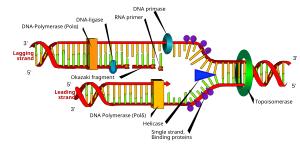
Bioinformatic Tools for Cancer Mutational Analysis: COSMIC and Beyond
Curator: Stephen J. Williams, Ph.D.
Updated 7/26/2019
Updated 04/27/2019
Signatures of Mutational Processes in Human Cancer (from COSMIC)
From The COSMIC Database
The genomic landscape of cancer. The COSMIC database has a fully curated and annotated database of recurrent genetic mutations founds in various cancers (data taken form cancer sequencing projects). For interactive map please go to the COSMIC database here: http://cancer.sanger.ac.uk/cosmic
Somatic mutations are present in all cells of the human body and occur throughout life. They are the consequence of multiple mutational processes, including the intrinsic slight infidelity of the DNA replication machinery, exogenous or endogenous mutagen exposures, enzymatic modification of DNA and defective DNA repair. Different mutational processes generate unique combinations of mutation types, termed “Mutational Signatures”.
In the past few years, large-scale analyses have revealed many mutational signatures across the spectrum of human cancer types [Nik-Zainal S. et al., Cell (2012);Alexandrov L.B. et al., Cell Reports (2013);Alexandrov L.B. et al., Nature (2013);Helleday T. et al., Nat Rev Genet (2014);Alexandrov L.B. and Stratton M.R., Curr Opin Genet Dev (2014)]. However, as the number of mutational signatures grows the need for a curated census of signatures has become apparent. Here, we deliver such a resource by providing the profiles of, and additional information about, known mutational signatures.
The current set of mutational signatures is based on an analysis of 10,952 exomes and 1,048 whole-genomes across 40 distinct types of human cancer. These analyses are based on curated data that were generated by The Cancer Genome Atlas (TCGA), the International Cancer Genome Consortium (ICGC), and a large set of freely available somatic mutations published in peer-reviewed journals. Complete details about the data sources will be provided in future releases of COSMIC.
The profile of each signature is displayed using the six substitution subtypes: C>A, C>G, C>T, T>A, T>C, and T>G (all substitutions are referred to by the pyrimidine of the mutated Watson–Crick base pair). Further, each of the substitutions is examined by incorporating information on the bases immediately 5’ and 3’ to each mutated base generating 96 possible mutation types (6 types of substitution ∗ 4 types of 5’ base ∗ 4 types of 3’ base). Mutational signatures are displayed and reported based on the observed trinucleotide frequency of the human genome, i.e., representing the relative proportions of mutations generated by each signature based on the actual trinucleotide frequencies of the reference human genome version GRCh37. Note that only validated mutational signatures have been included in the curated census of mutational signatures.
Additional information is provided for each signature, including the cancer types in which the signature has been found, proposed aetiology for the mutational processes underlying the signature, other mutational features that are associated with each signature and information that may be relevant for better understanding of a particular mutational signature.
The set of signatures will be updated in the future. This will include incorporating additional mutation types (e.g., indels, structural rearrangements, and localized hypermutation such as kataegis) and cancer samples. With more cancer genome sequences and the additional statistical power this will bring, new signatures may be found, the profiles of current signatures may be further refined, signatures may split into component signatures and signatures
See their COSMIC tutorial page here for instructional videos
Updated News: COSMIC v75 – 24th November 2015
COSMIC v75 includes curations across GRIN2A, fusion pair TCF3-PBX1, and genomic data from 17 systematic screen publications. We are also beginning a reannotation of TCGA exome datasets using Sanger’s Cancer Genome Project analyis pipeline to ensure consistency; four studies are included in this release, to be expanded across the next few releases. The Cancer Gene Census now has a dedicated curator, Dr. Zbyslaw Sondka, who will be focused on expanding the Census, enhancing the evidence underpinning it, and developing improved expert-curated detail describing each gene’s impact in cancer. Finally, as we begin to streamline our ever-growing website, we have combined all information for each gene onto one page and simplified the layout and design to improve navigation
may be found in cancer types in which they are currently not detected.
Mutational signatures across human cancer
Patterns of mutational signatures [Download signatures]
COSMIC database identifies 30 mutational signatures in human cancer
Please goto to COSMIC site to see bigger .png of mutation signatures
Signature 1
Cancer types:
Signature 1 has been found in all cancer types and in most cancer samples.
Proposed aetiology:
Signature 1 is the result of an endogenous mutational process initiated by spontaneous deamination of 5-methylcytosine.
Additional mutational features:
Signature 1 is associated with small numbers of small insertions and deletions in most tissue types.
Comments:
The number of Signature 1 mutations correlates with age of cancer diagnosis.
Signature 2
Cancer types:
Signature 2 has been found in 22 cancer types, but most commonly in cervical and bladder cancers. In most of these 22 cancer types, Signature 2 is present in at least 10% of samples.
Proposed aetiology:
Signature 2 has been attributed to activity of the AID/APOBEC family of cytidine deaminases. On the basis of similarities in the sequence context of cytosine mutations caused by APOBEC enzymes in experimental systems, a role for APOBEC1, APOBEC3A and/or APOBEC3B in human cancer appears more likely than for other members of the family.
Additional mutational features:
Transcriptional strand bias of mutations has been observed in exons, but is not present or is weaker in introns.
Comments:
Signature 2 is usually found in the same samples as Signature 13. It has been proposed that activation of AID/APOBEC cytidine deaminases is due to viral infection, retrotransposon jumping or to tissue inflammation. Currently, there is limited evidence to support these hypotheses. A germline deletion polymorphism involving APOBEC3A and APOBEC3B is associated with the presence of large numbers of Signature 2 and 13 mutations and with predisposition to breast cancer. Mutations of similar patterns to Signatures 2 and 13 are commonly found in the phenomenon of local hypermutation present in some cancers, known as kataegis, potentially implicating AID/APOBEC enzymes in this process as well.
Signature 3
Cancer types:
Signature 3 has been found in breast, ovarian, and pancreatic cancers.
Proposed aetiology:
Signature 3 is associated with failure of DNA double-strand break-repair by homologous recombination.
Additional mutational features:
Signature 3 associates strongly with elevated numbers of large (longer than 3bp) insertions and deletions with overlapping microhomology at breakpoint junctions.
Comments:
Signature 3 is strongly associated with germline and somatic BRCA1 and BRCA2 mutations in breast, pancreatic, and ovarian cancers. In pancreatic cancer, responders to platinum therapy usually exhibit Signature 3 mutations.
Signature 4
Cancer types:
Signature 4 has been found in head and neck cancer, liver cancer, lung adenocarcinoma, lung squamous carcinoma, small cell lung carcinoma, and oesophageal cancer.
Proposed aetiology:
Signature 4 is associated with smoking and its profile is similar to the mutational pattern observed in experimental systems exposed to tobacco carcinogens (e.g., benzo[a]pyrene). Signature 4 is likely due to tobacco mutagens.
Additional mutational features:
Signature 4 exhibits transcriptional strand bias for C>A mutations, compatible with the notion that damage to guanine is repaired by transcription-coupled nucleotide excision repair. Signature 4 is also associated with CC>AA dinucleotide substitutions.
Comments:
Signature 29 is found in cancers associated with tobacco chewing and appears different from Signature 4.
Signature 5
Cancer types:
Signature 5 has been found in all cancer types and most cancer samples.
Proposed aetiology:
The aetiology of Signature 5 is unknown.
Additional mutational features:
Signature 5 exhibits transcriptional strand bias for T>C substitutions at ApTpN context.
Comments:
Signature 6
Cancer types:
Signature 6 has been found in 17 cancer types and is most common in colorectal and uterine cancers. In most other cancer types, Signature 6 is found in less than 3% of examined samples.
Proposed aetiology:
Signature 6 is associated with defective DNA mismatch repair and is found in microsatellite unstable tumours.
Additional mutational features:
Signature 6 is associated with high numbers of small (shorter than 3bp) insertions and deletions at mono/polynucleotide repeats.
Comments:
Signature 6 is one of four mutational signatures associated with defective DNA mismatch repair and is often found in the same samples as Signatures 15, 20, and 26.
Signature 7
Cancer types:
Signature 7 has been found predominantly in skin cancers and in cancers of the lip categorized as head and neck or oral squamous cancers.
Proposed aetiology:
Based on its prevalence in ultraviolet exposed areas and the similarity of the mutational pattern to that observed in experimental systems exposed to ultraviolet light Signature 7 is likely due to ultraviolet light exposure.
Additional mutational features:
Signature 7 is associated with large numbers of CC>TT dinucleotide mutations at dipyrimidines. Additionally, Signature 7 exhibits a strong transcriptional strand-bias indicating that mutations occur at pyrimidines (viz., by formation of pyrimidine-pyrimidine photodimers) and these mutations are being repaired by transcription-coupled nucleotide excision repair.
Comments:
Signature 8
Cancer types:
Signature 8 has been found in breast cancer and medulloblastoma.
Proposed aetiology:
The aetiology of Signature 8 remains unknown.
Additional mutational features:
Signature 8 exhibits weak strand bias for C>A substitutions and is associated with double nucleotide substitutions, notably CC>AA.
Comments:
Signature 9
Cancer types:
Signature 9 has been found in chronic lymphocytic leukaemias and malignant B-cell lymphomas.
Proposed aetiology:
Signature 9 is characterized by a pattern of mutations that has been attributed to polymerase η, which is implicated with the activity of AID during somatic hypermutation.
Additional mutational features:
Comments:
Chronic lymphocytic leukaemias that possess immunoglobulin gene hypermutation (IGHV-mutated) have elevated numbers of mutations attributed to Signature 9 compared to those that do not have immunoglobulin gene hypermutation.
Signature 10
Cancer types:
Signature 10 has been found in six cancer types, notably colorectal and uterine cancer, usually generating huge numbers of mutations in small subsets of samples.
Proposed aetiology:
It has been proposed that the mutational process underlying this signature is altered activity of the error-prone polymerase POLE. The presence of large numbers of Signature 10 mutations is associated with recurrent POLE somatic mutations, viz., Pro286Arg and Val411Leu.
Additional mutational features:
Signature 10 exhibits strand bias for C>A mutations at TpCpT context and T>G mutations at TpTpT context.
Comments:
Signature 10 is associated with some of most mutated cancer samples. Samples exhibiting this mutational signature have been termed ultra-hypermutators.
Signature 11
Cancer types:
Signature 11 has been found in melanoma and glioblastoma.
Proposed aetiology:
Signature 11 exhibits a mutational pattern resembling that of alkylating agents. Patient histories have revealed an association between treatments with the alkylating agent temozolomide and Signature 11 mutations.
Additional mutational features:
Signature 11 exhibits a strong transcriptional strand-bias for C>T substitutions indicating that mutations occur on guanine and that these mutations are effectively repaired by transcription-coupled nucleotide excision repair.
Comments:
Signature 12
Cancer types:
Signature 12 has been found in liver cancer.
Proposed aetiology:
The aetiology of Signature 12 remains unknown.
Additional mutational features:
Signature 12 exhibits a strong transcriptional strand-bias for T>C substitutions.
Comments:
Signature 12 usually contributes a small percentage (<20%) of the mutations observed in a liver cancer sample.
Signature 13
Cancer types:
Signature 13 has been found in 22 cancer types and seems to be commonest in cervical and bladder cancers. In most of these 22 cancer types, Signature 13 is present in at least 10% of samples.
Proposed aetiology:
Signature 13 has been attributed to activity of the AID/APOBEC family of cytidine deaminases converting cytosine to uracil. On the basis of similarities in the sequence context of cytosine mutations caused by APOBEC enzymes in experimental systems, a role for APOBEC1, APOBEC3A and/or APOBEC3B in human cancer appears more likely than for other members of the family. Signature 13 causes predominantly C>G mutations. This may be due to generation of abasic sites after removal of uracil by base excision repair and replication over these abasic sites by REV1.
Additional mutational features:
Transcriptional strand bias of mutations has been observed in exons, but is not present or is weaker in introns.
Comments:
Signature 2 is usually found in the same samples as Signature 13. It has been proposed that activation of AID/APOBEC cytidine deaminases is due to viral infection, retrotransposon jumping or to tissue inflammation. Currently, there is limited evidence to support these hypotheses. A germline deletion polymorphism involving APOBEC3A and APOBEC3B is associated with the presence of large numbers of Signature 2 and 13 mutations and with predisposition to breast cancer. Mutations of similar patterns to Signatures 2 and 13 are commonly found in the phenomenon of local hypermutation present in some cancers, known as kataegis, potentially implicating AID/APOBEC enzymes in this process as well.
Signature 14
Cancer types:
Signature 14 has been observed in four uterine cancers and a single adult low-grade glioma sample.
Proposed aetiology:
The aetiology of Signature 14 remains unknown.
Additional mutational features:
Comments:
Signature 14 generates very high numbers of somatic mutations (>200 mutations per MB) in all samples in which it has been observed.
Signature 15
Cancer types:
Signature 15 has been found in several stomach cancers and a single small cell lung carcinoma.
Proposed aetiology:
Signature 15 is associated with defective DNA mismatch repair.
Additional mutational features:
Signature 15 is associated with high numbers of small (shorter than 3bp) insertions and deletions at mono/polynucleotide repeats.
Comments:
Signature 15 is one of four mutational signatures associated with defective DNA mismatch repair and is often found in the same samples as Signatures 6, 20, and 26.
Signature 16
Cancer types:
Signature 16 has been found in liver cancer.
Proposed aetiology:
The aetiology of Signature 16 remains unknown.
Additional mutational features:
Signature 16 exhibits an extremely strong transcriptional strand bias for T>C mutations at ApTpN context, with T>C mutations occurring almost exclusively on the transcribed strand.
Comments:
Signature 17
Cancer types:
Signature 17 has been found in oesophagus cancer, breast cancer, liver cancer, lung adenocarcinoma, B-cell lymphoma, stomach cancer and melanoma.
Proposed aetiology:
The aetiology of Signature 17 remains unknown.
Additional mutational features:
Comments:
Signature 1Signature 18
Cancer types:
Signature 18 has been found commonly in neuroblastoma. Additionally, Signature 18 has been also observed in breast and stomach carcinomas.
Proposed aetiology:
The aetiology of Signature 18 remains unknown.
Additional mutational features:
Comments:
Signature 19
Cancer types:
Signature 19 has been found only in pilocytic astrocytoma.
Proposed aetiology:
The aetiology of Signature 19 remains unknown.
Additional mutational features:
Comments:
Signature 20
Cancer types:
Signature 20 has been found in stomach and breast cancers.
Proposed aetiology:
Signature 20 is believed to be associated with defective DNA mismatch repair.
Additional mutational features:
Signature 20 is associated with high numbers of small (shorter than 3bp) insertions and deletions at mono/polynucleotide repeats.
Comments:
Signature 20 is one of four mutational signatures associated with defective DNA mismatch repair and is often found in the same samples as Signatures 6, 15, and 26.
Signature 21
Cancer types:
Signature 21 has been found only in stomach cancer.
Proposed aetiology:
The aetiology of Signature 21 remains unknown.
Additional mutational features:
Comments:
Signature 21 is found only in four samples all generated by the same sequencing centre. The mutational pattern of Signature 21 is somewhat similar to the one of Signature 26. Additionally, Signature 21 is found only in samples that also have Signatures 15 and 20. As such, Signature 21 is probably also related to microsatellite unstable tumours.
Signature 22
Cancer types:
Signature 22 has been found in urothelial (renal pelvis) carcinoma and liver cancers.
Proposed aetiology:
Signature 22 has been found in cancer samples with known exposures to aristolochic acid. Additionally, the pattern of mutations exhibited by the signature is consistent with the one previous observed in experimental systems exposed to aristolochic acid.
Additional mutational features:
Signature 22 exhibits a very strong transcriptional strand bias for T>A mutations indicating adenine damage that is being repaired by transcription-coupled nucleotide excision repair.
Comments:
Signature 22 has a very high mutational burden in urothelial carcinoma; however, its mutational burden is much lower in liver cancers.
Signature 23
Cancer types:
Signature 23 has been found only in a single liver cancer sample.
Proposed aetiology:
The aetiology of Signature 23 remains unknown.
Additional mutational features:
Signature 23 exhibits very strong transcriptional strand bias for C>T mutations.
Comments:
Signature 24
Cancer types:
Signature 24 has been observed in a subset of liver cancers.
Proposed aetiology:
Signature 24 has been found in cancer samples with known exposures to aflatoxin. Additionally, the pattern of mutations exhibited by the signature is consistent with that previous observed in experimental systems exposed to aflatoxin.
Additional mutational features:
Signature 24 exhibits a very strong transcriptional strand bias for C>A mutations indicating guanine damage that is being repaired by transcription-coupled nucleotide excision repair.
Comments:
Signature 25
Cancer types:
Signature 25 has been observed in Hodgkin lymphomas.
Proposed aetiology:
The aetiology of Signature 25 remains unknown.
Additional mutational features:
Signature 25 exhibits transcriptional strand bias for T>A mutations.
Comments:
This signature has only been identified in Hodgkin’s cell lines. Data is not available from primary Hodgkin lymphomas.
Signature 26
Cancer types:
Signature 26 has been found in breast cancer, cervical cancer, stomach cancer and uterine carcinoma.
Proposed aetiology:
Signature 26 is believed to be associated with defective DNA mismatch repair.
Additional mutational features:
Signature 26 is associated with high numbers of small (shorter than 3bp) insertions and deletions at mono/polynucleotide repeats.
Comments:
Signature 26 is one of four mutational signatures associated with defective DNA mismatch repair and is often found in the same samples as Signatures 6, 15 and 20.
Signature 27
Cancer types:
Signature 27 has been observed in a subset of kidney clear cell carcinomas.
Proposed aetiology:
The aetiology of Signature 27 remains unknown.
Additional mutational features:
Signature 27 exhibits very strong transcriptional strand bias for T>A mutations. Signature 27 is associated with high numbers of small (shorter than 3bp) insertions and deletions at mono/polynucleotide repeats.
Comments:
Signature 28
Cancer types:
Signature 28 has been observed in a subset of stomach cancers.
Proposed aetiology:
The aetiology of Signature 28 remains unknown.
Additional mutational features:
Comments:
Signature 29
Cancer types:
Signature 29 has been observed only in gingivo-buccal oral squamous cell carcinoma.
Proposed aetiology:
Signature 29 has been found in cancer samples from individuals with a tobacco chewing habit.
Additional mutational features:
Signature 29 exhibits transcriptional strand bias for C>A mutations indicating guanine damage that is most likely repaired by transcription-coupled nucleotide excision repair. Signature 29 is also associated with CC>AA dinucleotide substitutions.
Comments:
The Signature 29 pattern of C>A mutations due to tobacco chewing appears different from the pattern of mutations due to tobacco smoking reflected by Signature 4.
Signature 30
Cancer types:
Signature 30 has been observed in a small subset of breast cancers.
Proposed aetiology:
The aetiology of Signature 30 remains unknown.
Examples in the literature of deposits into or analysis from the COSMIC database
The Genomic Landscapes of Human Breast and Colorectal Cancers from Wood 318 (5853): 1108–1113 Science 2007
“analysis of exons representing 20,857 transcripts from 18,191 genes, we conclude that the genomic landscapes of breast and colorectal cancers are composed of a handful of commonly mutated gene “mountains” and a much larger number of gene “hills” that are mutated at low frequency. “
- found cellular pathways with multiple pathways
- analyzed a highly curated database (Metacore, GeneGo, Inc.) that includes human protein-protein interactions, signal transduction and metabolic pathways
- There were 108 pathways that were found to be preferentially mutated in breast tumors. Many of the pathways involved phosphatidylinositol 3-kinase (PI3K) signaling
- the cancer genome landscape consists of relief features (mutated genes) with heterogeneous heights (determined by CaMP scores). There are a few “mountains” representing individual CAN-genes mutated at high frequency. However, the landscapes contain a much larger number of “hills” representing the CAN-genes that are mutated at relatively low frequency. It is notable that this general genomic landscape (few gene mountains and many gene hills) is a common feature of both breast and colorectal tumors.
- developed software to analyze multiple mutations and mutation frequencies available from Harvard Bioinformatics at
http://bcb.dfci.harvard.edu/~gp/software/CancerMutationAnalysis/cma.htm
R Software for Cancer Mutation Analysis (download here)
CancerMutationAnalysis Version 1.0:
R package to reproduce the statistical analyses of the Sjoblom et al article and the associated Technical Comment. This package is build for reproducibility of the original results and not for flexibility. Future version will be more general and define classes for the data types used. Further details are available in Working Paper 126.
CancerMutationAnalysis Version 2.0:
R package to reproduce the statistical analyses of the Wood et al article. Like its predecessor, this package is still build for reproducibility of the original results and not for flexibility. Further details are available in Working Paper 126
Update 04/27/2019
Review 2018. The COSMIC Cancer Gene Census: describing genetic dysfunction across all human cancers. Z. Sondka et al. Nature Reviews. 2018.
The Catalogue of Somatic Mutations in Cancer (COSMIC) Cancer Gene Census (CGC) reevaluates the cancer genome landscape periodically and curates the findings into a database of genetic changes occurring in various tumor types. The 2018 CGC describes in detail the effect of 719 cancer driving genes. The recent expansion includes functional and mechanistic descriptions of how each gene contributes to disease etiology and in terms of the cancer hallmarks as described by Hanahan and Weinberg. These functional characteristics show the complexity of the cancer mutational landscape and genome and suggest ” multiple cancer-related functions for many genes, which are often highly tissue-dependent or tumour stage-dependent.” The 2018 CGC expands a second tier of genes, expanding the list of cancer related genes.
Criteria for curation of genes into CGC (curation process)
- choosing candidate genes are selected from published literature, conference abstracts, large cancer genome screens deposited in databases, and analysis of current COSMIC database
- COSMIC data are analyzed to determine presence of patterns of somatic mutations and frequency of such mutations in cancer
- literature review to determine the role of the gene in cancer
- Minimum evidence
– at least two publications from different groups shows increased mutation frequency in at least one type of cancer (PubMed)
– at least two publications from different groups showing experimental evidence of functional involvement in at least one hallmark of cancer in order to classify the mutant gene as oncogene, tumor suppressor, or fusion partner (like BCR-Abl)
- independent assessment by at least two postdoctoral fellows
- gene must be classified as either Tier 1 of Tier 2 CGC gene
- inclusion in database
- continued curation efforts
definitions:
Tier 1 gene: genes which have strong evidence from both mutational and functional analysis as being involved in cancer
Tier 2 gene: genes with mutational patterns typical of cancer drivers but not functionally characterized as well as genes with published mechanistic description of involvement in cancer but without proof of somatic mutations in cancer
Current Status of Tier 1 and Tier 2 genes in CGC
Tier 1 genes (574 genes): include 79 oncogenes, 140 tumor suppressor genes, 93 fusion partners
Tier 2 genes (719 genes): include 103 oncogenes, 181 tumor suppressors, 134 fusion partners and 31 with unknown function
Updated 7/26/2019
|
The COSMIC database is undergoing an extensive update and reannotation, in order to ensure standardisation and modernisation across COSMIC data. This will substantially improve the identification of unique variants that may have been described at the genome, transcript and/or protein level. The introduction of a Genomic Identifier, along with complete annotation across multiple, high quality Ensembl transcripts and improved compliance with current HGVS syntax, will enable variant matching both within COSMIC and across other bioinformatic datasets. As a result of these updates there will be significant changes in the upcoming releases as we work through this process. The first stage of this work was the introduction of improvedHGVS syntax compliance in our May release. The majority of the changes will be reflected in COSMIC v90, which will be released in late August or early September, and the remaining changes will be introduced over the next few releases. The significant changes in v90 include:
Key points for youCOSMIC variants have been annotated on all relevant Ensembl transcripts across both the GRCh37 and GRCh38 assemblies from Ensembl release 93. New genomic identifiers (e.g. COSV56056643) are used, which refers to the variant change at the genomic level rather than gene, transcript or protein level and can thus be used universally. Existing COSM IDs will continue to be supported and will now be referred to as legacy identifiers e.g. COSM476. The legacy identifiers (COSM) are still searchable. In the case of mutations without genomic coordinates, hence without a COSV identifier, COSM identifiers will continue to be used. All relevant Ensembl transcripts in COSMIC (which have been selected based on Ensembl canonical classification and on the quality of the dataset to include only GENCODE basic transcripts) will now have both accession and version numbers, so that the exact transcript is known, ensuring reproducibility. This also provides transparency and clarity as the data are updated. How these changes will be reflected in the download filesAs we are now mapping all variants on all relevant Ensembl transcripts, the number of rows in the majority of variant download files has increased significantly. In the download files, additional columns are provided including the legacy identifier (COSM) and the new genomic identifier (COSV). An internal mutation identifier is also provided to uniquely represent each mutation, on a specific transcript, on a given assembly build. The accession and version number for each transcript are included. File descriptions for each of the download files will be available from the downloads page for clarity. We have included an example of the new columns below. For example: COSMIC Complete Mutation Data (Targeted screens)
We will shortly have some sample data that can be downloaded in the new table structure, to give you real data to manipulate and integrate, this will be available on the variant updates page. How this affects youWe are aware that many of the changes we are making will affect integration into your pipelines and analytical platforms. By giving you advance notice of the changes, we hope much of this can be mitigated, and the end result of having clean, standardised data will be well worth any disruption. The variant updates page on the COSMIC website will provide a central point for this information and further technical details of the changes that we are making to COSMIC. |
|