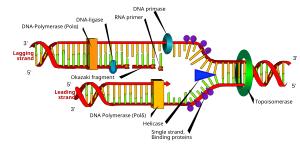
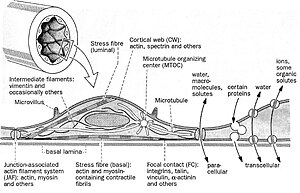
English: A diagram of cellular respiration including glycolysis, Krebs cycle, citric acid cycle, and the electron transport chain (Photo credit: Wikipedia)
Reporter and Curator: Larry H Bernstein, MD, FACP
Introduction
Mitochondria are essential for life, and are critical for the generation of ATP. Otto Warburg won the Nobel Prize in 1918 for his studies of respiration and he described a situation of impaired respiration in cancer cells causing them to produce lactic acid, like bacteria. This has been termed facultative anaerobic glycolysis. The metabolic explanation for mitochondrial respiration had to await the Nobel discoveries of the Krebs cycle and high energy ~P in acetyl CoA by Fritz Lippman. The Krebs cycle generates 16 ATPs I respiration compared to 2 ATPs through glycolysis. The discovery of the genetic code with the “Watson-Crick” model and the identification of DNA polymerase opened a window for contuing discovery leading to the human genome project at 20th century end that has now been followed by “ENCODE” in the 21st century. This review opens a rediscovery of the metabolic function of mitochondria and adaptive functions with respect to cancer and other diseases.
Function in aerobic and anaerobic metabolism
Two-carbon compounds – the TCA, the pentose phosphate pathway, together with gluconeogenesis and the glyoxylate cycle are essential for the provision of anabolic precursors. Yeast environmental diversity mostly leads to a vast metabolic complexity driven by carbon and the energy available in environmental habitats. This resulted in much early research on analysis of yeast metabolism associated with glucose catabolism in Saccharomyces cerevisiae, under both aerobic and anaerobic environments. Yeasts may be physiologically classified with respect to the type of energy-generating process involved in sugar metabolism, namely non-, facultative- or obligate fermentative. The nonfermentative yeasts have exclusively a respiratory metabolism and are not capable of alcoholic fermentation from glucose, while the obligate-fermentative yeasts – “natural respiratory mutants” – are only capable of metabolizing glucose through alcoholic fermentation. Most of the yeasts identified are facultative-fermentative ones, and depending on the growth conditions, the type and concentration of sugars and/or oxygen availability, may display either a fully respiratory or a fermentative metabolism or even both in a mixed respiratory-fermentative metabolism (e.g., S. cerevisiae). The sugar composition of the media and oxygen availability are the two main environmental conditions that have a strong impact on yeast metabolic physiology, and three frequently observed effects associated with the type of energy-generating processes involved in sugar metabolism and/or oxygen availability are Pasteur, Crabtree and Custer. In modern terms the Pasteur effect refers to an activation of anaerobic glycolysis in order to meet cellular ATP demands owing to the lower efficiency of ATP production by fermentation compared with respiration. In 1861 Pasteur observed that S. cerevisiae consume much more glucose in the absence of oxygen than in its presence. S. cerevisiae only shows a Pasteur at low growth rates and at resting-cell conditions, where a high contribution of respiration to sugar catabolism occurs owing to the loss of fermentative capacity. The Crabtree effect is defined as the occurrence of alcoholic fermentation under aerobic conditions, explained by a theory involving “limited respiratory capacities” in the branching point of pyruvate metabolism. The Custer effect is known as the inhibition of alcoholic fermentation by the absence of oxygen. It is thought that the Custer effect is caused by reductive stress.
Glycolysis
Once inside the cell, glucose is phosphorylated by kinases to glucose 6-phosphate and then isomerized to fructose 6-phosphate, by phosphoglucose isomerase. The next enzyme is phospho-fructokinase, which is subject to regulation by several metabolites, and further phosphorylates fructose 6-phosphate to fructose 1,6-bisphosphate. These steps of glycolysis require energy in the form of ATP. Glycolysis leads to pyruvate formation associated with a net production of energy and reducing equivalents. Approximately 50% of glucose 6-phosphate is metabolized via glycolysis and 30% via the pentose phosphate pathway in Crabtree negative yeasts. However, about 90% of the carbon going through the pentose phosphate pathway reentered glycolysis at the level of fructose 6-phosphate or glyceraldehyde 3-phosphate. The pentose phosphate pathway in Crabtree positive yeasts (S. cerevisiae) is predominantly used for NADPH production but not for biomass production or catabolic reactions.
Pyruvate branch point. At the pyruvate (the end product of glycolysis) branching point, pyruvate can follow three different metabolic fates depending on the yeast species and the environmental conditions. On the other hand, the carbon flux may be distributed between the respiratory and fermentative pathways. Pyruvate might be directly converted to acetyl–cofactor A (CoA) by the mitochondrial multienzyme complex pyruvate dehydrogenase (PDH) after its transport into the mitochondria by the mitochondrial pyruvate carrier. Alternatively, pyruvate can also be converted to acetyl–CoA in the cytosol via acetaldehyde and to acetate by the so-called PDH-bypass pathway. Compared with cytosolic pyruvate decarboxylase, the mitochondrial PDH complex has a higher affinity for pyruvate and therefore most of the pyruvate will flow through the PDH complex at low glycolytic rates. However, at increasing glucose concentrations, the glycolytic rate will increase and more pyruvate is formed, saturating the PDH bypass and shifting the carbon flux through ethanol production. In the yeast S. cerevisiae, the external glucose level controls the switch between respiration and fermentation.
Rodrigues F, Ludovico P and Leão C. Sugar Metabolism in Yeasts: an Overview of Aerobic and Anaerobic Glucose Catabolism. In Molecular and Structural Biology. Chapter 6. qxd 07/23/05 P117
Eriksson P, Andre L, Ansell R, Blomberg A, Adler L (1995) Cloning and characterization of GPD2, a second gene encoding sn-glycerol 3-phosphate dehydrogenase (NAD+) in Saccharomyces cerevisiae, and its comparison with GPD1. Mol Microbiol 17:95–107.
Flikweert MT, van der Zanden L, Janssen WM, Steensma HY, van Dijken JP, Pronk JT (1996)Pyruvate decarboxylase: an indispensable enzyme for growth of Saccharomyces cerevisiae on glucose. Yeast 12:247–257.
Biogenesis of mitochondrial structures from aerobically grown S. cerevisiae
Under aerobic conditions S. cerevisiae forms mitochondria which are classical in their properties,
but the number, morphology, and enzyme activity of these mitochondria are also affected by catabolite repression, but it cannot respire under anaerobic conditions and lacks cytochromes. These structures were isolated from anaerobically grown yeast cells and contain malate and succinate dehydrogenases, ATPase, and DNA characteristic of yeast mitochondria. These lipid-complete structures consist predominantly of double-membrane vesicles enclosing a dense matrix which contains a folded inner membrane system bordering electron-transparent regions similar to the cristae of mitochondria.
- The morphology of the structures is critically dependent on their lipid composition
- Their unsaturated fatty acid content is similar to that of mitochondria from aerobically grown cells
- The structures from cells grown without lipid supplements have simpler morphology – a dense granular matrix surrounded by a double membrane but have no obvious folded inner membrane system within the matrix
- The lipid-depleted structures are only isolated in intact form from protoplasts
- The synthesis of ergosterol and unsaturated fatty acids is oxygen-dependent and anaerobically grown cells may be depleted of these lipid components
- The cytology of anaerobically grown yeast cells is profoundly affected by both lipid-depletion and catabolite repression
- Lipid-depleted anaerobic cells, membranous mitochondrial profiles were not demonstrable
- The structures from the aerobically and anaerobically grown cells are markedly different in morphology and fatty acid composition, but both contain mitochondrial DNA and a number of mitochondrial enzymes
The phospholipid composition of various strains of Saccharomyces cerevisiae, wild type and petite (cytoplasmic respiratory deficient) yeasts and derived mitochondrial mutants grown under conditions designed to induce variations in the complement of mitochondrial were fractionated into various subcellular fractions and analyzed for cytochrome oxidase (in wild type) and phospholipid composition . 90% or more of the phospholipid, cardiolipin was found in the mitochondrial membranes of wild type and petite yeast . Cardiolipin content differed markedly under various growth conditions .
- Stationary yeast grown in glucose had better developed mitochondria and more cardiolipin than repressed log phase yeast .
- Aerobic yeast contained more cardiolipin than anaerobic yeast .
- Respiration-deficient cytoplasmic mitochondrial mutants, both suppressive and neutral, contained less cardiolipin than corresponding wild types .
- A chromosomal mutant lacking respiratory function had normal cardiolipin content .
- Log phase cells grown in galactose and lactate, which do not readily repress the development of mitochondrial membranes, contained as much cardiolipin as stationary phase cells grown in glucose .
- Cytoplasmic mitochondrial mutants respond to changes in the glucose concentration of the growth medium by variations in their cardiolipin content in the same way as wild type yeast does under similar growth conditions.
- It is of interest that the chromosomal petite, which as far as can be ascertained has qualitatively normal mitochondrial DNA and a normal cardiolipin content when grown under maximally derepressed conditions .
Thus, the genetic defect in this case probably does not diminish the mass of inner mitochondrial membrane under appropriate conditions . This suggests the cardiolipin content of yeast is a good indicator of the state of development of mitochondrial membrane.
Jakovcic S, Getz Gs, Rabinowitz M, Jakob H, Swift H. Cardiolipin Content Of Wild Type and Mutant Yeasts in Relation to Mitochondrial Function and Development. JCB 1971. jcb.rupress.org
Jakovcic S, Haddock J, Getz GS, Rabinowitz M, Swift H. Biochem J. 1971; 121 :341 .
EPHRUSSI, B . 1953 . Nucleocytoplasmic Relations in Microorganisms . Clarendon Press, Oxford.
Mitochondria, hydrogenosomes and mitosomes
Before and after the publication of an unnoticed article in 1905 by Mereschkowsky there were many publications dealing with plant “chimera’s” and cytoplasmic inheritance in plants, which should have favoured the interpretation of plastids as “semi-autonomous” symbiotic entities in the cytoplasm of the eukaryotic plant cell. Twenty years after Mereschkowsky’s plea for an endosymbiotic origin of plastids, Wallin (1925, 1927) postulated the “bacterial nature of mitochondria”. And so it is one of the mysteries of the 20th century that an endosymbiotic origin of plastids had not been generally accepted before the 1970s, primarily because one cannot experience the consequences of mutations in the mitochondrial genome by naked eye.
- Mitochondrial DNA is usually present in multiple copies in one and the same mitochondrion and those in the hundreds to thousands of mitochondria in a single cell are not necessarily identical.
- The random partitioning of the mitochondria in mitosis (and meiosis) frequently results in a more or less biased distribution of the diverent mitochondria in the daughter cells, eventually causing diverent phenotypes in different tissues obscuring the maternal inheritance
- It was not until the 1990s that certain diseases—which had been interpreted as being X-chromosomal with incomplete penetrance—eventually turned out to be
Lastly, the vast majority of mitochondrial proteins are encoded in the nucleus and, consequently, mutations in the corresponding genes exhibit a Mendelian, and not a cytoplasmic, maternal inheritance
In the 1970s and 1980s the unequivocal demonstration of mitochondrial DNA occurred
and mitochondrial mutations at the DNA level provided the final proof for the role of such mutations in a wealth of hereditary diseases in man.
- The genomics era provided the tools to prove the endosymbiont-hypothesis for the origin of the eukaryotic cell
Since DNA does not arise de novo, the genomes of organisms and organelles provide a historical record for the evolution of the eukaryotic cell and its organelles. The DNA sequences of two to three genomes of the eukaryotic cell turned out to be a record of the evolution of the eukaryotic life on earth. The analysis of organelle genomes unequivocally revealed a cyanobacterial origin for plastids and an -proteobacterial origin for mitochondria. Both plastids and mitochondria appear to be monophyletic, i.e. plastids derived from one and the same cyanobacterial ancestor, and mitochondria from one and the same -proteobacterial ancestor.
The evolution of the eukaryotic cell appears to have involved one (in the case of animals) or two (in the case of plants) events that took place 1.5 to 2 billion years ago. However, it appears that symbioses involving one or the other eubacterium arose repeatedly during the billions of years available. For example, photosynthetic algae by phagotrophic eukaryotes, negating the hypothesis of a single eukaryotic event, rather than stringent selection shaping the diversity of present-day life. Recent hypotheses for the origin of the nucleus have postulated that introns, which could be acquired by the uptake of the -proteobacterial endosymbiont, forced the nucleus-cytosol compartmentalization. Lateral gene transfer among eukaryotes is more frequent than was assumed earlier, and “mitochondrial genes” in the nuclear genomes of amitochondrial organisms are not necessarily the consequence of a transient presence of a DNA-containing mitochondrial-like organelle.
To cope with the obvious ubiquity of “mitochondrial” genes and the chimerism of the DNA of present day eukaryotes, the hydrogen hypothesis postulates that an archaeal host took up a eubacterial symbiont that became the ancestor of mitochondria and hydrogenosomes. The hydrogen hypothesis has the potential to explain both the monophyly of the mitochondria, and the existence of “anaerobic” and “aerobic” variants of one and the same original organelle. Based on these observations we have only the terms “mitochondrion”, “hydrogenosome” and “mitosome” to classify the various variants of the mitochondrial family.
Hackstein JHP, Joachim Tjaden J , Huynen M. Mitochondria, hydrogenosomes and mitosomes: products of evolutionary tinkering! Curr Genet (2006) 50:225–245. DOI 10.1007/s00294-006-0088-8.
Lineages
A look at the phylogenetic distribution of characterized anaerobic mitochondria among animal lineages shows that these are not clustered but spread across metazoan phylogeny. The biochemistry and the enzyme equipment used in the facultatively anaerobic mitochondria of metazoans is nearly identical across lineages, strongly indicating a common origin from an archaic metazoan ancestor. The organelles look like hydrogenosomes – anaerobic forms of mitochondria that generate H2 and adenosine triphosphate (ATP) from pyruvateoxidation and which were previously found only in unicellular eukaryotes. The animals harbor structures resembling prokaryotic endosymbionts, reminiscent of the methanogenic endosymbionts found in some hydrogenosome-bearing protists; fluorescence of F420, a typical methanogen cofactor, or lack thereof, will bring more insights as to what these structures are. If we follow the anaerobic lifestyle further back into evolutionary history, beyond the origin of the metazoans, we see that the phylogenetic distribution of eukaryotes with facultative anaerobic mitochondria, eukaryotes with hydrogenosomes and eukaryotes that possess mitosomes (reduced forms of mitochondria with no direct role in ATP synthesis) the picture is similar to that seen for animals. In all six of the major lineages (or supergroups) of eukaryotes that are currently recognized, forms with anaerobic mitochondria have been found. The newest additions to the growing collection of anaerobic mitochondrial metabolisms are the denitrifying foraminiferans. A handful of about a dozen enzymes make the difference between a ‘normal’ O2-respiring mitochondrion found in mammals, and the energy metabolism of eukaryotes with anaerobic mitochondria, hydrogenosomes or mitosomes. Notably, the full complement of those enzymes, once thought to be specific to eukaryotic anaerobes, surprisingly turned up in the green alga Chlamydomonas reinhardtii , which produces O2 in the light, has typical O2-respiring mitochondria but, within about 30 min of exposure to heterotrophic, anoxic and dark conditions, expresses its anaerobic biochemistry to make H2 in the same way as trichomonads, the group in which hydrogenosomes were discovered. Chlamydomonas provides evidence which indicates that the ability to inhabit oxygen-harbouring, as well as anoxic environments, is an ancestral feature of eukaryotes and their mitochondria. The prokaryote inhabitants have existed for well over a billion years, and have reached this new habitat by dispersal, not by adaptive evolution de novo and in situ. Indeed, geochemical evidence has shown that methanogenesis and sulphate reduction, and the niches in which they occur, are truly ancient.
Mentel and Martin. Anaerobic mitochondria: more common all the time. BMC Biology 2010; 8:32. BioMed Central Ltd. http://www.biomedcentral.com/1741-7007/8/32.
Anaerobic mitochondrial enzymes
Mitochondria from the muscle of the parasitic nematode Ascaris lumbricoides var. suum function anaerobically in electron transport-associated phosphorylations under physiological conditions. These helminth organelles have been fractionated into inner and outer membrane, matrix, and inter-membrane space fractions. The distributions of enzyme systems were determined and compared with corresponding distributions reported in mammalian mitochondria. Succinate and pyruvate dehydrogenases as well as NADH oxidase, Mg++-dependent ATPase, adenylate kinase, citrate synthase, and cytochrome c reductases were determined to be distributed as in mammalian mitochondria. In contrast with the mammalian systems, fumarase and NAD-linked “malic” enzyme were isolated primarily from the intermembrane space fraction of the worm mitochondria. These enzymes are required for the anaerobic energy-generating system in Ascaris and would be expected to give rise to NADH in the intermembrane space.
Pyruvate kinase activity is barely detectable in Ascaris muscle. Therefore, rather than giving rise to cytoplasmic pyruvate, CO2 is fixed into phosphoenolpyruvate, resulting in the formation of oxalacetate which, in turn, is reduced by NADH to form malate regenerating glycolytic NAD . Ascaris muscle mitochondria utilize malate anaerobically as their major substrate by means of a dismutation reaction. The “malic” enzyme in the mitochondrion catalyzes theoxidation of malate to form pyruvate, CO2, and NADH. This reaction serves to generate intramitochondrial reducing power in the form of NADH. Concomitantly, fumarase catalyzes thedehydration of an equivalent amount of malate to form fumarate which, in turn, is reduced by an NADH-linked fumarate reductase to succinate. The flavin-linked fumarate reductase reaction results in a site I electron transport-associated phosphorylation of ADP, giving rise to ATP. This identifies a proton translocation system to obtain energy generation.
Rew RS, Saz HJ. Enzyme Localization in the Anaerobic Mitochondria Of Ascaris Lumbricoides. The Journal Of Cell Biology 1974; 63: 125-135. jcb.rupress.org
Mitochondrial redox status
Tumor cells are characterized by accelerated growth usually accompanied by up-regulated pathways that ultimately increase the rate of ATP production. These cells can suffer metabolic reprogramming, resulting in distinct bioenergetic phenotypes, generally enhancing glycolysis channeled to lactate production. These investigators showed metabolic reprogramming by means of inhibitors of histone deacetylase (HDACis), sodium butyrate and trichostatin. This treatment was able to shift energy metabolism by activating mitochondrial systems such as the respiratory chain and oxidative phosphorylation that were largely repressed in the untreated controls.
Amoêdo ND, Rodrigues MF, Pezzuto P, Galina A, et al. Energy Metabolism in H460 Lung Cancer Cells: Effects of Histone Deacetylase Inhibitors. PLoS ONE 2011; 6(7): e22264. doi:10.1371/ journal.pone.0022264
Antioxidant pathways that rely on NADPH are needed for the reduction of glutathione and maintenance of proper redox status. The mitochondrial matrix protein isocitrate dehydrogenase 2 (IDH2) is a major source of NADPH. NAD+-dependent deacetylase SIRT3 is essential for the prevention of age related hearing loss of caloric restricted mice. Oxidative stress resistance by SIRT3 was mediated through IDH2. Inserting SIRT3 Nε-acetyl-lysine into position 413 of IDH2 and has an activity loss by as much as 44-fold. Deacetylation by SIRT3 fully restored maximum IDH2 activity. The ability of SIRT3 to protect cells from oxidative stress was dependent on IDH2, and the deacetylated mimic, IDH2K413R variant was able to protect Sirt3-/- MEFs from oxidative stress through increased reduced glutathione levels. The increased SIRT3 expression protects cells from oxidative stress through IDH2 activation. Together these results uncover a previously unknown mechanism by which SIRT3 regulates IDH2 under dietary restriction. Recent findings demonstrate that IDH2 activities are a major factor in cancer, and as such, these results implicate SIRT3 as a potential regulator of IDH2-dependent functions in cancer cell metabolism.
Wei Yu, Dittenhafer-Reed KE and JM Denu. SIRT3 Deacetylates Isocitrate Dehydrogenase 2 (IDH2) and Regulates Mitochondrial Redox Status. JBC Papers in Press. Published on March 13, 2012 as Manuscript M112.355206. http://www.jbc.org
Computationally designed drug small molecules targeted for metabolic processes: a bridge from the genome to repair of dysmetabolism
New druglike small molecules with possible anticancer applications were computationally designed. The molecules formed stable complexes with antiapoptotic BCL-2, BCL-W, and BFL-1 proteins. These findings are novel because, to the best of the author’s knowledge, molecules that bind all three of these proteins are not known. A drug based on them should be more economical and better tolerated by patients than a combination of drugs, each targeting a single protein. The calculated drug-related properties of the molecules were similar to those found in most commercial drugs. The molecules were designed and evaluated following a simple, yet effective procedure. The procedure can be used efficiently in the early phases of drug discovery to evaluate promising lead compounds in time- and cost-effective ways.
Keywords: small molecule mimetics, antiapoptotic proteins, computational drug design.
Tardigrades
Tardigrades have unique stress-adaptations that allow them to survive extremes of cold, heat, radiation and vacuum. To study this, encoded protein clusters and pathways from an ongoing transcriptome study on the tardigrade Milnesium tardigradum were analyzed using bioinformatics tools and compared to expressed sequence tags (ESTs) from Hypsibius dujardini, revealing major pathways involved in resistance against extreme environmental conditions. ESTs are available on the Tardigrade Workbench along with software and databank updates. Our analysis reveals that RNA stability motifs for M. tardigradum are different from typical motifs known from higher animals. M. tardigradum and H. dujardini protein clusters and conserved domains imply metabolic storage pathways for glycogen, glycolipids and specific secondary metabolism as well as stress response pathways (including heat shock proteins, bmh2, and specific repair pathways). Redox-, DNA-, stress- and protein protection pathways complement specific repair capabilities to achieve the strong robustness of M. tardigradum. These pathways are partly conserved in other animals and their manipulation could boost stress adaptation even in human cells. However, the unique combination of resistance and repair pathways make tardigrades and M. tardigradum in particular so highly stress resistant.
Keywords: RNA, expressed sequence tag, cluster, protein family, adaptation, tardigrada, transcriptome
Epicrisis
This discussion has disparate pieces that are tied together by dysfunctional changes that are
- adaptations from metabolic process in the channeling of energy dependent of mitochondrial enzymes in interaction with three to 6 carbon carbohydrates, high energy phosphate, oxygen and membrane lipid structures, as well as
- proteins rich or poor in sulfur linked with genome specific targets, and semisynthetic modifications, oxidative stress
- leading to a new approach to pharmaceutical targeted drug design.
Related articles
- Determinants of Brain Cell Metabolic Phenotypes and Energy Substrate Utilization Unraveled with a Modeling Approach (ploscompbiol.org)
- Mitochondria and Cancer: An overview of mechanisms (pharmaceuticalintelligence.com)
- Making a molecular micromap: Imaging the yeast 26S proteasome at near-atomic resolution (phys.org)
- Fermentation (ahschoolapbio2013.wordpress.com)
- Nitric Oxide has a ubiquitous role in the regulation of glycolysis -with a concomitant influence on mitochondrial function (pharmaceuticalintelligence.com)
- What Causes Aging and How Does Dietary Restriction Increase Life Span? (understandnutrition.com)
- Diaz-Cano SJ. Tumor heterogeneity: mechanisms and bases for a reliable application of molecular marker design. Int J Mol Sci 2012; 13(2):1951-2011 PMID 22408433Bernstein LH. Expanding the Genetic Alphabet and linking the genome to the metabolome.PharmaIntell.wordpress.com. luly 24, 2012.
- Sainani K. Meet the Skeptics: Why Some Doubt Biomedical Models – and What it Takes to Win Them Over. IMAG Futures meeting. 2009
- Ritusaxena. ENCODE: the key to unlocking the secrets of complex genetic diseases. PharmaIntell.wordpress.com 2012.
- Ritusaxena. Scientists use natural agents for prostate cancer bone metastasis treatment. PharmaIntell.wordpress.com. 2012
- Ritusaxena. β Integrin emerges as an important player in mitochondrial dysfunction associated Gastric Cancer. PharmaIntell.wordpress.com 2012