Your Own
NGS in Clinical Use
1:20-1:25 Chairperson’s Remarks
1:25-1:50 College of American Pathologists’ Standards and Proficiency
Testing for Next-Generation Sequencing for the Clinical Laboratory
Nazneen Aziz, Ph.D., Director, Molecular Medicine, Transformation Program
Office, College of American Pathologists
The rapid and ongoing advances in the genetic test market, spurred by the
opportunities of Next-Generation Sequencing (NGS), necessitate many
facets of the health care industry to work cohesively. Adoption of NGS as
a clinical test requires the adoption of many processes and procedures,
such as the analytic and clinical validation of the test, CLIA certification/CAP
accreditation, standards for reference materials, availability for proficiency
testing, genetic counseling, and questions regarding reimbursement,
informed consent and incidental findings. This talk will focus on the
laboratory requirements developed at CAP for CLIA/CAP accreditation and
the plans for proficiency testing for NGS.
1:50-2:15 Assuring the Quality of Next-Generation Sequencing in
Clinical Laboratory Practice
Ira M. Lubin, Ph.D., Team Lead, Genetics Laboratory Research and Evaluation
Branch, Division of Laboratory Science and Standards, Laboratory Science,
Policy, and Practice Program Office, Office of Surveillance, Epidemiology, and
Laboratory Services, Centers for Disease Control and Prevention
Integration of next-generation sequencing (NGS) into the clinical laboratory
requires test validation, establishment of quality control procedures, and
the independent assessment of test performance by proficiency testing
or alternate approaches. Existing regulatory requirements and professional
guidance do not adequately address these quality issues for clinical NGS
testing. This talk will describe the outcomes of a national workgroup
organized by the Centers for Disease Control and Prevention tasked to
identify principles and develop guidance to promote good clinical laboratory
practices for NGS and meet regulatory and professional standards.
2:15-2:40 Clinical NGS: Validation, Reporting and Economics
Seth Crosby, M.D., Director, Partnerships & Alliances, Washington University
School of Medicine
As NGS enters the clinic, matters of analytic and clinical validation are just
the start of the medical director’s worries. How should results be quickly
generated and communicated to a physician in a meaningful and actionable
manner? What are the new rules for billing and reimbursement?
2:40-3:40 Refreshment Break in the Exhibit Hall with Poster Viewing
3:40-4:05 Exome Sequencing in a Clinical Setting to Guide Patient Care
Madhuri Hegde, Ph.D., Associate Professor, Human Genetics; Senior Director,
Emory Genetics Laboratory, Emory University
Advances in genomic medicine have made it necessary for clinical
laboratories to rapidly implement new technologies to guide patient care.
Exome sequencing is being rapidly being implemented across different
specialties such as inherited diseases, cancer and infectious diseases. This
talk will focus on the clinical utility of exome sequencing in patient care
with real case examples.
10 | Biomarkers & Diagnostics World Congress BiomarkerWorldCongress.com
Track 3: Cancer Tissue Diagnostics
4:05-4:30 Interpreting Clinical Next-Generation Sequencing Data:
Current Challenges and Hope for the Future
Elaine Lyon, Ph.D., Medical Director, Molecular Genetics; Co-Medical Director,
Pharmacogenomics, ARUP Laboratories; Associate Professor, University of Utah
With the complexity of genomic scale sequencing (next-generation
sequencing or NGS) and the massive amounts of data obtained, informatics
is essential. Two challenges in evaluating a variant are 1) is it real and 2)
is it clinically significant. Informatics allow alignment and variant calling
(differences from a reference sequence), and sifting of probable clinically
insignificant variants. More challenging is prioritizing variants that are likely
to be associated with the clinical symptoms. In addition to the symptomguided
analysis approach, NGS data can reveal variants in genes related to
drug metabolism that may affect efficacy or response. This presentation will
discuss approaches to prioritize symptom-related variants as well as the
potential of NGS data for companion diagnostics or therapeutics.
4:30-5:00 Sponsored Presentations
(Opportunities available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
5:00-6:00 Networking Reception in the Exhibit Hall with Poster Viewing
6:00-9:00 Dinner Courses
Fit-for-Purpose Biomarker Assay Development and Validation
Next-Generation Sequencing as a Clinical Test
(Separate registration required. See Page 4 for additional information.)
Tuesday, May 7
7:30-8:15 am Breakfast Presentation Sponsored by
Identifying Non-Invasive Biomarkers of Smoking-
Related Parenchymal Lung Disease (i.e. COPD and
IPF) to Detect Subclinical Lung Disease
Ivan O. Rosas, M.D., Assistant Professor, Medicine Division, Pulmonary &
Critical Care Medicine, Brigham & Women’s Hospital, Harvard Medical School
Advances in Immunohistochemistry:
Guiding Therapy Decisions
8:25-8:30 Chairperson’s Opening Remarks
8:30-8:55 Quality Assurance/Quality Control for Immunohistochemistry
in the Era of Personalized Medicine
Emina Torlakovic, M.D., Ph.D., Associate Professor, Laboratory Medicine and
Pathobiology, University of Toronto
Immunohistochemistry (IHC) enables in situ detection of protein expression
(right tissue, right cells, right cellular compartment) and evaluation of expression
levels. Biomarker discovery increases demands on biomarker testing by IHC.
IHC incorporates >20 parameters and requires expert interpretation. Key
challenges include clinical trial study design, tissue processing parameters
and parameters related to expert interpretation. IHC testing challenges remain
significant due to widely spread lack of awareness that IHC QA/QC needs to
evolve to match IHC intended use in personalized medicine.
8:55-9:20 Detection of ALK Gene Rearrangement (ALK+) in Non-Small
Cell Lung Cancers
Eunhee S. Yi, M.D., Consultant, Anatomic Pathology, Mayo Clinic; Professor,
Pathology, Mayo Clinic College of Medicine
Currently, ALK FISH is regarded as the gold standard to select the ALK+
patients eligible for crizotinib therapy, and FISH confirmation is required for
“on-label” crizotinib treatment. ALK IHC can be useful to limit the number
of patients to be tested for ALK FISH by identification of a high probability
population whose tumors are likely to be ALK+. Current status of ALK IHC will
be reviewed along with the data from a molecular study on discordant cases
for ALK status by ALK IHC and FISH in a Mayo Clinic Lung Cancer Cohort.
9:20-9:45 Molecular Profiling and Immunohistochemistry: The Interface
for Identification of Tissue of Origin in Occult Primary Cancers
Charles R. Handorf, M.D., Ph.D., Professor and Chair, Pathology and Laboratory
Medicine, University of Tennessee
Metastatic tumors with an uncertain primary site can be a difficult clinical
problem. In thousands of patients every year, no confident diagnosis is ever
issued making standard-of-care treatment difficult. Newer gene expression
profiling (GEP) tests currently available to analyze these difficult-to-diagnose
tumors are now being compared head-to-head with immunochemistry (IHC),
which has long been held as a gold standard. The interface between these
techniques will be discussed and practical approaches will be explored.
9:45-10:00 Sponsored Presentation
(Opportunity available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
Tissue Biomarkers for Targeted Therapy
10:00-11:00 Coffee Break in the Exhibit Hall with Poster Viewing
11:00-11:25 In situ Measurement of Tissue Biomarkers for Companion
Diagnostics in Cancer
Kurt A. Schalper, M.D., Ph.D., Associate Research Scientist, Pathology, Yale
School of Medicine
Measurement of tissue biomarkers has been shown to be a valuable tool
for companion diagnostics and is an essential component of personalized
cancer medicine. Several technical limitations surround commonly used
testing methods. In situ measurement of protein and mRNA transcripts
using automated quantitative immunofluorescence and novel hybridization
techniques provides increased sensitivity, specificity and reproducibility.
More quantitative approaches could open new opportunities for biomarker
discovery and patient selection for anti-cancer treatments.
11:25-11:50 Biomarkers and Targeted Therapy for Kaposi Sarcoma
Liron Pantanowitz, M.D., Associate Professor, Pathology and Biomedical
Informatics, University of Pittsburgh Medical Center
Kaposi sarcoma (KS) is an enigmatic vascular neoplasm that arises from
the initial infection of an endothelial or progenitor cell by Kaposi Sarcoma
Herpesvirus/Human Herpesvirus-8 (KSHV/HHV8). KS represents an ideal
model to investigate the interplay between viral oncogenesis, angiogenesis
and host immunity. The discovery of KSHV and related data about the
pathogenesis of KS has resulted in the identification of multiple novel
therapeutic targets. This talk will educate participants about KS biomarkers
being applied for diagnostic work, and also address newer therapeutic
agents aimed at molecular targets being evaluated in clinical trials.
11:50-12:15 pm Access to Human Tissue in the Age of Targeted
Therapies—Impact on Patient Care and Drug Development
Carol Cheung, M.D., Ph.D., Department of Pathology, Canadian University Health
Network
Access to human tissue is paramount in this age of targeted therapies.
Demand for this biological substrate, which is necessary for development
of innovative new tests and potentially blockbuster new therapies, is ever
increasing. The distinction between the two broad classes of excised human
tissue, research tissue that resides in research biobanks and diagnostic
tissue that resides in the clinical archives of institutional departments of
pathology, is important because the rules governing access differ depending
on this fundamental classification.
12:15-1:45 Enjoy Lunch on Your Own
BiomarkerWorldCongress.com Biomarkers & Diagnostics World Congress | 11
Track 4: Executive Summit: Companion Diagnostics
Sunday, May 5
5:00-6:00 pm Conference Pre-Registration
Monday, May 6
7:30-8:30 am Conference Registration and Morning Coffee
8:30-8:40 Welcome Remarks from Conference Director
Julia Boguslavsky, Executive Director, Conferences, Cambridge Healthtech Institute
Commercialization of Companion Diagnostics
8:40-8:45 Chairperson’s Opening Remarks
8:45-9:10 Companion Diagnostic Success: Biomarker Discovery to
Global Commercialization
Chris Jowett, General Manager, Commercial Operations, Abbott Molecular
Developing a successful global commercialization strategy for a companion
diagnostic can be a significant challenge. Critical capability factors need
to be discussed prior to entering into the partnership to minimize risk.
Understanding the IVD manufacturers’ capabilities to develop, manage the
required clinical trials, navigate the regulatory environment for approval,
and drive sales and marketing efforts in all targeted countries for the
therapeutic launch is essential. This talk will focus on a variety of strategies
to support a successful launch of a companion diagnostic program.
9:10-9:35 The Payor’s Role in Personalized Medicine
Carol S. Palackdharry, M.D., MS, Medical Director, ActiveHealth Management;
Clinical Lead, Oncology Condition Analysis, Aetna
Targeted cancer treatment is already changing the standard of care for many
cancers. Personalized therapies are costly and generally have anti-tumor activity
only in patients with the specific targeted abnormality. Most targeted agents
require pre-certification, with coverage dependent on appropriate results on
approved companion diagnostic tests. Development of rigorous, evidencebased
recommendations for usage of such tests, as well as new contracting
strategies with high-quality laboratories, will avoid wasted expenditures and
assure access to personalized therapies for all qualified patients.
9:35-10:00 Meeting Evidence Demands for Diagnostics in an Evolving
Payment Environment
Andrew C. Fish, Executive Director, AdvaMedDx
Payer reimbursement of diagnostics is critical to ensuring a robust
market for innovation. As advanced molecular diagnostics proliferate, a
growing appreciation of the importance of these tests is tempered by
rising payer concerns about coding transparency, evidence of clinical
utility, and utilization of and payment for these tests. This talk will review
the reimbursement challenges faced by test developers and initiatives
underway by payers and in Congress to address those challenges.
10:00-10:30 Networking Coffee Break
10:30-10:55 Creating a Companion Diagnostic Regulatory Strategy:
Biomarker to Commercial Test
Debra Rasmussen, Senior Director, Regulatory Affairs, Johnson & Johnson
Validated biomarkers (diagnostic tests) that can serve as intermediate or
surrogate endpoints to acquire rapid regulatory approval are needed to
help move research into the clinic. This is especially true if such biomarkers
could be measured easily, rapidly and were generally accessible.
Pharmaceutical companies could gain from biomarkers and diagnostic
co-development efforts. In an increasingly challenging regulatory
environment, diagnostic led treatment can improve the chance that drugs
are reimbursed or approved in the first place. As companion diagnostics
these could also potentially identify patient benefits from a novel
therapeutic strategy earlier, assist in early discontinuation of ineffective
strategies, and identify active drugs more efficiently. New concerns could
include: 1) designing a definitive clinical study for a joint therapeutic–
diagnostic that allows for assessment of the therapeutic’s safety and
efficacy, as well as for validation of the clinical utility of the biomarker
guiding the therapeutic’s use or 2) regulatory bodies requiring a diagnostic
test before a prescription may be written for a patient.
10:55-11:25 Sponsored Presentations
(Opportunities available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
11:25-11:50 Panel Discussion: Strategies for Regulatory and
Reimbursement Challenges in Commercialization of CDx
Panelists:
Chris Jowett, General Manager, Commercial Operations, Abbott Molecular
Carol S. Palackdharry, M.D., MS, Medical Director, ActiveHealth Management;
Clinical Lead, Oncology Condition Analysis, Aetna
Debra Rasmussen, Senior Director, Regulatory Affairs, Johnson & Johnson
Andrew C. Fish, Executive Director, AdvaMedDx
11:50-1:20 pm Enjoy Lunch on Your Own
Strategies for Rx-Dx Partnerships
1:20-1:25 Chairperson’s Remarks
1:25-1:50 Synchronizing Drug Development and Companion
Diagnostics: Challenges and Solutions
Hakan Sakul, Ph.D., Executive Director and Head, Diagnostics, Worldwide R&D,
Clinical Research and Precision Medicine, Pfizer
Last year witnessed simultaneous regulatory approvals of Rx and Dx and
it is expected that such approvals will be more commonplace in the near
future. Synchronizing the drug development phases with those for Dx
development presents many challenges. This talk will attempt to outline
these challenges and offer solutions based on the Xalkori Rx/Dx program.
1:50-2:15 Managing Pharma/Diagnostic Partnerships in Companion
Diagnostic Development
George A. Green IV, Ph.D., Director, Pharmacodiagnostics, Bristol-Myers Squibb
The development of CDx assays minimally requires a partnership between a
pharmaceutical and a diagnostic company. It is not uncommon for the drug
to be developed through an alliance of two pharmaceutical companies, and
diagnostic assay development programs may include separate companies
for design of the assay and development of the platform. To ensure effective
delivery of the CDx within this complex environment, highly matrixed teams
must be formed to address strategic and technical issues, and to deliver a
quality product coordinated with the drug development schedule.
2:15-2:40 Presentation to be Announced
2:40-3:40 Refreshment Break in the Exhibit Hall with Poster Viewing
3:40-4:10 Key Considerations for Selecting a Diagnostic Sponsored by
Partner in Rx-Dx Program Commercialization
Jeremy Bridge-Cook, Ph.D., Senior Vice President, Research &
Development, Luminex
What are the optimal capabilities and expertise required of diagnostic
partners for the development and commercialization of companion
diagnostic devices? Key considerations include prototype assay
development, analytical and clinical validation, regulatory filing, approval
and market launch. The speaker will discuss how each of these elements
can impact the success of a companion diagnostic program.
4:10-4:25 Sponsored Presentation
(Opportunity available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
12 | Biomarkers & Diagnostics World Congress BiomarkerWorldCongress.com
Track 4: Executive Summit: Companion Diagnostics
4:25-5:00 Panel Discussion: Strategies for Initiating and Managing
Successful Rx-Dx Partnerships
Panelists:
Hakan Sakul, Ph.D., Executive Director and Head, Diagnostics, Worldwide R&D,
Clinical Research and Precision Medicine, Pfizer
George A. Green IV, Ph.D., Director, Pharmacodiagnostics, Bristol-Myers Squibb
Panelist to be Announced
5:00-6:00 Networking Reception in the Exhibit Hall with Poster Viewing
6:00-9:00 Dinner Courses
Fit-for-Purpose Biomarker Assay Development and Validation
Next-Generation Sequencing as a Clinical Test
(Separate registration required. See Page 4 for additional information.)
Tuesday, May 7
7:30-8:15 am Breakfast Presentation Sponsored by
Identifying Non-Invasive Biomarkers of Smoking-
Related Parenchymal Lung Disease (i.e. COPD and
IPF) to Detect Subclinical Lung Disease
Ivan O. Rosas, M.D., Assistant Professor, Medicine Division, Pulmonary &
Critical Care Medicine, Brigham & Women’s Hospital, Harvard Medical School
Choosing a Platform for Companion Diagnostics
8:25-8:30 Chairperson’s Opening Remarks
8:30-8:55 Validating Biomarker Assays as a Prelude to Companion
Diagnostic Development: Emerging Platform-Specific Considerations
Michael Burczynski, Ph.D., Executive Director, Biomarker Technologies,
Discovery Medicine and Clinical Pharmacology, Bristol-Myers Squibb
Timely implementation of companion diagnostics alongside therapeutic
products has amplified the need to validate predictive biomarkers in earlier
phases of drug development. Today, biomarker strategies are more complex
and require more diverse platforms than ever before. Ensuring that analytical
validation strategies for these exploratory predictive biomarker assays
are aligned with the downstream requirements for full-blown companion
diagnostic development is a critical activity that ultimately helps determine
the efficiency with which targeted medicines can be brought to market.
8:55-9:20 Choosing a Platform for Companion Diagnostic Development
Ron Mazumder, Ph.D., MBA, Global Head, Research and Product Development,
Janssen Diagnostics, Janssen Pharmaceutical Companies of Johnson &
Johnson
One of the early considerations in developing a companion diagnostic
is choice of platform. Several factors, such as technical performance,
regulatory and reimbursement path, and commercial access will be
discussed in this context. Examples from the literature and case studies
will be presented.
9:20-9:45 Thoughts and Considerations for Choosing a Companion
Diagnostic Technology and Platform Delivery System
Patrick Groody, Ph.D., Divisional Vice President, Research & Development,
Abbott
Choosing a diagnostic technology and testing platform for the
development of a companion diagnostic test can be a significant challenge.
A wide variety of factors including the development time, capabilities of
potential partners and the ability of laboratories and physicians to access
and perform the test routinely in a clinical setting are key factors in
developing a companion diagnostic program. This talk will focus on variety
of strategies for developing commercial companion diagnostic tests.
9:45-10:00 Sponsored Presentation
(Opportunity available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
10:00-11:00 Coffee Break in the Exhibit Hall with Poster Viewing
11:00-12:00 pm Panel Discussion: Next-Generation CDx Platforms
Panelists:
Michael Burczynski, Ph.D., Executive Director, Biomarker Technologies,
Discovery Medicine and Clinical Pharmacology, Bristol-Myers Squibb
Ron Mazumder, Ph.D., MBA, Global Head, Research and Product Development,
Janssen Diagnostics, Janssen Pharmaceutical Companies of Johnson & Johnson
Elaine Lyon, Ph.D., Medical Director, Molecular Genetics; Co-Medical Director,
Pharmacogenomics, ARUP Laboratories; Associate Professor, University of Utah
Patrick Groody, Ph.D., Divisional Vice President, Quality Assurance and
Operations, Abbott
12:00-1:45 Enjoy Lunch on Your Own
Biomarkers to Diagnostics
1:45-1:50 Chairperson’s Remarks
1:50-2:15 Investing in Biomarkers and Turning Them into Diagnostics
Felix Frueh, Entrepreneur-in-Residence, Third Rock Ventures
The translation of biomarkers into useful clinical diagnostics requires the
demonstration of clinical benefit and cost effectiveness. Investing in new
technology is not sufficient without the realization that discovery and
development of a new biomarker needs to include the demonstration that
the biomarker makes a difference in clinical outcomes or decision-making,
preferably tested in the environment in which ultimately a diagnostic will
be used.
2:15-2:40 From Biomarker Research to Diagnostic Development—Our
Challenges
Yoshi Oda, Ph.D., President, Biomarkers and Personalized Medicine Core
Function Unit, Eisai
Biomarkers play important roles for drug development as a part of
translational research. Several examples about biomarkers for 1) the
evidence of target engagement, 2) patient stratification, 3) drug efficacy
and 4) disease diagnostics will be discussed.
2:40-3:45 Refreshment Break in the Exhibit Hall with Poster Viewing
Timeline for CDx Development
3:45-3:50 Chairperson’s Remarks
3:50-4:15 Timeline Considerations for Incorporating a Companion
Diagnostic into the Drug Development Process
Luigi Catanzariti, Ph.D., Executive Director and Global Program Director,
Diagnostics, Novartis
Rx/Dx co-development provides new opportunities for Pharma with respect to
targeted therapeutics. It also comes with considerable clinical, technical and
regulatory challenges. While both drug and diagnostic development processes
have their own rules and regulations, this new codependency requires
significant adjustments in what can be considered quintessentially clinical
(Rx) and technical (Dx) development cultures. Mutual understanding and
integration of both cultures early in the development process is an important
aspect for minimizing development timelines and achieving success.
4:15-4:40 Nothing Ventured, Nothing Gained: The Timeline Challenge
for Companion Diagnostics
Scott Patterson, Ph.D., Executive Director, Medical Sciences, Amgen
The identification of patients who are most likely to benefit from therapy
is an important component of any drug development strategy. Other
than when the target of the therapeutic is also the diagnostic for patient
selection, the generation of evidence to test a biomarker patient selection
hypothesis occurs during the drug development process. That data may
not become available until late in the development process. Strategies that
could be pursued to address this issue, with examples, will be presented.
BiomarkerWorldCongress.com Biomarkers & Diagnostics World Congress | 13
Track 4: Executive Summit: Companion Diagnostics
4:40-5:05 Strategic and Computational Considerations in
Development of Complex Companion Diagnostics
Amir Handzel, Ph.D., Associate Director, Translational and Clinical Sciences, OSI
Pharma (Astellas)
Successful development of CDx requires special attention to diverse
factors, as well as to their seamless integration. These challenges in
developing validated complex diagnostic biomarkers have been highlighted
by several failures in the last decade. The universe of molecular entities
from which markers can be chosen is rich, comprising genetic mutations,
the transcriptome, proteins and emerging non-coding RNA and epigenetic
entities. Their extremely large numbers present difficult problems of
selection and validation in a statistically robust and consistent way.
In order to address them, an array of technical, as well as operational
and organizational approaches must be employed. For example, the
characteristics of the experimental platforms used to acquire data
influence biomarker selection and design and these in turn necessitate a
multidisciplinary team structure. I will discuss these strategic and technical
elements while pointing to pitfalls and how to avoid them to reach the
desired goal.
5:05-5:30 Companion Diagnostics: Challenges in Bridging the Chasm
between Diagnostics and Drugs
Steven Gutman, M.D., MBA, Strategic Advisor, Myraqa
An IVD companion diagnostic device is an in vitro diagnostic device
that provides information essential for the safe and effective use of a
corresponding therapeutic product. This pairing of products has generated
intense interest because 1) it offers a clear model for the implementation
of personalized health care and 2) it may contribute to more informed
choices about how to manage the pipeline for new drugs. This talk will
focus on potential roadmaps for use in drug-diagnostic co-development.
6:00-9:00 Dinner Course
Laboratory-Developed Tests
(Separate registration required. See Page 4 for additional information.)
Wednesday, May 8
7:30-8:15 am Breakfast Presentation or Morning Coffee
(Sponsorship opportunity available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
Advancing Personalized Medicine
8:25-8:30 Chairperson’s Opening Remarks
8:30-8:55 Personalized Health Care: ‘One Size Does Not Fit All’ Applies
to Patients and Products
M.J. Finley Austin, Ph.D., Personalized Healthcare & Biomarker Strategy
Director, AstraZeneca
The essence of Personalized Health Care (PHC) is identifying,
understanding and partioning drug response variation to improve clinical
outcomes. Existing PHC examples demonstrate diversity in source of
variation, path to market as well as market delivery and uptake. Current
examples will be used to elucidate the implications of differing sources
and degrees of variation, clinical utility, and timing of discovery all have for
clinical trial design, regulatory strategy and market delivery.
8:55-9:20 Molecular Subtyping of Patients for Drug Development
Eric Lai, Ph.D., Senior Vice President and Head, Pharmacogenomics, Takeda
Pharmaceuticals International
While the concept of drug-diagnostic co-development (CDx) has been
around for awhile, most companion diagnostics are still an afterthought
and not an integrated component of drug development. To benefit from
the full potential of CDx, we have to change the strategy of drug target
identification from the single target approach to systematic understanding
of a patient’s disease phenotypes. I will discuss some of the potentials
steps that we have made to the drug development process.
9:20-9:45 Co-Diagnostics in Autoimmune Disorders: Improving
Outcomes in RA and IBD
Mark E. Curran, Ph.D., Vice President, Immunology Biomarkers, Janssen
Research & Development
Rheumatoid arthritis and inflammatory bowel disease are severe immune
diseases with significantly reduced quality of life for patients. Despite
advances in treatment with the evolution of antibody and recombinant
protein based therapeutics, there remains a significant unmet clinical need
for new therapies and integrated treatment solutions. At Janssen we are
focused on transforming therapy in these diseases by applying systems
pharmacology, precision medicine principles and developing companion
diagnostics to create new treatment paradigms. Our objective is to
provide for higher response rates, deeper remission, early interception and
eventually prevention of these diseases. Progress toward these objectives
will be discussed.
9:45-10:15 Complex microRNA Signatures of Response Sponsored by
and Resistance as Powerful Biomarkers
E. Robert Wassman, M.D., CMO, Rosetta Genomics
10:15-10:30 Sponsored Presentation
(Opportunity available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
10:30-11:30 Coffee Break in the Exhibit Hall with Poster Viewing
11:30-11:55 Precision Medicine: Triumphs and Tribulations
Claudio Carini, M.D., Global Clinical Immunology and Biomarkers Lead,
Bioenhancement Development Unit, Pfizer
The current model for drug development is failing. Failures often occur
either during the phase II trials, where either the candidate drug did not
meet the expected pharmacological requirements or the targeted drug
mechanism did not play a role in the patients population studied. Thus,
a new “personalized medicine” strategy is needed to develop predictive
biomarkers to assist in the decision making process during the pre-clinical
phase of drug development and use biomarkers as companion diagnostics
for stratifying patients in hypothesis-driven clinical trials.
11:55-12:20 pm Towards Personalized Medicine in Metabolic Diseases
Mark Broenstrup, Ph.D., Director, Biomarker and Diagnostics, R&D Diabetes
Division, Sanofi
Currently, more than 346 million people worldwide have diabetes. The
identification of the most effective drug(s) for the individual patient
is guided by a few selection criteria and a trial-and-error approach.
Consequently, the introduction of personalized approaches, accounting
for the heterogeneity of the disease, is regarded as a key enabler for
improved health care. An overview on biomarkers for assessing risk,
monitoring disease progression and predicting response to drugs is
provided, with a focus on beta cell imaging and systems biology solutions.
Finally, major public-private partnerships aiming at personalized solutions in
diabetes will be highlighted.
12:20-12:45 Translating Molecular Targets for Cancer Therapeutics
Glen J. Weiss, M.D., Co-Head, Lung Cancer Unit, The Translational Genomics
Research Institute (TGen); Director, Clinical Research, Cancer Treatment
Centers of America; CMO, CRAB-Clinical Trials Consortium
The presentation will focus on why there is a push to individualize cancer
therapy, past failures and successes, and how to define the tumor context
of vulnerability (COV). The talk will also describe the steps from pre-clinical
to new drug application and show how to optimize the drug development
path with knowledge of biomarker-based COV.
12:45 Close of Conference
14 | Biomarkers & Diagnostics World Congress BiomarkerWorldCongress.com
Track 5: Biomarkers for Patient Selection
tuesday, May 7
12:15-1:45 Conference Registration
Biomarkers to Diagnostics
1:45-1:50 Chairperson’s Opening Remarks
1:50-2:15 Investing in Biomarkers and Turning Them into Diagnostics
Felix Frueh, Entrepreneur-in-Residence, Third Rock Ventures
The translation of biomarkers into useful clinical diagnostics requires the
demonstration of clinical benefit and cost effectiveness. Investing in new
technology is not sufficient without the realization that discovery and
development of a new biomarker needs to include the demonstration that the
biomarker makes a difference in clinical outcomes or decision-making, preferably
tested in the environment in which ultimately a diagnostic will be used.
2:15-2:40 From Biomarker Research to Diagnostic Development—Our
Challenges
Yoshi Oda, Ph.D., President, Biomarkers and Personalized Medicine Core
Function Unit, Eisai
Biomarkers play important roles for drug development as a part of
translational research. Several examples about biomarkers for 1) the
evidence of target engagement, 2) patient stratification, 3) drug efficacy
and 4) disease diagnostics will be discussed.
2:40-3:45 Refreshment Break in the Exhibit Hall with Poster Viewing
Molecular Profiling of Tumor Heterogeneity to
Guide Therapy
3:45-3:50 Chairperson’s Remarks
3:50-4:15 Liquid Biopsies to Monitor Response and Resistance to
Targeted Therapies
Luis Alberto Diaz, M.D., Associate Professor of Oncology, Johns Hopkins
Sidney Kimmel Comprehensive Cancer Center
The simplest hypothesis to account for the development of resistance
to EGFR blockade is that rare cells with KRAS mutations pre-exist at low
levels in tumors with ostensibly wild-type KRAS genes. Although this
hypothesis would seem readily testable, there is no evidence in preclinical
models to support it, nor is there data from patients. To test this
hypothesis, we determined whether mutant KRAS DNA could be detected
in the circulation of 28 patients receiving monotherapy with panitumumab,
a therapeutic anti-EGFR antibody. The results suggest that the emergence
of KRAS mutations is a mediator of acquired resistance to EGFR blockade
and that these mutations can be detected in a non-invasive manner. They
explain why solid tumors develop resistance to targeted therapies in a
highly reproducible fashion.
4:15-4:40 Application of Clinical Tumor Genotyping in Targeted Cancer
Therapy
Darrell R. Borger, Ph.D., Co-Director, Translational Research Laboratory,
Massachusetts General Hospital Cancer Center
Multiplexed tumor genotyping has been offered as a physician-ordered
clinical test at a major U.S. cancer center. Over 3,000 patients have been
evaluated and these new capabilities have fostered a genotype-directed
approach to clinical trial design. By testing a broad spectrum of tumor
types, new molecular signatures have been revealed and mechanisms
of de novo and acquired resistance to targeted therapies have been
uncovered. This has provided the foundation for expanding clinical cancer
genotyping approaches for personalizing cancer care.
4:40-5:05 Quantitative Tumor Protein Profiling for Therapy-Relevant
Stratification of Breast Cancer Patients
Hallgeir Rui, M.D., Ph.D., Professor, Cancer Biology, Medical Oncology and
Pathology; Scientific Director, Jefferson Breast Care Center; Program Leader,
Biology of Breast Cancer, Kimmel Cancer Center; Co-Director, Pathology
Translational Research Core, Thomas Jefferson University
Breast cancer is a heterogeneous group of malignancies driven by diverse
oncogenic pathways. Ongoing consortium efforts are to map breast cancer
subtypes at high resolution based on quantitative immunofluorescence
(QIF) profiling of druggable target proteins within carcinoma cells of a
panel of 5,000 untreated primary breast cancer specimens. Progress
with prolactin-receptor-Jak-Stat pathway profiling will be highlighted using
complementary QIF technologies. Utility of resulting protein-based breast
cancer subclassification maps for rational recruitment of patients into
biomarker-driven, adaptive clinical trials will be discussed.
5:05-5:30 Clinical Validation of Predictive Biomarkers and Next-
Generation Personalized Medicine Treatment Strategies Incorporating
Genetic Dynamics
Robert A. Beckman, M.D., External Faculty, Center for Evolution and Cancer,
Helen Diller Family Cancer Center, University of California at San Francisco;
Executive Director, Clinical Development Oncology, Daiichi Sankyo Pharma
Development
The future of oncology drug development lies in personalized therapy
using predictive biomarkers. However, examples of the failure of predictive
biomarkers also exist. In these cases the use of biomarkers increased
the costs, complexity and duration of clinical trials, and narrowed the
treated population unnecessarily. We present methods to adaptively
integrate predictive biomarkers into clinical programs in a data-driven
manner, wherein these biomarkers are emphasized in exact proportion
to the evidence supporting their clinical predictive value. Next-generation
personalized treatment strategies, which emphasize tumor heterogeneity,
evolutionary dynamics and possible future tumor states, will also
be presented.
6:00-9:00 Dinner Course
Laboratory-Developed Tests
(Separate registration required. See Page 4 for additional information.)
Wednesday, May 8
7:30-8:15 am Breakfast Presentation or Morning Coffee
(Sponsorship opportunity available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
Advancing Personalized Medicine
8:25-8:30 Chairperson’s Opening Remarks
8:30-8:55 Personalized Health Care: ‘One Size Does Not Fit All’ Applies
to Patients and Products
M.J. Finley Austin, Ph.D., Personalized Healthcare & Biomarker Strategy
Director, AstraZeneca
The essence of Personalized Health Care (PHC) is identifying,
understanding and partioning drug response variation to improve clinical
outcomes. Existing PHC examples demonstrate diversity in source of
variation, path to market as well as market delivery and uptake. Current
examples will be used to elucidate the implications of differing sources
and degrees of variation, clinical utility, and timing of discovery all have for
clinical trial design, regulatory strategy and market delivery.
8:55-9:20 Molecular Subtyping of Patients for Drug Development
Eric Lai, Ph.D., Senior Vice President and Head, Pharmacogenomics, Takeda
Pharmaceuticals International
While the concept of drug-diagnostic co-development (CDx) has been
around for awhile, most companion diagnostics are still an afterthought
and not an integrated component of drug development. To benefit from
the full potential of CDx, we have to change the strategy of drug target
identification from the single target approach to systematic understanding
of a patient’s disease phenotypes. I will discuss some of the potentials
steps that we have made to the drug development process.
BiomarkerWorldCongress.com Biomarkers & Diagnostics World Congress | 15
Track 5: Biomarkers for Patient Selection
9:20-9:45 Co-Diagnostics in Autoimmune Disorders: Improving
Outcomes in RA and IBD
Mark E. Curran, Ph.D., Vice President, Immunology Biomarkers, Janssen
Research & Development
Rheumatoid arthritis and inflammatory bowel disease are severe immune
diseases with significantly reduced quality of life for patients. Despite
advances in treatment with the evolution of antibody and recombinant
protein based therapeutics, there remains a significant unmet clinical need
for new therapies and integrated treatment solutions. At Janssen we are
focused on transforming therapy in these diseases by applying systems
pharmacology, precision medicine principles and developing companion
diagnostics to create new treatment paradigms. Our objective is to
provide for higher response rates, deeper remission, early interception and
eventually prevention of these diseases. Progress toward these objectives
will be discussed.
9:45-10:15 Complex microRNA Signatures of Response
Sponsored by
and Resistance as Powerful Biomarkers
E. Robert Wassman, M.D., CMO, Rosetta Genomics
10:15-10:30 Sponsored Presentation
(Opportunity available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com)
10:30-11:30 Coffee Break in the Exhibit Hall with Poster Viewing
11:30-11:55 Precision Medicine: Triumphs and Tribulations
Claudio Carini, M.D., Global Clinical Immunology and Biomarkers Lead,
Bioenhancement Development Unit, Pfizer
The current model for drug development is failing. Failures often occur
either during the Phase II trials, where either the candidate drug did not
meet the expected pharmacological requirements or the targeted drug
mechanism did not play a role in the patients population studied. Thus,
a new “personalized medicine” strategy is needed to develop predictive
biomarkers to assist in the decision making process during the pre-clinical
phase of drug development and use biomarkers as companion diagnostics
for stratifying patients in hypothesis-driven clinical trials.
11:55-12:20 pm Towards Personalized Medicine in Metabolic Diseases
Mark Broenstrup, Ph.D., Director, Biomarker and Diagnostics, R&D Diabetes
Division, Sanofi
Currently, more than 346 million people worldwide have diabetes. The
identification of the most effective drug(s) for the individual patient
is guided by a few selection criteria and a trial-and-error approach.
Consequently, the introduction of personalized approaches, accounting
for the heterogeneity of the disease, is regarded as a key enabler for
improved health care. An overview on biomarkers for assessing risk,
monitoring disease progression and predicting response to drugs is
provided, with a focus on beta cell imaging and systems biology solutions.
Finally, major public-private partnerships aiming at personalized solutions in
diabetes will be highlighted.
12:20-12:45 Translating Molecular Targets for Cancer Therapeutics
Glen J. Weiss, M.D., Co-Head, Lung Cancer Unit, The Translational Genomics
Research Institute (TGen); Director, Thoracic Oncology, Virginia G. Piper Cancer
Center Clinical Trials at Scottsdale Healthcare; CMO, CRAB-Clinical Trials
Consortium
The presentation will focus on why there is a push to individualize cancer
therapy, past failures and successes, and how to define the tumor context
of vulnerability (COV). The talk will also describe the steps from pre-clinical
to new drug application and show how to optimize the drug development
path with knowledge of biomarker-based COV.
12:45 Close of Conference
SPONSORSHIP, EXHIBIT & LEAD GENERATION OPPORTUNITIES
CHI offers comprehensive sponsorship packages which include presentation
opportunities, exhibit space and branding, as well as the use of the pre and postshow
delegate lists. Customizable sponsorship packages allow you to achieve
your objectives before, during, and long after the event. Signing on early will
allow you to maximize exposure to hard-to-reach decision makers!
Agenda Presentations
Showcase your solutions to a guaranteed, highly-targeted audience. Package
includes a 15- or 30-minute podium presentation within the scientific agenda,
exhibit space, on-site branding and access to cooperative marketing efforts
by CHI.
Breakfast & Luncheon Presentations
Opportunity includes a 30-minute podium presentation. Boxed lunches are
delivered into the main session room, which guarantees audience attendance
and participation. A limited number of presentations are available for sponsorship
and they will sell out quickly. Sign on early to secure your talk!
Invitation-Only VIP Dinner/Hospitality Suite
Sponsors will select their top prospects from the conference pre-registration
list for an evening of networking at the hotel or at a choice local venue. CHI will
extend invitations and deliver prospects. Evening will be customized according to
sponsor’s objectives i.e.:
• Purely social
• Focus group
• Reception style
• Plated dinner with specific
conversation focus
Exhibit
Exhibitors will enjoy facilitated networking opportunities with high-level
conference delegates. Speak face-to-face with prospective clients and showcase
your latest product, service, or solution.
*Inquire about additional branding opportunities!
Looking for additional ways to drive leads to your sales team?
Cambridge Healthtech Institute can help!
We offer clients numerous options for custom lead generation programs to
address their marketing and sales needs, including:
• Live Webinars
• White Papers
• Market Surveys
• Podcasts
• And More!
Benefits of working with Cambridge Healthtech Institute for your lead
generation needs:
• Your campaign will receive targeted promotion to Cambridge Healthtech
Institute’s unparalleled database of over 800,000 individuals, all of which are
involved in all sectors of the life sciences – lists can be segmented based on
geography, research area, title and industry.
• All custom lead generation programs are promoted through our experienced
marketing team that will develop and drive targeted campaigns to drive
awareness and leads to your lead generation program.
• For our webinar programs, we offer assistance in procuring speakers for
your web symposia through our extensive roster of industry recognized
speakers across multiple disciplines within life sciences, as well as provide
an experienced moderator and dedicated operations team who will
coordinate all efforts.
• If choosing a white paper program, we can offer editorial experience and
provide an industry recognized author to write your white paper.
To customize your participation at this event, please contact:
Ilana Quigley – Business Development Manager
781-972-5457 | iquigley@healthtech.com
16 | Biomarkers & Diagnostics World Congress BiomarkerWorldCongress.com
Track 6: Cancer Drug Resistance
Tuesday, May 8
12:15-1:45 Conference Registration
Biomarkers to Diagnostics
1:45-1:50 Chairperson’s Opening Remarks
1:50-2:15 Investing in Biomarkers and Turning Them into Diagnostics
Felix Frueh, Entrepreneur-in-Residence, Third Rock Ventures
The translation of biomarkers into useful clinical diagnostics requires the
demonstration of clinical benefit and cost effectiveness. Investing in new
technology is not sufficient without the realization that discovery and
development of a new biomarker needs to include the demonstration that the
biomarker makes a difference in clinical outcomes or decision-making, preferably
tested in the environment in which ultimately a diagnostic will be used.
2:15-2:40 From Biomarker Research to Diagnostic Development—Our
Challenges
Yoshi Oda, Ph.D., President, Biomarkers and Personalized Medicine Core
Function Unit, Eisai
Biomarkers play important roles for drug development as a part of
translational research. Several examples about biomarkers for 1) the
evidence of target engagement, 2) patient stratification, 3) drug efficacy
and 4) disease diagnostics will be discussed.
2:40-3:45 Refreshment Break in the Exhibit Hall with Poster Viewing
Molecular Profiling of Tumor Heterogeneity to
Guide Therapy
3:45-3:50 Chairperson’s Opening Remarks
3:50-4:15 Liquid Biopsies to Monitor Response and Resistance to
Targeted Therapies
Luis Alberto Diaz, M.D., Associate Professor of Oncology, Johns Hopkins
Sidney Kimmel Comprehensive Cancer Center
The simplest hypothesis to account for the development of resistance
to EGFR blockade is that rare cells with KRAS mutations pre-exist at low
levels in tumors with ostensibly wild-type KRAS genes. Although this
hypothesis would seem readily testable, there is no evidence in preclinical
models to support it, nor is there data from patients. To test this
hypothesis, we determined whether mutant KRAS DNA could be detected
in the circulation of 28 patients receiving monotherapy with panitumumab,
a therapeutic anti-EGFR antibody. The results suggest that the emergence
of KRAS mutations is a mediator of acquired resistance to EGFR blockade
and that these mutations can be detected in a non-invasive manner. They
explain why solid tumors develop resistance to targeted therapies in a
highly reproducible fashion.
4:15-4:40 Application of Clinical Tumor Genotyping in Targeted Cancer
Therapy
Darrell R. Borger, Ph.D., Co-Director, Translational Research Laboratory,
Massachusetts General Hospital Cancer Center
Multiplexed tumor genotyping has been offered as a physician-ordered
clinical test at a major U.S. cancer center. Over 3,000 patients have been
evaluated and these new capabilities have fostered a genotype-directed
approach to clinical trial design. By testing a broad spectrum of tumor
types, new molecular signatures have been revealed and mechanisms
of de novo and acquired resistance to targeted therapies have been
uncovered. This has provided the foundation for expanding clinical cancer
genotyping approaches for personalizing cancer care.
4:40-5:05 Quantitative Tumor Protein Profiling for Therapy-Relevant
Stratification of Breast Cancer Patients
Hallgeir Rui, M.D., Ph.D., Professor, Cancer Biology, Medical Oncology and
Pathology; Scientific Director, Jefferson Breast Care Center; Program Leader,
Biology of Breast Cancer, Kimmel Cancer Center; Co-Director, Pathology
Translational Research Core, Thomas Jefferson University
Breast cancer is a heterogeneous group of malignancies driven by diverse
oncogenic pathways. Ongoing consortium efforts are to map breast cancer
subtypes at high resolution based on quantitative immunofluorescence
(QIF) profiling of druggable target proteins within carcinoma cells of a
panel of 5,000 untreated primary breast cancer specimens. Progress
with prolactin-receptor-Jak-Stat pathway profiling will be highlighted using
complementary QIF technologies. Utility of resulting protein-based breast
cancer subclassification maps for rational recruitment of patients into
biomarker-driven, adaptive clinical trials will be discussed.
5:05-5:30 Clinical Validation of Predictive Biomarkers and Next-
Generation Personalized Medicine Treatment Strategies Incorporating
Genetic Dynamics
Robert A. Beckman, M.D., External Faculty, Center for Evolution and Cancer,
Helen Diller Family Cancer Center, University of California at San Francisco;
Executive Director, Clinical Development Oncology, Daiichi Sankyo Pharma
Development
The future of oncology drug development lies in personalized therapy
using predictive biomarkers. However, examples of the failure of predictive
biomarkers also exist. In these cases the use of biomarkers increased
the costs, complexity and duration of clinical trials, and narrowed the
treated population unnecessarily. We present methods to adaptively
integrate predictive biomarkers into clinical programs in a data-driven
manner, wherein these biomarkers are emphasized in exact proportion
to the evidence supporting their clinical predictive value. Next-generation
personalized treatment strategies, which emphasize tumor heterogeneity,
evolutionary dynamics and possible future tumor states, will also
be presented.
6:00-9:00 Dinner Course
Laboratory-Developed Tests
(Separate registration required. See Page 4 for additional information.)
Wednesday, May 8
7:30-8:05 am Morning Coffee
Secondary Resistance to Targeted Cancer Therapy
8:05-8:30 Biomarkers and Trastuzumab Resistance
Wen Jin Wu, M.D., Ph.D., Principal Investigator, Division of Monoclonal Antibodies,
Office of Biotechnology Products, Center for Drug Evaluation and Research, FDA
Trastuzumab is an anti-HER2 antibody indicated for the treatment of
HER2-positive breast cancer. Approximately two-thirds of HER2-positive
breast cancers show primary resistance to trastuzumab treatment, and
a majority of patients who achieve an initial response to trastuzumab
acquire resistance to trastuzumab within one year. However, there are
no clinically useful predictive biomarkers that can be used in conjunction
with HER2 expression to predict the outcome of trastuzumab treatment
in the HER2-positive breast cancer patients. We recently found that the
phosphorylation of HER2-Y1248 was associated with the sensitivity of
trastuzumab treatment, suggesting that the phosphorylation status of
HER2-Y1248 may be a predictive biomarker for trastuzumab treatment.
8:30-8:55 Resistance to MAPK Pathway Inhibitors in Melanoma:
Insights and Future Challenges
Jessie Villanueva, Ph.D., Assistant Professor, Molecular and Cellular
Oncogenesis Program, The Wistar Institute
The mitogen-activated protein kinase (MAPK) pathway is a key
therapeutic target for melanoma due to its activation in the majority of
tumors. Numerous small molecule inhibitors aimed at controlling MAPK
activity, such as BRAF and MEK inhibitors, are currently undergoing
clinical investigation. However, their therapeutic success is limited by
the development of drug resistance. To develop effective therapies for
melanoma patients, it is critical to uncover the mechanisms of resistance
to BRAF and MEK inhibitors. This presentation will discuss recent studies
on the molecular mechanisms of resistance to inhibitors of the MAPK
pathway and potential strategies to treat drug-resistant melanomas.
BiomarkerWorldCongress.com Biomarkers & Diagnostics World Congress | 17
8:55-9:20 A Pre-Clinical Model of BRAF Inhibitor Resistance in
Melanoma Reveals a Novel Approach to Forestall Drug Resistance
Meghna Das Thakur, Ph.D., Presidential Postdoctoral Fellow, Novartis Institutes
for BioMedical Research
BRAF inhibitors such as vemurafenib have shown promising effects in
patients with mutant BRAF(V600E) melanomas, but the tumors generally
develop resistance. Interestingly, the vemurafenib-resistant melanomas
become drug dependent for their continued proliferation, such that
cessation of drug administration leads to regression of established drugresistant
tumors. Thus, a discontinuous dosing strategy exploiting the
fitness disadvantage shown by drug-resistant cells in the absence of the
drug forestalls the onset of lethal drug-resistant disease.
9:20-9:45 Non Cell-Autonomous Mechanisms of Resistance against
Anti-EGFR Therapy
Janghee Woo, M.D., Albert Einstein Medical Center; Recipient of AACRGlaxoSmithKline
Clinical Cancer Research Scholar Award and Dana-Farber/
Harvard Cancer Center Award
Our findings suggest that stroma-derived MMP9 may help tumors bypass
common mutational mechanisms for constitutive growth factor pathway
activation and confer resistance to anti-EGFR therapy through activation of
the ERBB2/ERK/JUN pathway. Stromal MMP9 expression may therefore
have value as a predictive marker for cetuximab response and in stratifying
patients before treatment.
9:45-10:30 Sponsored Presentations
(Opportunities available. Contact Ilana Quigley at 781-972-5457 or
iquigley@healthtech.com)
10:30-11:30 Coffee Break in the Exhibit Hall with Poster Viewing
11:30-11:55 Managing Secondary Drug Resistance in the Clinic: The
Memorial Sloan-Kettering Approach
Maria E. Arcila, M.D., Department of Pathology, Memorial Sloan-Kettering
Cancer Center
Resistance to Various Therapies:
Cancer Does Not Discriminate
11:55-12:00 Chairperson’s Remarks
12:00-12:25 A20 Ubiquitin E3 Ligase is a Biomarker of the Cancer
Stem Cell Resistance to Apoptotic Drugs
Chunhai “Charlie” Hao, M.D., Ph.D., Associate Professor, Neuropathology
Attending, Department of Pathology and Laboratory Medicine, Emory
University School of Medicine
The TRAIL (tumor necrosis factor-related apoptosis-inducing ligand)
apoptosis pathway has emerged as a cancer therapeutic target; however,
Phase II trials recently completed have showed limited if any antitumor
activities of TRAIL pathway-targeted therapies. Molecular and functional
examination of patients’ glioblastoma tissues and derived cancer stem
cells reveals the resistance mechanism by which the ubiquitin E3 ligase
A20 mediated poly-ubiquitination inhibits the cleavage of apoptosisinitiating
caspase-8 and the initiation of TRAIL-induced apoptosis. The
study suggests that the full characterization of patients’ cancer tissues
and derived cancer stem cells can predict the cancer responsiveness
to treatment and thus should be a critical pre-clinical trial step in
drug development.
12:25-12:50 pm Molecular Determinants of Hormone-Refractory
Prostate Cancer
Atish Choudhury, M.D., Instructor in Medicine, Medical Oncology, Dana-Farber
Cancer Institute
To identify novel genes that can confer androgen independence to
prostate cancer cells in vivo, we performed an unbiased screen for kinases
conferring androgen-independent tumor formation to androgen-dependent
transformed prostate epithelial cells in vivo. These kinases are likely
to activate signaling pathways that are relevant for conferring castrate
resistance in patients with advanced prostate cancer, and inhibiting these
genes is likely to result in inhibition of cancer cell proliferation and/or
restoration of hormone sensitivity. Integration of our ambitious functional
studies with gene expression and sequencing data in CRPC from tumor
samples being generated through collaborations between DFCI and the
Broad Institute will provide us a more comprehensive understanding of the
development of castrate resistance and novel targets for therapy.
12:50-1:15 Impact of microRNAs in Chemoresistance
Jingfang Ju, Ph.D., Co-Director, Translational Research, Pathology, Stony Brook
University
Non-coding miRNAs contribute to both intrinsic and extrinsic
chemoresistance mechanism, particularly in colon cancer stem cells.
We first discovered several miRNAs suppressing the expression of both
thymidylate synthase and dihydrofolate reductase to impact 5-FU and MTX
sensitivity. The expression of miR-215 was significantly associated with
colorectal cancer patient survival. Our recent studies also show miRNAs
impact intrinsic apoptotic pathways and autophagy. We believe miRNA
based therapeutics, diagnosis and prognosis may emerge in the near
future to benefit patients.
1:15 Close of Conference
Track 6: Cancer Drug Resistance
Lead Media Partners
Media Partners Web Partner
Lead Sponsoring Publications
Sponsoring Publications
18 | Biomarkers & Diagnostics World Congress BiomarkerWorldCongress.com
Track 7: Exosomes and Microvesicles as Biomarkers and Diagnostics
Tuesday, May 8
12:15-1:45 Conference Registration
Exosome Biomarkers in Drug Development
1:45-1:50 Chairperson’s Opening Remarks
1:50-2:15 Exosomes as Biomarkers for Translational Medicine
Holly Hilton, Ph.D., Head, Disease and Translational Genomics, Hoffmann-La
Roche; Adjunct Professor, Graduate School of Biomedical Sciences, University
of Medicine and Dentistry New Jersey
The need for new, relevant biomarkers for translational drug discovery
research is critical. Exosomes are small microvesicles secreted by a wide
range of mammalian cell types under normal and pathological conditions.
The unique signature of exosomal membrane and cytoplasmic proteins as
well as mRNAs and miRNAs can reveal the cell of origin and the condition
of those cells. Isolation and profiling of exosomes from accessible patient
biofluids, such as urine, blood, BALF and CSF, make them ideal candidates
as biomarkers. Examples of their utility as disease biomarkers of chronic
kidney disease and Alzheimer’s as well as possible applications of patient
stratification will be discussed. The current state of challenges to the
widespread use of fluid-based biomarkers will be explored.
2:15-2:40 Investigation of Microparticles as Potential Translatable
Biomarkers of Vascular Injury
Sharon Sokolowski, Ph.D., Principal Scientist, Pfizer Global Research &
Development
Endothelial cells (EC) are thin, flattened cells that line blood and lymph vessel
walls. Endothelial microparticles (EMPs) are small vesicles (0.1-1 mm) that are
released into circulating blood from activated, injured or apoptotic endothelial
cells and are found at elevated levels in a number of diseases associated with
vascular/endothelial dysfunction. The EMPs are being investigated as potential
translatable biomarkers of drug-induced vascular injury.
2:40-3:45 Refreshment Break in the Exhibit Hall with Poster Viewing
3:45-4:10 Utilization of Next-Gen Genomics Technologies for
Unraveling Exosomal Biomarker Potential
Saumya Pant, Ph.D., Research Fellow, Merck
4:10-4:35 CNS Exosomes and the Art of Eavesdropping
Reyna Favis, Ph.D., Scientific Director, Janssen Pharmaceutical Companies of
Johnson & Johnson
Gaining insight into both genomic changes and differences in the central
nervous system of living humans is currently pursued via investigation of
post mortem brain tissue and lymphocytes from living donors. Analyses
of both tissue types suffer from numerous caveats. There is an urgent
need to develop non-invasive methods that can accurately report temporal
changes, as well as inter-individual differences, in the CNS that may
elucidate neurological and neuropsychiatric disease and drug response.
4:35-5:00 Technology Assessment for Evaluation of Exosomal
microRNA as Novel Biomarkers
Shidong Jia, Ph.D., Scientist, Oncology Biomarker Development, Genentech
Dr. Jia’s lab has developed working procedures to evaluate exosomal
microRNA as novel biomarkers for cancer prognosis, prediction and patient
stratification. In particular, their work has refreshed current practice and
demonstrated a new approach for studying microRNA signature in patient
blood samples.
5:00-5:25 The Exosome Factor in Cancer
Lorraine O’Driscoll, Ph.D., Associate Professor, Pharmacology; Director, Research,
School of Pharmacy and Pharmaceutical Sciences, Trinity College Dublin
Our research at Trinity College Dublin supports exosomes cargo having
relevance as diagnostic, prognostic and predictive biomarkers. Evidence
indicates they are also causative in cancer spread and drug resistance.
Here we will discuss examples of this research in relation to breast cancer
and prostate cancer.
6:00-9:00 Dinner Course
Laboratory-Developed Tests
(Separate registration required. See Page 4 for additional information.)
Wednesday, May 8
7:30-8:15 am Breakfast Presentation or Morning Coffee
(Sponsorship opportunity available. Contact Ilana Quigley at 781-972-5457
or iquigley@healthtech.com).
Exosomes as Disease Markers
8:25-8:30 Chairperson’s Opening Remarks
8:30-8:55 Salivary Exosomes and Biomarkers Development
David T.W. Wong, D.M.D., D.M.Sc., Professor, Associate Dean, Research, UCLA
School of Dentistry and Director of Dental Research Institute
Extracellular RNA is an emerging concept in cellular communication
and biomarker development. Salivary extracellular RNA, microRNA and
snoRNA have recently been shown to be contained within exosomes
and can be developed to be discriminatory biomarkers for oral as well as
systemic diseases.
8:55-9:20 Circulating Exosomes in Liver Disease
Gyongyi Szabo, M.D., Ph.D., Professor, Gastroenterology, University of
Massachusetts Medical School
microRNAs (miRNAs) are fine tuners of diverse biological responses and
are expressed in various cell types of the liver. They can also serve as
biomarkers of liver damage and inflammation. We studied miRNA-122 that
is abundant in hepatocytes and miR-155, -146a and -125b that regulate
inflammation in immune cells in mouse models of various types of liver
diseases and found that serum/plasma miR-122 correlated with ALT
increases in the liver damage. miR-155, a regulator of inflammation, was
increased in serum/plasma liver injury associated with inflammation.
Depending on the type of liver injury, circulating miRNAs showed
association either with the exosome-rich or protein-rich compartments.
Our results suggest that circulating miRNAs may serve as biomarkers
to differentiate between hepatocyte injury and inflammation and the
exosome versus protein association of miRNAs may provide further
specificity to mechanisms of liver pathology.
9:20-9:45 Microvesicles: Linking the Bone Marrow and Endothelium in
Pulmonary Vascular Disease
Jason M. Aliotta, M.D., Assistant Professor, Medicine, Warren Alpert Medical
School, Brown University
Extracellular vesicles (EVs) represent potentially important mediators of
cell-to-cell communication and, depending on their source, facilitate tissue
repair or remodeling. We’ve demonstrated that EVs isolated from mice
with monocrotaline-induced pulmonary hypertension (PH) induce features
of PH in normal mice. This may be due to EV-induced apoptosis resistance
of pulmonary vascular endothelial cells or EV-induced differentiation
of marrow cells into progenitor cells which, in turn, induce vascular
remodeling. Conversely, we have found that mesenchymal stem cellderived
EVs may reverse monocrotaline-induced PH.
9:45-10:30 Sponsored Presentations
(Opportunities available. Contact Ilana Quigley at 781-972-5457 or
iquigley@healthtech.com)
10:30-11:30 Coffee Break in the Exhibit Hall with Poster Viewing
BiomarkerWorldCongress.com Biomarkers & Diagnostics World Congress | 19
Track 7: Exosomes and Microvesicles as Biomarkers and Diagnostics
Exosomes as Novel Cancer Biomarkers
11:30-11:35 Chairperson’s Remarks
11:35-12:00 pm The Exosome Platform as a Real-Time Tumor Status Monitor
Douglas D. Taylor, Ph.D., Professor, Obstetrics and Gynecology, University of
Louisville School of Medicine
12:00-12:25 Customized Heterogeneity of Breast Cancer Microvesicles
Dominik Duelli, Ph.D., Assistant Professor, Cellular and Molecular
Pharmacology, Rosalind Franklin University of Medicine & Science, Chicago
Medical School
Breast cancer cells, unlike normal cells, release a heterogeneous
population of circulating microvesicles. Resolving this heterogeneity
suggests that individual microvesicle subclasses have different subcellular
origin, different contents, and different destinations. Each subclass
contains mutually exclusive, functional marker microRNA species, and
some proteins with different functions in docking and lysis resistance in
blood plasma. Additionally, organ-site of metastasis influences the ratio of
these proteins, suggesting that these differences could be used to detect
the presence of malignant cells in the body.
12:25-12:50 Tumor-Derived Microvesicles: Biology and Clinical Potential
Crislyn D’Souza-Schorey, Ph.D., Professor, Biological Sciences, University of
Notre Dame
Tumor-derived microvesicles (TMVs) are heterogeneous membrane-bound
sacs that are shed from tumor cells into the extracellular environment.
The formation of these shed vesicles likely involves the vertical trafficking
of intracellular cargo to the cell surface. The complexity of bioactive
cargo contained in TMVs suggests multi-pronged mechanisms by which
shed TMVs can condition the extracellular milieu to facilitate disease
progression. It also demonstrates the potential to translate this knowledge
into innovative approaches for cancer diagnostics and therapy.
12:50-1:15 Exosome Biomarkers of Brain Tumors
Fred H. Hochberg, M.D., Associate Professor, Neurology, Massachusetts
General Hospital
We explore technology for detection of plasma and CSF exosomal
mutations specific to brain tumors. The analytics for mutations EGFrvIII
and IDH1.132 offer the potential to provide a diagnostic biomarker for
low grade and high grade gliomas. An eighteen member consortium,
collaborating with the ABC2 Foundation and the company Exosome
Diagnostics, will validate the sensitivity of these biomarker assays. The
presentation will include discussion of pre-clinical detection, SOPs for
specimen handling and the rationale for use of these biomarkers.
1:15 Close of Conference
Conference Hotel :
Loews Philadelphia Hotel
1200 Market Street
Philadelphia, PA 19107
Phone: 215-627-1200
HOTEL & TRAVEL
INFORMATION
Discounted Room Rate: $229 s/d
Discounted Room Rate Cut-off Date:
April 8, 2013
Please visit our conference website
to make your reservation online or
call the hotel directly to reserve your
sleeping accommodations. You will need
to identify yourself as a Cambridge
Healthtech Institute conference attendee
to receive the discounted room rate with
the host hotel. Reservations made after
the cut-off date or after the group room
block has been filled (whichever comes
first) will be accepted on a space and
rate-availability basis. Rooms are limited,
so please book early.
Flight Discounts:
Special discount rentals have been established with American Airlines
for this conference.
• Call American Airlines 1-800-433-1790 and use Conference code 8353BL.
• Go to http://www.aa.com/group and enter Conference code 8353BL in promotion
discount box.
• Contact our dedicated travel agents at 1-877-559-5549 or chi@protravelinc.com.
Car Rental Discounts:
Special discount rentals have been established with Hertz for this conference.
• Call Hertz 1-800-654-3131 and use our Hertz Convention Number (CV): 04KL0003
• Go to http://www.hertz.com and use our Hertz Convention Number (CV): 04KL0003
Top Reasons to Stay at The Loews Philadelphia
• Minutes from Amtrak 30th Street Station and 20 minutes from
Philadelphia Airport
• Complimentary wireless internet in your guest room
• Close to many of Philadelphia’s historical sites, including the Liberty
Bell and Independence Hall
• Steps from Reading Terminal Market, which offers an exhilarating selection
of baked goods, meats, poultry, seafood, produce, flowers and more
• Pet-friendly accommodations including specialty pet menus, gifts
upon arrival and dog-walking services
• Located in the historic PSFS Building: A 20th Century Masterpiece
BIOMARKERS & DIAGNOSTICS
world congress 2013
Additional registrat ion deta ils
Each registration includes all conference
sessions, posters and exhibits, food
functions, and access to the conference
proceedings link.
Handicapped Equal Access: In accordance
with the ADA, Cambridge Healthtech
Institute is pleased to arrange special
accommodations for attendees with
special needs. All requests for such
assistance must be submitted in writing
to CHI at least 30 days prior to the start
of the meeting.
To view our Substitutions/
Cancellations Policy, go to
http://www.healthtech.com/regdetails
Video and or audio recording of any kind
is prohibited onsite at all CHI events.
Receive a FREE eNewsletter by signing up
at chimediagroup.com
The latest industry news, commentary
and highlights from Bio-IT World
Innovative management in clinical trials
A series of diverse reports designed to
keep life science professionals informed
of the salient trends in pharmaceutical
technology, business, clinical development,
and therapeutic disease markets.
For a detailed list of reports, visit
InsightPharmaReports.com, or contact
Rose LaRaia, rlaraia@healthtech.com,
+1-781-972-5444.
Barnett is a recognized leader in clinical
education, training, and reference guides
for life science professionals involved in
the drug development process. For more
information, visit barnettinternational.com.
Cambridge Healthtech Associates™
(CHA™) uses its collaborative model to
improve the speed and economics of life
sciences R&D, leveraging its consulting,
technology evaluations and communities.
Visit http://www.chacorporate.com.
DINNER courses
Academic, Government,
Commercial Hospital-affiliated
Single Dinner Course Pricing $595 $295
Double Dinner Course Pricing $895 $495
May 6, 2013 May 7, 2013
Fit-for-Purpose Biomarker Assay Development and Validation Laboratory-Developed Tests
Next-Generation Sequencing as a Clinical Test: It Takes a Community
Conference Pricing
ALL ACCESS Executive Pricing: Includes access to entire 3-days of Congress programs, including Executive Summit. (Does not include
access to dinner courses.)
Advance Registration Discount until March 29, 2013 $2445 $1145
Registrations after March 29, 2013, and on-site $2695 $1195
BEST VALUE Main Conference Pricing: Includes access to entire 3-days of Congress programs. (Does not include access to Executive
Summit or dinner courses.)
Advance Registration Discount until March 29, 2013 $2195 $1095
Registrations after March 29, 2013, and on-site $2395 $1145
Single Conference Pricing: Includes access to 1 program. (Does not include access to Executive Summit or dinner courses.)
Advance Registration Discount until March 29, 2013 $1495 $695
Registrations after March 29, 2013, and on-site $1745 $775
May 6-7, 2013 May 7-8, 2013
Track 1: Translational Biomarkers in Drug Development Track 5: Biomarkers for Patient Selection
Track 2: Clinical Assay Development Track 6: Cancer Drug Resistance
Track 3: Cancer Tissue Diagnostics Track 7: Exosomes and Microvesicles as Biomarkers and Diagnostics
Track 4: Executive Summit: Companion Diagnostics (May 6-8, 2013)
Conference Discounts
Poster Submission-Discount ($50 Off)
Poster abstracts are due by March 29, 2013. Once your registration has been fully processed, we will send an email containing a unique link allowing you to submit
your poster abstract. If you do not receive your link within 5 business days, please contact jring@healthtech.com. *CHI reserves the right to publish your poster title
and abstract in various marketing materials and products.
REGISTER 3 –
4th IS FREE: Individuals must register for the same conference or conference combination and submit completed registration form together for discount to apply.
Additional discounts are available for multiple attendees from the same organization. For more information on group rates contact David Cunningham at +1-781-972-5472
How to Register: BiomarkerWorldCongress.com
reg@healthtech.com • P: 781.972.5400 or Toll-free in the U.S. 888.999.6288
Please use keycode
BMC F
when registering!
If you are unable to attend but would like to purchase the Biomarker and Diagnostic World Congress CD for $750 (plus shipping), please visit
BiomarkerWorldCongress.com. Massachusetts delivery will include sales tax.
Please refer to the Registration Code below: Cambridge Healthtech Institute
250 First Avenue, Suite 300
Needham, MA 02494
http://www.healthtech.com
Fax: 781-972-5425
Pricing and Registration Information
Read Full Post »




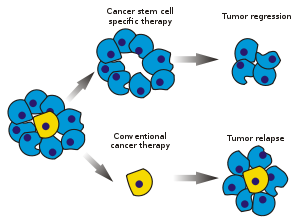

 Physiologically, the proteins play a role in calcium and Pi homeostasis as demonstrated by studies on mouse transgenic models. In addition to cancer, the protein has been linked to atherosclerosis, hypoxia response and in wound repair (Lal et al., 2001; Iyer et al., 1999). Pharmacologically, an STC1 receptor has been deduced from studies and thought to be localized to the mitochondria where it has been shown to have a relationship with the mitochondrial electron transfer (McCudden et al., 2002). Recent studies show that STC1 activates the mitochondrial antioxidant pathway through its regulation of intracellular calcium (Sheikh-Hamad, 2010). Overall, STC1 and STC2 are thought to be secreted as phosphoproteins as demonstrated by coimmunoprecipitations of cellular lysates. And, it’s thought the proteins play a role in mineralizing tissues (e.g. bone) to control the levels of calcium and Pi via their influence on calcium channels and sodium/Pi co-transporters. A schematic diagram showing how stannniocalcin might be have pro-apoptic functions is shown in Figure 1 (Yeung et al., 2012).
Physiologically, the proteins play a role in calcium and Pi homeostasis as demonstrated by studies on mouse transgenic models. In addition to cancer, the protein has been linked to atherosclerosis, hypoxia response and in wound repair (Lal et al., 2001; Iyer et al., 1999). Pharmacologically, an STC1 receptor has been deduced from studies and thought to be localized to the mitochondria where it has been shown to have a relationship with the mitochondrial electron transfer (McCudden et al., 2002). Recent studies show that STC1 activates the mitochondrial antioxidant pathway through its regulation of intracellular calcium (Sheikh-Hamad, 2010). Overall, STC1 and STC2 are thought to be secreted as phosphoproteins as demonstrated by coimmunoprecipitations of cellular lysates. And, it’s thought the proteins play a role in mineralizing tissues (e.g. bone) to control the levels of calcium and Pi via their influence on calcium channels and sodium/Pi co-transporters. A schematic diagram showing how stannniocalcin might be have pro-apoptic functions is shown in Figure 1 (Yeung et al., 2012). However, stanniocalcin’s more prominent role is arguably as a cancer biomarker. Its expression has been shown to be affected in a number of different cancer pathologies. Table 1 shows a representative list of cancers where stanniocalcin levels are differentially expressed depending on the cancer. Thus, it appears that stanniocalcin is a good candidate cancer biomarker. It is hypothesized that this is due in part to its role in tumor vasculature (Chang et al., 2003). It should be noted that the list is but a brief compilation while stanniocalcin has been linked to other cancers as well.
However, stanniocalcin’s more prominent role is arguably as a cancer biomarker. Its expression has been shown to be affected in a number of different cancer pathologies. Table 1 shows a representative list of cancers where stanniocalcin levels are differentially expressed depending on the cancer. Thus, it appears that stanniocalcin is a good candidate cancer biomarker. It is hypothesized that this is due in part to its role in tumor vasculature (Chang et al., 2003). It should be noted that the list is but a brief compilation while stanniocalcin has been linked to other cancers as well.