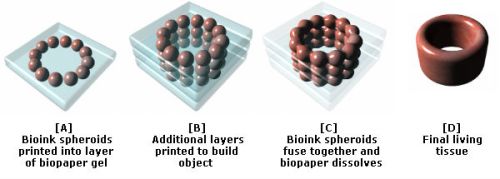
Use of 3D Bioprinting for Development of Toxicity Prediction Models
Curator: Stephen J. Williams, PhD
SOT FDA Colloquium on 3D Bioprinted Tissue Models: Tuesday, April 9, 2019
The Society of Toxicology (SOT) and the U.S. Food and Drug Administration (FDA) will hold a workshop on “Alternative Methods for Predictive Safety Testing: 3D Bioprinted Tissue Models” on Tuesday, April 9, at the FDA Center for Food Safety and Applied Nutrition in College Park, Maryland. This workshop is the latest in the series, “SOT FDA Colloquia on Emerging Toxicological Science: Challenges in Food and Ingredient Safety.”
Human 3D bioprinted tissues represent a valuable in vitro approach for chemical, personal care product, cosmetic, and preclinical toxicity/safety testing. Bioprinting of skin, liver, and kidney is already appearing in toxicity testing applications for chemical exposures and disease modeling. The use of 3D bioprinted tissues and organs may provide future alternative approaches for testing that may more closely resemble and simulate intact human tissues to more accurately predict human responses to chemical and drug exposures.
A synopsis of the schedule and related works from the speakers is given below:
| 8:40 AM–9:20 AM |
Overview and Challenges of Bioprinting
Sharon Presnell, Amnion Foundation, Winston-Salem, NC |
| 9:20 AM–10:00 AM |
Putting 3D Bioprinting to the Use of Tissue Model Fabrication
Y. Shrike Zhang, Brigham and Women’s Hospital, Harvard Medical School and Harvard-MIT Division of Health Sciences and Technology, Boston, MA |
| 10:00 AM–10:20 AM |
Break |
| 10:20 AM–11:00 AM |
Uses of Bioprinted Liver Tissue in Drug Development
Jean-Louis Klein, GlaxoSmithKline, Collegeville, PA |
| 11:00 AM–11:40 AM |
Biofabrication of 3D Tissue Models for Disease Modeling and Chemical Screening
Marc Ferrer, National Center for Advancing Translational Sciences, NIH, Rockville, MD |
Sharon Presnell, Ph.D. President, Amnion Foundation
Dr. Sharon Presnell was most recently the Chief Scientific Officer at Organovo, Inc., and the President of their wholly-owned subsidiary, Samsara Sciences. She received a Ph.D. in Cell & Molecular Pathology from the Medical College of Virginia and completed her undergraduate degree in biology at NC State. In addition to her most recent roles, Presnell has served as the director of cell biology R&D at Becton Dickinson’s corporate research center in RTP, and as the SVP of R&D at Tengion. Her roles have always involved the commercial and clinical translation of basic research and early development in the cell biology space. She serves on the board of the Coulter Foundation at the University of Virginia and is a member of the College of Life Sciences Foundation Board at NC State. In January 2019, Dr. Presnell will begin a new role as President of the Amnion Foundation, a non-profit organization in Winston-Salem.
A few of her relevant publications:
Bioprinted liver provides early insight into the role of Kupffer cells in TGF-β1 and methotrexate-induced fibrogenesis
Integrating Kupffer cells into a 3D bioprinted model of human liver recapitulates fibrotic responses of certain toxicants in a time and context dependent manner. This work establishes that the presence of Kupffer cells or macrophages are important mediators in fibrotic responses to certain hepatotoxins and both should be incorporated into bioprinted human liver models for toxicology testing.
Bioprinted 3D Primary Liver Tissues Allow Assessment of Organ-Level Response to Clinical Drug Induced Toxicity In Vitro
Abstract: Modeling clinically relevant tissue responses using cell models poses a significant challenge for drug development, in particular for drug induced liver injury (DILI). This is mainly because existing liver models lack longevity and tissue-level complexity which limits their utility in predictive toxicology. In this study, we established and characterized novel bioprinted human liver tissue mimetics comprised of patient-derived hepatocytes and non-parenchymal cells in a defined architecture. Scaffold-free assembly of different cell types in an in vivo-relevant architecture allowed for histologic analysis that revealed distinct intercellular hepatocyte junctions, CD31+ endothelial networks, and desmin positive, smooth muscle actin negative quiescent stellates. Unlike what was seen in 2D hepatocyte cultures, the tissues maintained levels of ATP, Albumin as well as expression and drug-induced enzyme activity of Cytochrome P450s over 4 weeks in culture. To assess the ability of the 3D liver cultures to model tissue-level DILI, dose responses of Trovafloxacin, a drug whose hepatotoxic potential could not be assessed by standard pre-clinical models, were compared to the structurally related non-toxic drug Levofloxacin. Trovafloxacin induced significant, dose-dependent toxicity at clinically relevant doses (≤ 4uM). Interestingly, Trovafloxacin toxicity was observed without lipopolysaccharide stimulation and in the absence of resident macrophages in contrast to earlier reports. Together, these results demonstrate that 3D bioprinted liver tissues can both effectively model DILI and distinguish between highly related compounds with differential profile. Thus, the combination of patient-derived primary cells with bioprinting technology here for the first time demonstrates superior performance in terms of mimicking human drug response in a known target organ at the tissue level.
A great interview with Dr. Presnell and the 3D Models 2017 Symposium is located here:
Please click here for Web based and PDF version of interview
Some highlights of the interview include
- Exciting advances in field showing we can model complex tissue-level disease-state phenotypes that develop in response to chronic long term injury or exposure
- Sees the field developing a means to converge both the biology and physiology of tissues, namely modeling the connectivity between tissues such as fluid flow
- Future work will need to be dedicated to develop comprehensive analytics for 3D tissue analysis. As she states “we are very conditioned to get information in a simple way from biochemical readouts in two dimension, monocellular systems” however how we address the complexity of various cellular responses in a 3D multicellular environment will be pertinent.
- Additional challenges include the scalability of such systems and making such system accessible in a larger way
- Shrike Zhang, Brigham and Women’s Hospital, Harvard Medical School and Harvard-MIT Division of Health Sciences and Technology
Dr. Zhang currently holds an Assistant Professor position at Harvard Medical School and is an Associate Bioengineer at Brigham and Women’s Hospital. His research interests include organ-on-a-chip, 3D bioprinting, biomaterials, regenerative engineering, biomedical imaging, biosensing, nanomedicine, and developmental biology. His scientific contributions have been recognized by >40 international, national, and regional awards. He has been invited to deliver >70 lectures worldwide, and has served as reviewer for >400 manuscripts for >30 journals. He is serving as Editor-in-Chief for Microphysiological Systems, and Associate Editor for Bio-Design and Manufacturing. He is also on Editorial Board of Bioprinting, Heliyon, BMC Materials, and Essays in Biochemistry, and on Advisory Panel of Nanotechnology.
Some relevant references from Dr. Zhang
Multi-tissue interactions in an integrated three-tissue organ-on-a-chip platform.
Skardal A, Murphy SV, Devarasetty M, Mead I, Kang HW, Seol YJ, Shrike Zhang Y, Shin SR, Zhao L, Aleman J, Hall AR, Shupe TD, Kleensang A, Dokmeci MR, Jin Lee S, Jackson JD, Yoo JJ, Hartung T, Khademhosseini A, Soker S, Bishop CE, Atala A.
Sci Rep. 2017 Aug 18;7(1):8837. doi: 10.1038/s41598-017-08879-x.
Reconstruction of Large-scale Defects with a Novel Hybrid Scaffold Made from Poly(L-lactic acid)/Nanohydroxyapatite/Alendronate-loaded Chitosan Microsphere: in vitro and in vivo Studies.
Wu H, Lei P, Liu G, Shrike Zhang Y, Yang J, Zhang L, Xie J, Niu W, Liu H, Ruan J, Hu Y, Zhang C.
Sci Rep. 2017 Mar 23;7(1):359. doi: 10.1038/s41598-017-00506-z.
A liver-on-a-chip platform with bioprinted hepatic spheroids.
Bhise NS, Manoharan V, Massa S, Tamayol A, Ghaderi M, Miscuglio M, Lang Q, Shrike Zhang Y, Shin SR, Calzone G, Annabi N, Shupe TD, Bishop CE, Atala A, Dokmeci MR, Khademhosseini A.
Biofabrication. 2016 Jan 12;8(1):014101. doi: 10.1088/1758-5090/8/1/014101.
Marc Ferrer, National Center for Advancing Translational Sciences, NIH
Marc Ferrer is a team leader in the NCATS Chemical Genomics Center, which was part of the National Human Genome Research Institute when Ferrer began working there in 2010. He has extensive experience in drug discovery, both in the pharmaceutical industry and academic research. Before joining NIH, he was director of assay development and screening at Merck Research Laboratories. For 10 years at Merck, Ferrer led the development of assays for high-throughput screening of small molecules and small interfering RNA (siRNA) to support programs for lead and target identification across all disease areas.
At NCATS, Ferrer leads the implementation of probe development programs, discovery of drug combinations and development of innovative assay paradigms for more effective drug discovery. He advises collaborators on strategies for discovering small molecule therapeutics, including assays for screening and lead identification and optimization. Ferrer has experience implementing high-throughput screens for a broad range of disease areas with a wide array of assay technologies. He has led and managed highly productive teams by setting clear research strategies and goals and by establishing effective collaborations between scientists from diverse disciplines within industry, academia and technology providers.
Ferrer has a Ph.D. in biological chemistry from the University of Minnesota, Twin Cities, and completed postdoctoral training at Harvard University’s Department of Molecular and Cellular Biology. He received a B.Sc. degree in organic chemistry from the University of Barcelona in Spain.
Some relevant references for Dr. Ferrer
Fully 3D Bioprinted Skin Equivalent Constructs with Validated Morphology and Barrier Function.
Derr K, Zou J, Luo K, Song MJ, Sittampalam GS, Zhou C, Michael S, Ferrer M, Derr P.
Tissue Eng Part C Methods. 2019 Apr 22. doi: 10.1089/ten.TEC.2018.0318. [Epub ahead of print]
Determination of the Elasticity Modulus of 3D-Printed Octet-Truss Structures for Use in Porous Prosthesis Implants.
Bagheri A, Buj-Corral I, Ferrer M, Pastor MM, Roure F.
Materials (Basel). 2018 Nov 29;11(12). pii: E2420. doi: 10.3390/ma11122420.
Mutation Profiles in Glioblastoma 3D Oncospheres Modulate Drug Efficacy.
Wilson KM, Mathews-Griner LA, Williamson T, Guha R, Chen L, Shinn P, McKnight C, Michael S, Klumpp-Thomas C, Binder ZA, Ferrer M, Gallia GL, Thomas CJ, Riggins GJ.
SLAS Technol. 2019 Feb;24(1):28-40. doi: 10.1177/2472630318803749. Epub 2018 Oct 5.
A high-throughput imaging and nuclear segmentation analysis protocol for cleared 3D culture models.
Boutin ME, Voss TC, Titus SA, Cruz-Gutierrez K, Michael S, Ferrer M.
Sci Rep. 2018 Jul 24;8(1):11135. doi: 10.1038/s41598-018-29169-0.
A High-Throughput Screening Model of the Tumor Microenvironment for Ovarian Cancer Cell Growth.
Lal-Nag M, McGee L, Guha R, Lengyel E, Kenny HA, Ferrer M.
SLAS Discov. 2017 Jun;22(5):494-506. doi: 10.1177/2472555216687082. Epub 2017 Jan 31.
Exploring Drug Dosing Regimens In Vitro Using Real-Time 3D Spheroid Tumor Growth Assays.
Lal-Nag M, McGee L, Titus SA, Brimacombe K, Michael S, Sittampalam G, Ferrer M.
SLAS Discov. 2017 Jun;22(5):537-546. doi: 10.1177/2472555217698818. Epub 2017 Mar 15.
RNAi High-Throughput Screening of Single- and Multi-Cell-Type Tumor Spheroids: A Comprehensive Analysis in Two and Three Dimensions.
Fu J, Fernandez D, Ferrer M, Titus SA, Buehler E, Lal-Nag MA.
SLAS Discov. 2017 Jun;22(5):525-536. doi: 10.1177/2472555217696796. Epub 2017 Mar 9.
Other Articles on 3D Bioprinting on this Open Access Journal include:
Global Technology Conferences on 3D BioPrinting 2015 – 2016
3D Medical BioPrinting Technology Reporting by Irina Robu, PhD – a forthcoming Article in “Medical 3D BioPrinting – The Revolution in Medicine, Technologies for Patient-centered Medicine: From R&D in Biologics to New Medical Devices”
Bio-Inks and 3D BioPrinting
New Scaffold-Free 3D Bioprinting Method Available to Researchers
Gene Editing for Gene Therapies with 3D BioPrinting
Read Full Post »