Stem Cells and Cancer
Larry H. Bernstein, MD, FCAP, Curator
Leaders in Pharmaceutical Intelligence
Series E. 2; 8.09
Cancer cells programmed back to normal by US scientists
By Sarah Knapton, Science Editor
Scientists have turned cancerous cells back to normal by switching back on the process which stops normal cells from replicating too quickly. Cancer cells could be stopped from replicating after scientists found how to switch on the brakes.
http://www.telegraph.co.uk/news/science/science-news/11821334/Cancer-cells-programmed-back-to-normal-by-US-scientists.html
Cancer cells have been programmed back to normal by scientists in a breakthrough which could lead to new treatments and even reverse tumour growth.
For the first time aggressive breast, lung and bladder cancer cells have been turned back into harmless benign cells by restoring the function which prevents them from multiplying excessively and forming dangerous growths.
Scientists at the Mayo Clinic in Florida, US, said it was like applying the brakes to a speeding car.
So far it has only been tested on human cells in the lab, but the researchers are hopeful that the technique could one day be used to target tumours so that cancer could be ‘switched off’ without the need for harsh chemotherapy or surgery.
“We should be able to re-establish the brakes and restore normal cell function,” said Profesor Panos Anastasiadis, of the Department for Cancer Biology.
“Initial experiments in some aggressive types of cancer are indeed very promising.
“It represents an unexpected new biology that provides the code, the software for turning off cancer.”
Cells need to divide constantly to replace themselves. But in cancer the cells do not stop dividing leading to huge cell reproduction and tumour growth.
The scientists discovered that the glue which holds cells together is regulated by biological microprocessors called microRNAs. When everything is working normally the microRNAs instruct the cells to stop dividing when they have replicated sufficiently. They do this by triggering production of a protein called PLEKHA7 which breaks the cell bonds. But in cancer that process does not work.
Scientists discovered they could switch on cancer in cells by removing the microRNAs from cells and preventing them from producing the protein.
And, crucially they found that they could reverse the process switching the brakes back on and stopping cancer. MicroRNAs are small molecules which can be delivered directly to cells or tumours so an injection to increase levels could switch off disease.
“We have now done this in very aggressive human cell lines from breast and bladder cancer,” added Dr Anastasiadis.
“These cells are already missing PLEKHA7. Restoring either PLEKHA7 levels, or the levels of microRNAs in these cells turns them back to a benign state. We are now working on better delivery options.”
Cancer experts in Britain said the research solved a riddle that biologists had puzzled over for decades, why cells did not naturally prevent the proliferation of cancer.
“This is an unexpected finding,” said Dr Chris Bakal, a specialist in how cells change shape to become cancerous, at the Institute for Cancer Research in London.
“We have been trying to work out how normal cells might be suppressing cancer, and stopping dividing when they form contacts with each other, which has been a big mystery.
“Normal cells touch each other and form junctions then they shut down proliferation. If there is a way to turn that back on then that would be a way to stop tumours from growing.
“I think in reality it is unlikely that you could reverse tumours by reversing just one mechanism, but it’s a very interesting finding.”
Henry Scowcroft, Cancer Research UK’s senior science information manager, said: “This important study solves a long-standing biological mystery, but we mustn’t get ahead of ourselves.
“There’s a long way to go before we know whether these findings, in cells grown in a laboratory, will help treat people with cancer. But it’s a significant step forward in understanding how certain cells in our body know when to grow, and when to stop. Understanding these key concepts is crucial to help continue the encouraging progress against cancer we’ve seen in recent years.”
The research was published in the journal Nature Cell Biology.
Biomaterial Sponge-Like Impant Traps Spreading Cancer Cells
September 9, 2015 by mburatov http://wp.me/ptV19-1vG
Prof Lonnie Shea, from the Department of Biomedical Engineering at the University of Michigan and his team have designed a small sponge-like implant that has the ability to mop up cancer cells as they move through the body. This device has been tested in mice, but there is hope that the device could act as an early warning system in patients, alerting doctors to cancer spread. The sponge-like implant also seemed to stop rogue cancer cells from reaching other areas where they could establish the growth of new tumors. Shea and others published their findings in the journal Nature Communications.
According to Cancer Research UK, nine in 10 cancer deaths are caused by the disease-spreading to other areas of the body. Stopping the spread of cancer cells, or metastasis, is one of the ways to prevent cancers from becoming worse. Complicating this venture is the fact that cancer cells that circulate in the bloodstream are rare and difficult to detect.
Shea’s device is about 5mm or 0.2 inches in diameter and made of a “biomaterial” already approved for use in medical devices. So far, this implant has so far been tested in mice with breast cancer. Implantation experiments showed that it can be placed either in the abdominal fat or under the skin and that it tended to suck up cancer cells that had started to circulate in the body.
The implant mimicked a process known as chemoattraction in which cells that have broken free from a tumor are attracted to other areas in the body by immune cells. Shea and others found that these immune cells are drawn to the implant where they “set up shop.” This is actually a natural reaction to any foreign body, and the presence of the immune cells also attracts the cancer cells to the implant.
Initially, Shea and others labeled cancer cells with fluorescent proteins that caused them to glow under certain lights, which allowed them to be easily spotted. However, they eventually went on to use a special imaging technique that can distinguish between cancerous and normal cells. They discovered that they could definitively detect cancer cells that had been caught within the implant.
Unexpectedly, when they measured cancer cells that had spread in mice with and without the implant, they showed that the implant not only captured circulating cancer cells, but it also reduced the numbers of cancer cells present at other sites in the body.
Shea, said that he and his team were planning the first clinical trials in humans fairly soon: “We need to see if metastatic cells will show up in the implant in humans like they did in the mice, and if it’s a safe procedure and that we can use the same imaging to detect cancer cells.”
Shea and his coworkers are continuing their work in animals to determine what the outcomes if the spread of the cancer spread was detected at a very early stage, which is something that is presently not yet fully understood.
Lucy Holmes, Cancer Research UK’s science information manager, said: “We urgently need new ways to stop cancer in its tracks. So far this implant approach has only been tested in mice, but it’s encouraging to see these results, which could one day play a role in stopping cancer spread in patients.”
U of Penn Group Releases Hopeful Results of CAR T-Cells Trial
Sept 8, 2015 by mburatov
https://beyondthedish.wordpress.com/2015/09/08/u-of-penn-group-releases-hopeful-results-of-car-t-cells-trial/
Chimeric Antigen Receptor T-Cells (CART-cells) are a type of genetically engineered type of immune cell that represents one of the most promising avenues of cancer therapy. Such treatments can induce sustained remissions in patients with stubborn disease.
Studies with CART-cells have been tested in patients with relapsed and stubborn chronic lymphocytic leukemia (CLL). Now a new publication by Porter and others reports the results of a clinical trial that examined CART-cells as a treatment for blood-based cancers. This study reports that infused CART-cells were functional up to 4 years after treatment. Patients also achieved completely remission, and no patient who achieved complete remission relapsed, and no minimal residual disease was detected, suggesting that in a subset of patients, CAR T cells may drive disease eradication.
Patients enrolled in this study suffered from CLL and had a poor prognosis. The CART-cells employed in this study targeted the molecule CD19. Porter and others report the mature results of the treatment of 14 patients with relapsed and refractory CLL.
The patient’s own T-Cells were extracted from circulating blood, and genetically engineered to express a CD19-directed receptor. Patients received doses of 0.14 × 10[8] to 11 × 10[8] CTL019 cells. Patients were monitored for toxicity, response, expansion, and persistence of circulating CTL019 T cells.
The overall response rate in these heavily pretreated CLL patients was 8 of 14 (57%), and there were 4 complete remissions (CR) and 4 partial remissions (PR). The expansion of the CAR T-cells in culture correlated with clinical responses; the better the engineered T-cells grew in culture the better they performed in the Patient’s bodies. Furthermore, the CAR T-cells persisted and remained functional beyond 4 years in the first two patients achieving Complete Remission. None of the patients who experienced Complete Remission have relapsed.
All the patients who responded to the treatment developed “B cell aplastic” (abnormally low B-cell levels) and experienced cytokine release syndrome, which was part and partial of T cell proliferation.
Minimal residual disease was not detectable in patients who achieved Complete Remission, suggesting that disease eradication may be possible in some patients with advanced CLL.
New Method to Regulate Stem Cell Differentiation
GEN News Highlights Sep 2, 2015
http://www.genengnews.com/gen-news-highlights/new-method-developed-to-regulate-stem-cell-differentiation/81251707/
|
| Researchers have developed a method that enables the regulation of a single gene’s behavior without changing the genome itself. [Professor Otonkoski Lab, University of Helsinki] |
http://www.genengnews.com/Media/images/GENHighlight/thumb_Sep0915_UnivHelsinki_StemCellDifferentiationGraph3620321462.jpg
Scientists at the University of Helsinki in Finland say they have developed a new method that enables the activation of genes in a cell without changing the genome. Applications of the method include directing the differentiation of stem cells.
The method was developed by researchers Diego Balboa and Jere Weltner, who are working on their doctoral dissertations in the lab of Timo Otonkoski, Ph.D., at the Meilahti medical campus of the University of Helsinki. The research study (“Conditionally Stabilized dCas9 Activator for Controlling Gene Expression in Human Cell Reprogramming and Differentiation”) was published in Stem Cell Reports.
The hottest topics in stem cell research at the moment are methods that can regulate the differentiation of cells. The differentiation process is based on how genes in a cell are activated and deactivated, so researchers are looking for ways to control the activation of the genes. The researchers dream of being able to activate and deactivate genes precisely at specific moments.
“We can produce undifferentiated stem cells from specialized cells, also known as iPS, or induced pluripotent stem cells, and we can regulate the differentiation of these cells by providing them with the right kinds of growth environments. However, we cannot control the differentiation process sufficiently. The process may go smoothly, but then at the very end, a single gene won’t activate at the necessary time, and the cell remains immature,” Dr. Otonkoski explains.
Researchers in Dr. Otonkoski’s laboratory have now developed a method that enables the regulation of a single gene’s behavior without changing the genome itself. The method employs CRISPR technology, but the regulation itself is controlled by the addition of chemicals. The desired gene is made receptive to the drug by introducing bits of RNA into the cell that will bind to the activator protein and the gene’s regulatory area. The gene will then activate in the desired way when the chemicals that regulates the activator protein are provided to the cell.
“In our research, we used two common antibiotics, doxycycline and trimethoprim, and these chemicals enabled us to regulate the expression of many genes precisely and effectively. The method worked on all cells we tested, including stem cells. We used human cells in our development,” continued Dr. Otonkoski, who emphasized that the method is currently being used in experimental models. It is far too early to discuss therapeutic applications.
“The basic idea has now been developed, and the method has been demonstrated to be viable, and I believe that it can become a very important research tool. In my laboratory we use the method to regulate the differentiation of stem cells, but it has many potential applications in other research fields, for example, in cancer biology.”
Single Cell Analysis (SCA): Expanding in Importance in Life Science Research — circa 2015
Technologies Impacting SCA and Driving Translation Towards Single Cell-based Diagnostics
GEN Sep 2, 2015 http://www.genengnews.com/insight-and-intelligence/single-cell-analysis-sca-expanding-in-importance-in-life-science-research-circa-2015/77900516/
The focus of this GEN Market & Tech Analysis report is Single Cell Analysis (SCA) Trends.
- Select Biosciences performed a study of the en bloc Single Cell Analysis (SCA) space in August 2015 to reveal trends in this evolving field—the results from these analyses are presented in this GENReport
- The field is evolving as it is permeating into life sciences research as well as diagnostics development — this represents the translation of SCA and is evidenced for instance by the increasing penetrance of circulating tumor cell (CTC) research in the SCA space
- The field of SCA is intersecting with nucleic acid and protein characterizing approaches/technologies which suggests that the “cargo” of single cells is a current area of study
- The utilization of microfluidics approaches in SCA is a key and growing theme and suggests that the use of microfluidics for single cell capture and interrogation is gaining momentum
Shedding Light On Century-Old Biochemical Mystery
Aug 20, 2015 http://www.technologynetworks.com/Metabolomics/news.aspx?ID=182141
Yale scientists have used magnetic resonance measurements to show how glucose is metabolized in yeast to answer the puzzle of the “Warburg Effect.”
Given plenty of glucose and oxygen, yeast and cancer cells do not burn it all to produce energy but convert much of it to the byproducts ethanol and lactate, respectively.
In the 1920s Nobel laureate Otto Heinrich Warburg asked why these cells were so wasteful of energy. He suggested that this seemingly inefficient cellular use of resources was a root cause of cancer, a hypothesis that has been the subject of research ever since.
Almost a century later, two Yale scientists have used magnetic resonance measurements showing how glucose is metabolized in yeast to answer the puzzle of the “Warburg Effect.” The production of these byproducts is a result of the cell’s need to keep its internal state constant during glucose consumption, they report.
This biochemical response is an example of homeostasis, a fundamental need of all life forms.
“It’s the cell’s way of saying it has enough to eat,” said Robert Shulman, professor emeritus of molecular biophysics and biochemistry.
In the 1980s, Shulman conducted pioneering studies of metabolism in yeast using magnetic resonance spectroscopy, a method then confined to the study of cells but now used routinely in patients.
More recently, Shulman and co-author Douglas Rothman, professor of diagnostic radiology and of biomedical engineering, reviewed the data applying new methods of analyzing metabolic control. They found key intermediate molecular steps involved in the conversion of glucose to ethanol as well as to ATP, the chief energy source of cells. When these molecular switches that maintained homeostasis were disabled by mutations, the cells died from accumulated excesses of both byproducts and ATP.
This chemical balancing act explains why yeast and likely cancer cells do not convert all available fuel to energy that they could use to divide and flourish.
“Cancer cells have to survive first,” Rothman said.
Shulman and Rothman point out that their results open a new direction for cancer researchers — identifying metabolic homeostasis mechanisms and targeting them for treatment.
“By taking another look at the in vivo data available from magnetic resonance experiments, I think we can revitalize research efforts in a host of areas,” Shulman said.
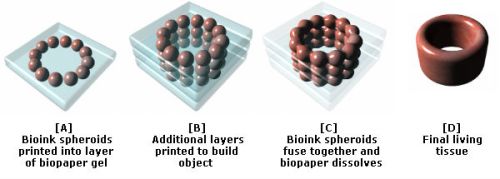
Orchestrating Organoids
A guide to crafting tissues in a dish that reprise in vivo organs
By Kelly Rae Chi | Sep 1, 2015 http://www.the-scientist.com//?articles.view/articleNo/43842/title/Orchestrating-Organoids/
In 2009, at the Hubrecht Institute in Utrecht, Netherlands, Hans Clevers and postdoc Toshiro Sato took adult stem cells from the mouse intestine and created the first mini-guts they called organoids—three-dimensional organized clusters of cells that would allow the researchers to glean new insights into the biology of gut health and disease, including colorectal cancer.
This method inspired many other scientists, working with both mouse and human tissues, to create a rapidly expanding palette of organoids that now includes kidney, brain, liver, prostate, and pancreas. These cultured clumps are tiny enough to be sustained without a blood supply, but large and diverse enough in their cell compositions to tell us something about tissue development and whole-organ physiology.
A typical organoid protocol starts with isolated embryonic or pluripotent stem cells. Scientists culture the cells in a proteinaceous matrix (such as Matrigel) that supports three-dimensional growth. After a set period of time the organoids grow mature enough for study, or for engrafting into a mouse to allow them to further develop. Researchers then harvest the organoids and slice them for immunohistochemistry, funnel them through a flow cytometer to study their cell surface markers, or blend them for PCR.
Of course, the devil’s in the details. Although the field of organoid research is maturing rapidly (see “2013’s Big Advances in Science,” The Scientist, December 24, 2013), with some organoids already moving into clinical studies to test drug efficacy, culture methods are still in their infancy, says Michael Shen, professor of medicine and of genetics and development at Columbia University in New York City. “Certainly there are different ways to pursue organoid culture, and some of these are just beginning to be explored. I don’t think we’re at the point yet where this is all entirely cookbook.”
The Scientist talked with researchers about how they’re producing organoids, and what beginners should know. Here’s what we learned.
BRAIN BEADS
Researcher: Madeline Lancaster, group leader, MRC Laboratory of Molecular Biology, Cambridge, U.K.
Project: Understanding early brain development and disease using organoids cultured from human stem cells
Background: In 2013, as a postdoctoral researcher in the lab of Jürgen Knoblich at the Institute of Molecular Biotechnology in Vienna, Austria, Lancaster developed organoids from neural stem cells that she had been studying in 2-D culture conditions. She used the method to coax human induced pluripotent stem cells into brain organoids in order to understand the biology of microcephaly, a disorder that is difficult to re-create in animal models (Nature, 501:373-79, 2013).
Researchers have adopted Lancaster’s methods to create models of embryonic brain development, analogous to what happens in the first trimester of pregnancy, and to probe the molecular mechanisms of brain disorders, including autism, schizophrenia, and neurodegenerative diseases such as Parkinson’s and Alzheimer’s.
Getting started: The group’s protocol addresses some of the common questions asked by new users and provides photos showing the appearance of healthy organoids (Nat Protoc, 9:2329-40, 2014).
For those well versed in cell and tissue culture, the time and financial investment required to delve into organoids is minimal, Lancaster says. You need two main things: Matrigel (the supportive structure that allows the organoids to develop into more complex tissue) and equipment that will allow you to agitate the organoids to enhance nutrient and oxygen exchange in the media, making bigger organoids possible. If you don’t have a spinning bioreactor, you can use an orbital shaker set inside a standard tissue culture incubator.
Considerations: You should closely characterize the first few batches using RT-PCR or immunofluorescence to check for the expression of certain genes that indicate the organoids are indeed brain cells, Lancaster says.
Researchers studying neurodegeneration might consider examining their organoids starting at about four months. Although the organoids survive for up to 15 months, by that time they don’t look healthy. They start to decline at around six or seven months, as the neurons begin to disappear and are replaced by glia.
Tip: It takes some time and practice to develop an eye for healthy organoids. A good way to learn is to take pictures of your organoids as they develop. “You can always look back and say, ‘Oh, at that point I think it started going bad,’” Lancaster says.
Cost: Roughly $150 per organoid (not including equipment), according to Lancaster’s calculations
Looking ahead: Lancaster has already tweaked the method to improve the reproducibility, using a combination of timing and media formulations, and some new additives. She expects to publish a revised protocol by the end of the year.
GUTSY GLOBS
INTIMATING INTESTINE: Mini-gut methods are the most established of organoid protocols. Proliferating epithelial cells in small intestinal aggregations from mouse (green, left) and human (pink, right) will pave the way for patient-specific organoids.COURTESY OF HELMRATH LABResearcher: Maxime Mahé, postdoctoral research fellow inMichael Helmrath’s lab at Cincinnati Children’s Hospital Medical Center, Ohio
Project: Understanding gastrointestinal development and homeostasis and generating patient-specific organoids for study
Background: The intestinal epithelial layer is made up of tiny, slender projections, called villi, resembling the strands of a shag carpet. The nooks formed at the bases of the villi, known as crypts, are home to intestinal stem cells responsible for constant renewal of the intestinal lining. Building on Sato’s protocol, Mahé added two new twists: he used manual dissection to extract the crypts, rather than shaking the tissue to dissociate the cells; and he added a small-molecule activator of the Wnt3A pathway to boost expansion of the cells (Curr Protoc Mouse Biol, 3:217-40, 2013).
Helmrath’s group grew such “enteroids” from intestinal stem cells isolated from the crypts of surgically removed human intestine. In principle, such organoids could be developed from the tissue of specific patients for diagnostic and clinical uses. A video protocol is available in the Journal of Visualized Experiments (doi: 10.3791/52483, 2015).
Getting started: It takes five or six attempts to get comfortable with the procedure, especially mastering the hardest part: the initial dissection. “The tissue is not always the same; it’s not something you can standardize,” Mahé says. “Sometimes you get a high number of crypts, sometimes you have a few.”
Tip: Many questions about cell proliferation, migration, and differentiation can be answered using in vitro organoids, Mahé says. “You save time, you save money, you save animals as well.” After that, you might consider moving into an animal model, depending on your goals: for example, to see muscle development, you should work in vivo, Mahé adds.
Looking ahead: The group is still working to be able to efficiently engraft human adult intestinal stem cell–derived organoids into mice. Although their first attempts were unsuccessful, they have since generated organoids for research from human embryonic stem cells (ESCs) and human induced pluripotent stem cells (iPSCs) derived by reprogramming fibroblasts. When organoids created from the either type of pluripotent stem cells are engrafted into immunodeficient mice to allow the cells to mature further, they develop into a human intestine (Nat Med, 20:1310-14, 2014), which may eventually lead to bioengineering a custom human intestine.
Cost: The Helmrath group spends roughly $150/sample in reagents to culture their organoids for a month. The medical center’s Pluripotent Stem Cell Facility provides training for a fee, and sells human intestinal organoids for roughly $400/plate (which contains 20–30 organoids).
B-CELL BALLS
PROSTRATE PROGRESS: Researchers have grown prostate organoids that consist of basal cells (green/blue) and luminal cells (red/blue).MAHO SHIBATAResearcher: Ankur Singh, assistant professor of mechanical and aerospace engineering, Cornell University
Project: In vitro modeling of immune reactions in mice
Background: When naive B cells in the body are exposed to antigens, they form clumps of cells called germinal centers in a lymph node or the spleen, where they proliferate, mutate to generate high-affinity antibodies, and undergo clonal expansion. Until now, this process has been difficult to recapitulate in vitro. Adding the necessary (stromal) support cells to primary naive B cells and culturing them in 2-D does not enable them to differentiate into cells resembling those from germinal centers, Singh says. Unlike stem cells, naive B cells do not tend to grow in clusters, so they need a little extra help.
Rather than using the conventional Matrigel for 3-D culture, Singh and his collaborators developed a gelatin and silicate-nanoparticle mix that mimics the softness of the body’s lymphoid organs. Within four to six days, the B cells in these organoids mature—100 times faster than B cells in 2-D culture—and produce two classes of antibodies important for fighting infections. The scientists use collagenase to dissolve the gel and harvest the organoid’s cells for analysis using flow cytometry. These new germinal center organoids were described this year in Biomaterials (63:24-34).
Getting started: Making the gelatin-nanoparticle mix is as easy as making Jell-O at home, Singh says, and the ingredients are commercially available. You’ll need experience with animal dissection (the necessary starting point is isolation of naive B cells from the spleen) and with cell culture. Once these techniques have been mastered, it takes roughly one week to get your first batch of organoids with mature antibody-producing cells.
Considerations: Singh’s group has already determined an optimal gelatin-nanoparticle ratio (2% gelatin/1.5% nanoparticle), but if you you’re using genetically mutated B cells, you may need to tweak the ratios. “It can be easily tuned,” Singh says.
Tip: After four days of incubating the cells with gel, you will see dark spots—a sign that the cells are proliferating and that you’re on the right track.
Cost: Not including the cost of generating immortalized stromal cell lines, it costs roughly $1 to produce one germinal center.
Looking ahead: Eventually, Singh’s group hopes to adapt the technique for use with patient-specific stem cells, though it has proven challenging to produce immune cells from stem cells. “It’s a very complicated process,” says Singh, “[but] it will happen one day in the context of this system.”
PROSTATE PELLETS
Researcher: Michael Shen, professor of medicine and of genetics and development, Columbia University Medical Center, New York
Project: Understanding basic prostate regeneration and prostate cancer
Background: In 2009, Shen’s group discovered a rare population of stem cells from which prostate cancer can originate (Nature, 461:495-500, 2009). Calling them CARNS, for castration-resistant Nkx3.1-expressing cells, the group knew they would face challenges culturing the cells because they are a type of luminal epithelial cell, which had historically proven difficult to expand using 2-D methods. “We thought if any type of approach would succeed it would be 3-D,” Shen recalls.
Through a trial-and-error approach, postdoctoral researcher Chee Wai Chua eventually converted mouse CARNS into organoids (Nat Cell Biol, 16:951-61, 2014). The resulting cell types and tissue architecture resembled those characteristic of normal prostate epithelium. The researchers then engrafted the organoids into mice to generate prostatic tissues.
Getting started: Shen’s group has made their method available via the Nature Protocol Exchange. The most difficult part for beginners is the initial tissue-dissociation step, which is typical of any organoid protocol. “To work out the details of how to do this is not straightforward,” Shen says. “In our case, we’re still working on this. We’re continually seeking to improve dissociation conditions.”
Considerations: When applied to the prostate, Clevers’s conditions seem to favor the growth of a different type of prostate cell known as a basal cell, though his group also grew luminal cells. Shen’s conditions are less defined than those of Clevers, using serum instead of specific growth factors. Shen’s group doesn’t know exactly which growth factors in the serum drive organoid growth and development.
Tip: If you are making the organoids from normal prostate for the first time, you might consider assessing their response to androgen deprivation. They should lose expression of Nkx3.1 in response to this condition.
Cost: It costs $1 or less for one mouse prostate organoid (not counting animal, equipment or labor costs).
Looking ahead: The group has been able to create organoids derived from human prostate cells, but determining the ideal conditions for these cells is still a work in progress, Shen says.
Tags
techniques, organoids, disease/medicine and 3-D cell culture
Aurelian Udristioiu commented on your update
“The human body emits low levels light, heat, and acoustical energy, these wavelengths of radiations having the electrical and magnetic properties and may also to be transformed in kinds of energy that cannot be easily defined by classical physical sciences and chemistry. In last time most researches has focused on electromagnetic aspects of the bio-magnetic field Bio-energetic fluids can be used in technology of preparation of drugs, from homeopath medicine and in laboratory medicine by the changes of pH in liquid medium with cultivated stem cells for to prolong the span life of cells, in view of cell-stem transplantation in chronic diseases. ”
Umbilical Cord Blood Contains c-kit+ Cells that Can Differentiate into Heart-like Cells
https://beyondthedish.wordpress.com/2015/09/10/umbilical-cord-blood-contains-c-kit-cells-that-can-differentiate-into-heart-like-cells/
Bone contains a wide variety of stem cells whose potential are only beginning to be tapped. One cell population possesses a cell surface protein called c-kit, and these c-kit+ progenitor cells seem to support myocardial regeneration. Do c-kit+ cells from umbilical cord blood have the same capacity?
Luciana Teofili from the Catholic University of the Sacred Heart in Rome, Italy and her colleagues purified c-kit+ cells from umbilical cord blood by means of magnetic beads that were coated with c-kit-binding antibodies. Teofili and others induced heart muscle differentiation in these cells with several different protocols. Then the expression of cardiac markers (GATA4, GATA6, β-myosin heavy chain, α-sarcomeric actin and cardiac Troponin T) was investigated, and whole-cell current and voltage-clamp recordings were performed.
The c-kit+ cells from umbilical cord blood showed a rather immature gene profile, and by themselves, they did not differentiate into heart muscle-like cells in culture. In contrast, if whole mononuclear cells from umbilical cord blood were subjected to the same treatment, several if the employed protocols produced large, adherent cells that expressed several heart muscle-specific genes and exhibited an excitability much like that of heart muscle cells.
Formation of these heart muscle-like cells was prevented if the c-kit+ cells were removed from the other cells. Tracking studies showed that the c-kit+ cells were the ones that differentiated into heart muscle-like cells, but they only did so when they were together with c-kit– cells.
Thus umbilical cord blood contains progenitors endowed with the ability to differentiate into heart muscle-like cells. The cells with this potential reside in the c-kit+ fraction but they require the presence of abundant accessory cells to differentiate properly.
These preliminary observations suggest that it is a good idea to consider the storage of the umbilical cord blood of patients with prenatal diagnosis of congenital heart diseases. Such conditions might be potentially amenable to myocardial regenerative therapies with umbilical blood-based stem cells.
This paper was published in the journal Cytotherapy, but it must be said that the evidence that these cells differentiated into heart muscle cells was not completely convincing.
Directed Neural Differentiation of Induced Pluripotent Stem Cells in the Marmoset
Peter J. Hornsby Ph.D. | 10th-Sep-2015
http://medical.wesrch.com/paper-details/pdf-ME1XXFT06ILUR-directed-neural-differentiation-of-induced-pluripotent-stem-cells-in-the-marmoset#page1
|
|
Description: Personalized cell therapy: The marmoset as a model- Before personalized cell therapy is used in humans, need to move beyond rodent models, Beyond rodents, nonhuman primates play key roles, Within nonhuman primates, the marmoset is a suitable size and life span for stem cell studies, Has been used in drug studies and in disease models, e.g. Parkinson’s disease, The marmoset was the first nonhuman primate to have transgenics with germline transmission, The second nonhuman primate (after the rhesus macaque) for which induced pluripotent stem cells were derived (our work, 2010).
DMSO treatment/differentiation: Conclusions- Despite some differences in growth characteristics of 3 marmoset iPS cell lines, all can be directed to a uniform pattern of neural differentiation by prior exposure to 24 h DMSO, The optimal DMSO concentration should be determined for each cell line, Therefore we should be able to differentiate any given (newly created) iPS cell population “on demand” by a protocol similar to the one used here.
Progress so far; next step- Marmoset iPS cells generated by a reproducible reprogramming method, Many marmoset iPS cell lines continuously grown for >1 year – immortal; maintain pluripotency, Rapid differentiation into the neural lineage using combinations of drugs with iterative testing, Rapid reprogramming of samples from living individuals, Rapid differentiation of living individual iPS cells. .
|
Like this:
Like Loading...
Read Full Post »