Is the Warburg Effect the Cause or the Effect of Cancer: A 21st Century View?
Author: Larry H. Bernstein, MD, FCAP

word cloud by Danielle Smolyar
A Critical Review
What is the Warburg effect?
“Warburg Effect” describes the preference of glycolysis and lactate fermentation rather than oxidative phosphorylation for energy production in cancer cells. Mitochondrial metabolism is an important and necessary component in the functioning and maintenance of the organelle, and accumulating evidence suggests that dysfunction of mitochondrial metabolism plays a role in cancer. Progress has demonstrated the mechanisms of the mitochondrial metabolism-to-glycolysis switch in cancer development and how to target this metabolic switch.
In vertebrates, food is digested and supplied to cells mainly in the form of glucose. Glucose is broken down further to make Adenosine Triphosphate (ATP) by two pathways. One is via anaerobic metabolism occurring in the cytoplasm, also known as glycolysis. The major physiological significance of glycolysis lies in making ATP quickly, but in a minuscule amount. The breakdown process continues in the mitochondria via the Krebs’s cycle coupled with oxidative phosphorylation, which is more efficient for ATP production. Cancer cells seem to be well-adjust to glycolysis. In the 1920s, Otto Warburg first proposed that cancer cells show increased levels of glucose consumption and lactate fermentation even in the presence of ample oxygen (known as “Warburg Effect”). Based on this theory, oxidative phosphorylation switches to glycolysis which promotes the proliferation of cancer cells. Many studies have demonstrated glycolysis as the main metabolic pathway in cancer cells.
Why cancer cells prefer glycolysis, an inefficient metabolic pathway?
It is now accepted that glycolysis provides cancer cells with the most abundant extracellular nutrient, glucose, to make ample ATP metabolic intermediates, such as ribose sugars, glycerol and citrate, nonessential amino acids, and the oxidative pentose phosphate pathway, which serve as building blocks for cancer cells.
Since, cancer cells have increased rates of aerobic glycolysis, investigators argue over the function of mitochondria in cancer cells. Mitochondrion, a one of the smaller organelles, produces most of the energy in the form of ATP to supply the body. In Warburg’s theory, the function of cellular mitochondrial respiration is dampened and mitochondria are not fully functional. There are many studies backing this theory. A recent review on hypoxia nicely summarizes some current studies and speculates that the “Warburg Effect” provides a benefit to the tumor not by increasing glycolysis but by decreasing mitochondrial activity.
Glycolysis
Glycolysis is enhanced and beneficial to cancer cells. The mammalian target of rapamycin (mTOR) has been well discussed in its role to promote glycolysis; recent literature has revealed some new mechanisms of how glycolysis is promoted during skin cancer development.
On the other hand, Akt is not only involved in the regulation of mitochondrial metabolism in skin cancer but also of glycolysis. Activation of Akt has been found to phosphorylate FoxO3a, a downstream transcription factor of Akt, which promotes glycolysis by inhibiting apoptosis in melanoma. In addition, activated Akt is also associated with stabilized c-Myc and activation of mTOR, which both increase glycolysis for cancer cells.
Nevertheless, ras mutational activation prevails in skin cancer. Oncogenic ras induces glycolysis. In human squamous cell carcinoma, the c-Jun NH(2)-terminal Kinase (JNK) is activated as a mediator of ras signaling, and is essential for ras-induced glycolysis, since pharmacological inhibitors if JNK suppress glycolysis. CD147/basigin, a member of the immunoglobulin superfamily, is high expressed in melanoma and other cancers.
Glyoxalase I (GLO1) is a ubiquitous cellular defense enzyme involved in the detoxification of methylglyoxal, a cytotoxic byproduct of glycolysis. In human melanoma tissue, GLO1 is upregulated at both the mRNA and protein levels.
Knockdown of GLO1 sensitizes A375 and G361 human metastatic melanoma cells to apoptosis.
The transcription factor HIF-1 upregulates a number of genes in low oxygen conditions including glycolytic enzymes, which promotes ATP synthesis in an oxygen independent manner. Studies have demonstrated that hypoxia induces HIF-1 overexpression and its transcriptional activity increases in parallel with the progression of many tumor types. A recent study demonstrated that in malignant melanoma cells, HIF-1 is upregulated, leading to elevated expression of Pyruvate Dehydrogenase Kinase 1 (PDK1), and downregulated mitochondrial oxygen consumption.
The M2 isoform of Pyruvate Kinase (PKM2), which is required for catalyzing the final step of aerobic glycolysis, is highly expressed in cancer cells; whereas the M1 isoform (PKM1) is expressed in normal cells. Studies using the skin cell promotion model (JB6 cells) demonstrated that PKM2 is activated whereas PKM1 is inactivated upon tumor promoter treatment. Acute increases in ROS inhibited PKM2 through oxidation of Cys358 in human lung cancer cells. The levels of ROS and stage of tumor development may be pivotal for the role of PKM2.
Mitochondrial metabolism and glycolysis targeting for cancer drug delivery
In cancer cells including skin cancer cells, the metabolic shift is composed of increased glycolysis, activation of anabolic pathways including amino acid and pentose phosphate production, and increased fatty acid biosynthesis. More and more studies have converged on particular glycolytic and mitochondrial metabolic targets for cancer drug discovery.
A marker for increased glycolysis in melanoma is the elevated levels of Lactate Dehydrogenase (LDH) in the blood of patients with melanoma, which has proven to be an accurate predictor of prognosis and response to treatments. LDH converts pyruvate, the final product of glycolysis, to lactate when oxygen is absent. High concentrations of lactate, in turn, negatively regulate LDH. Therefore, targeting acid excretion may provide a feasible and effective therapeutic approach for melanoma. For instance, JugloSne, a main active component in walnut, has been used in traditional medicines. Studies have shown that Juglone causes cell membrane damage and increased LDH levels in a concentration-dependent manner in cultured melanoma cells. As one of the rate-limiting enzyme of glycolysis, 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase isozyme 3 (PFKFB3) is activated in neoplastic cells. Studies have confirmed that an inhibitor of PFKFB3, 3-(3-pyridinyl)-1-(4-pyridinyl)-2-propen-1-one (3PO), suppresses glycolysis in neoplastic cells. In melanoma cell lines, the concentrations of Fru-2, 6-BP, lactate, ATP, NAD+, and NADH are diminished by 3PO. Therefore, targeting PFKFB3 using 3PO and other PFKFB3 specific inhibitors could be effective in melanoma chemotherapy.
A new NO (nitric oxide) donating compound [(S,R)-3-phenyl-4,5-dihydro-5-isoxazole acetic acid–nitric oxide (GIT-27NO)] has been tested in treating melanoma cells. The results suggest that GIT-27/NO causes a dose-dependent reduction of mitochondrial respiration in treated A375 human melanoma cells.
At least two mitochondrial enzymes are affected by angiostatin which include malate dehydrogenase, a member of the Kreb’s cycle enzymes; and adenosine triphosphate synthase. Both are identified potential angiostatin-binding partners. Treated with angiostatin, the ATP concentrations of A2058 cells were decreased. Meanwhile, using siRNA of these two enzymes also inhibited the ATP production. PKM2 is up regulated in the early stage of skin carcinogenesis, therefore, targeting PKM2 could serve as a new approach for skin cancer prevention and therapy.
The signaling pathways critical for this glycolytic activation could serve as preventive and therapeutic targets for human skin cancer.
The Historical Challenge posed by the Warburg Hypothesis.
Impaired cellular energy metabolism is the defining characteristic of nearly all cancers regardless of cellular or tissue origin. In contrast to normal cells, which derive most of their usable energy from oxidative phosphorylation, most cancer cells become heavily dependent on substrate level phosphorylation to meet energy demands. Evidence is reviewed supporting a general hypothesis that genomic instability and essentially all hallmarks of cancer, including aerobic glycolysis (Warburg effect), can be linked to impaired mitochondrial function and energy metabolism.
In a landmark review, six essential alterations in cell physiology could underlie malignant cell growth. These six alterations were described as the hallmarks of nearly all cancers and included,
- self-sufficiency in growth signals,
- insensitivity to growth inhibitory (antigrowth) signals,
- evasion of programmed cell death (apoptosis),
- limitless replicative potential,
- sustained vascularity (angiogenesis), and
- tissue invasion and metastasis.
Genome instability, leading to increased mutability, was considered the essential enabling characteristic for manifesting the six hallmarks. The loss of genomic “caretakers” or “guardians”, involved in sensing and repairing DNA damage, was proposed to explain the increased mutability of tumor cells. The loss of these caretaker systems would allow genomic instability thus enabling pre-malignant cells to reach the six essential hallmarks of cancer.
In addition to the six recognized hallmarks of cancer, aerobic glycolysis or the Warburg effect is also a robust metabolic hallmark of most tumors. Aerobic glycolysis in cancer cells involves elevated glucose uptake with lactic acid production in the presence of oxygen. This metabolic phenotype is the basis for tumor imaging using labeled glucose analogues and has become an important diagnostic tool for cancer detection and management. Genes for glycolysis are overexpressed in the majority of cancers examined.
Although aerobic glycolysis and anaerobic glycolysis are similar in that lactic acid is produced under both situations, aerobic glycolysis can arise in tumor cells from damaged respiration whereas anaerobic glycolysis arises from the absence of oxygen. As oxygen will reduce anaerobic glycolysis and lactic acid production in most normal cells (Pasteur effect), the continued production of lactic acid in the presence of oxygen can represent an abnormal Pasteur effect. This is the situation in most tumor cells.
Warburg proposed with considerable certainty and insight that irreversible damage to respiration was the prime cause of cancer. Warburg’s biographer, Hans Krebs, mentioned that Warburg’s idea on the primary cause of cancer, i.e., the replacement of respiration by fermentation (glycolysis), was only a symptom of cancer and not the cause. While there is renewed interest in the energy metabolism of cancer cells, it is widely thought that the Warburg effect and the metabolic defects expressed in cancer cells arise primarily from genomic mutability selected during tumor progression. Emerging evidence, however, questions the genetic origin of cancer and suggests that cancer is primarily a metabolic disease.
Genomic mutability and essentially all hallmarks of cancer, including the Warburg effect, can be linked to impaired respiration and energy metabolism. In brief, damage to cellular respiration precedes and underlies the genome instability that accompanies tumor development. Once established, genome instability contributes to further respiratory impairment, genome mutability, and tumor progression. In other words, effects become causes. This hypothesis is based on evidence that nuclear genome integrity is largely dependent on mitochondrial energy homeostasis and that all cells require a constant level of useable energy to maintain viability. While Warburg recognized the centrality of impaired respiration in the origin of cancer, he did not link this phenomenon to what are now recognize as the hallmarks of cancer.
Abnormal metabolism of tumors, a selective advantage
The initial observation of Warburg 1916 on tumor glycolysis with lactate production is still a crucial observation. Two fundamental findings complete the metabolic picture:
- the discovery of the M2 pyruvate kinase (PK) typical of tumors
- and the implication of tyrosine kinase signals and subsequent phosphorylations in the M2 PK blockade.
A typical feature of tumor cells is a glycolysis associated to an inhibition of apoptosis. Tumors overexpress the high affinity hexokinase 2, which strongly interacts with the mitochondrial ANT-VDAC-PTP complex. In this position, close to the ATP/ADP exchanger (ANT), the hexokinase receives efficiently its ATP substrate. As long as hexokinase occupies this mitochondria site, glycolysis is efficient. However, this has another consequence, hexokinase pushes away from the mitochondria site the permeability transition pore (PTP), which inhibits the release of cytochrome C, the apoptotic trigger. The site also contains a voltage dependent anion channel (VDAC) and other proteins. The repulsion of PTP by hexokinase would reduce the pore size and the release of cytochrome C. Thus, the apoptosome-caspase proteolytic structure does not assemble in the cytoplasm. The liver hexokinase or glucokinase, is different it has less interaction with the site, has a lower affinity for glucose; because of this difference, glucose goes preferentially to the brain.
Further, phosphofructokinase gives fructose 1-6 bisphosphate; glycolysis is stimulated if an allosteric analogue, fructose 2-6 bis phosphate increases in response to a decrease of cAMP. The activation of insulin receptors in tumors has multiple effects, among them; a decrease of cAMP, which will stimulate glycolysis. Another control point is glyceraldehyde P dehydrogenase that requires NAD+ in the glycolytic direction. If the oxygen supply is normal, the mitochondria malate/aspartate (MAL/ASP) shuttle forms the required NAD+ in the cytosol and NADH in the mitochondria. In hypoxic conditions, the NAD+ will essentially come via lactate dehydrogenase converting pyruvate into lactate. This reaction is prominent in tumor cells; it is the first discovery of Warburg on cancer.
At the last step of glycolysis, pyruvate kinase (PK) converts phospho-enolpyruvate (PEP) into pyruvate, which enters in the mitochondria as acetyl- CoA, starting the citric acid cycle and oxidative metabolism. To explain the PK situation in tumors we must recall that PK only works in the glycolytic direction, from PEP to pyruvate, which implies that gluconeogenesis uses other enzymes for converting pyruvate into PEP. In starvation, when cells need glucose, one switches from glycolysis to gluconeogenesis and ketogenesis; PK and pyruvate dehydrogenase (PDH) are off, in a phosphorylated form, presumably following a cAMP-glucagon-adrenergic signal. In parallel, pyruvate carboxylase (Pcarb) becomes active.
Moreover, in starvation, much alanine comes from muscle protein proteolysis, and is transaminated into pyruvate. Pyruvate carboxylase first converts pyruvate to OAA and then, PEP carboxykinase converts OAA to PEP etc…, until glucose. The inhibition of PK is necessary, if not one would go back to pyruvate. Phosphorylation of PK, and alanine, inhibit the enzyme.
PK and a PDH of tumors are inhibited by phosphorylation and alanine, like for gluconeogenesis, in spite of an increased glycolysis! Moreover, in tumors, one finds a particular PK, the M2 embryonic enzyme [2,9,10] the dimeric, phosphorylated form is inactive, leading to a “bottleneck “. The M2 PK has to be activated by fructose 1-6 bis P its allosteric activator, whereas the M1 adult enzyme is a constitutive active form. The M2 PK bottleneck between glycolysis and the citric acid cycle is a typical feature of tumor cell glycolysis.
Above the bottleneck, the massive entry of glucose accumulates PEP, which converts to OAA via mitochondria PEP carboxykinase, an enzyme requiring biotine-CO2-GDP. This source of OAA is abnormal, since Pcarb, another biotin-requiring enzyme, should have provided OAA. Tumors may indeed contain “morule inclusions” of biotin-enzyme suggesting an inhibition of Pcarb, presumably a consequence of the maintained citrate synthase activity, and decrease of ketone bodies that normally stimulate Pcarb. The OAA coming via PEP carboxykinase and OAA coming from aspartate transamination or via malate dehydrogenase condenses with acetyl CoA, feeding the elevated tumoral citric acid condensation starting the Krebs cycle.
Thus, tumors have to find large amounts of acetyl CoA for their condensation reaction; it comes essentially from lipolysis and β oxidation of fatty acids, and enters in the mitochondria via the carnitine transporter. This is the major source of acetyl CoA. It is as if the mechanism switching from gluconeogenesis to glycolysis was jammed in tumors, PK and PDH are at rest, like for gluconeogenesis, but citrate synthase is on. Thus, citric acid condensation pulls the glucose flux in the glycolytic direction, which needs NAD+; it will come from the pyruvate to lactate conversion by lactate dehydrogenase (LDH) no longer in competition with a quiescent Pcarb.
Since the citrate condensation consumes acetyl CoA, ketone bodies do not form; while citrate will support the synthesis of triglycerides via ATP citrate lyase and fatty acid synthesis… The cytosolic OAA drives the transaminases in a direction consuming amino acid. The result of these metabolic changes is that tumors burn glucose while consuming muscle protein and lipid stores of the organism. In a normal physiological situation, one mobilizes stores for making glucose or ketone bodies, but not while burning glucose!
The 21st Century Genomic Challenge?
According to the modern understanding of cancer, it is a disease caused by genetic and epigenetic alterations. Although this is now widely accepted, perhaps more emphasis has been given to the fact that cancer is a genetic disease. Numerous studies, including our earlier works, have supported the notion that carcinogenesis involves the activation of tumor-promoting oncogenes and the inactivation of growth-inhibiting tumor suppressor genes. It should be noted that in the post-genome sequencing project period of the 21st century, an in depth investigation of the factors associated with tumorigenesis is required for achieving it. Extensive research is warranted in two areas, namely, tumor bioenergetics and the cancer stem cell (CSC) hypothesis, neither of which received the required attention after the success of the genome sequencing project. An investigation of these two concepts would give rise to a new era in the study of cancer biology. Indeed, recent studies have indicated that the two apparently distinct fields might be related to each other and can converge more rapidly than previously recognized.
Warburg Effect Revisited
Cancer cells rarely depend on mitochondria for respiration and obtain almost half of their ATP by directly metabolizing glucose to lactic acid, even in the presence of oxygen. However, with the discovery that tumors do not show any shift to glycolysis, Warburg’s cancer theory (high lactate production and low mitochondrial respiration in tumor under normal oxygen pressure) was gradually discredited. Otto Warburg won a Nobel Prize in 1931 for the discovery of tumor bioenergetics, which is now commonly used as the basis of positron emission tomography (PET), a highly sensitive noninvasive technique used in cancer diagnosis. The increasing number of recent reports on the Warburg effect has reestablished the significance of this effect in tumorigenesis, indicating that bioenergetics may play a critical role in malignant transformation. Furthermore, it has been reported that TP53, which is one of the most commonly mutated genes in cancer, can trigger the Warburg effect. Glycolytic conversion is initiated in the early stages in cells that are genetically engineered to become cancerous, and the conversion was enhanced as the cells became more malignant. Therefore, the Warburg effect might directly contribute to the initiation of cancer formation not only by enhanced glycolysis but also via decreased respiration in the presence of oxygen, which suppresses apoptosis. This effect may also produce a metabolic shift to enhanced glycolysis and play a role in the early stages of multistep tumorigenesis in vivo.
Cancer Stem Cells (CSC) and Embryonic Stem Cells (ESC)
The importance of the cancer stem cell (CSC) hypothesis in therapy-related resistance and metastasis has been recognized during the past 2 decades. Accumulating evidence suggests that tumor bioenergetics plays a critical role in CSC regulation; this finding has opened up a new era of cancer medicine, which goes beyond cancer genomics.
Embryonic stem (ES) cells and immortalized primary and cancerous cells show a common concerted metabolic shift, including:
- enhanced glycolysis,
- decreased apoptosis, and
- reduced mitochondrial respiration.
This finding reinforces the use of somatic stem cells or metastatic tumor cells in hypoxic niches. Hypoxia appears to regulate the functions of hematopoietic stem cells in the bone marrow and metastatic tumor cells by preserving important stem cell functions, such as:
- cell cycle control,
- survival,
- metabolism, and
- protection against oxidative stress.
Several companies and laboratories are now attempting to evaluate the bioenergetics associated with tumorigenesis by testing and challenging the available anticancer drugs.
A small population of cancer-initiating cells plays a very important role in current investigations. These CSCs may cause resistance to chemotherapy or radiation therapy or lead to post-therapy recurrence even when most of the cancer cells appear to be dead. In addition to their genetic alterations, CSCs are believed to mimic normal adult stem cells with regard to properties like self-renewal and undifferentiated status, which eventually leads to the formation of differentiated cells. Unlike well-differentiated daughter cells, small populations of CSCs are believed to be more resistant to toxic injuries and chemoradiotherapy. Indeed, the conventional cancer therapies have always been targeted toward proliferating cells. The control of CSCs, which is often exercised in the dormant phase of the cell cycle, can now be applied to achieve complete tumor regression.
Identification of cancer-specific markers
Due to their potential use in clinical applications, the surface markers of CSCs have been studied and identified. Adult stem cells and their malignant counterparts share similar intrinsic and extrinsic factors that regulate the
- self renewal,
- differentiation, and
- proliferation pathways.
The following are the examples of candidate markers: musashi-1 (Msi-1), hairy and enhancer of split homolog-1 (Hes-1), CD133 (prominin-1, Prom1), epithelial cellular adhesion molecule (EpCam), claudin-7,29 CD44 variant isoforms, Lgr5,30Hedgehog (Hh), bone morphogenic protein (Bmp), Notch, and Wnt.
Is cancer a metabolic disease and genomic instability a secondary effect?
Bioenergetics of Cancer Stem Cells
The bioenergetics associated with the adaptation of CSCs to their micro-environment still requires extensive research. Although numerous studied suggested the association between Warburg effect and reduced oxidative stress in cancer, the relevant molecular mechanism was not known until very recently when Ruckenstuhl, et al. reported their findings in a yeast model.
How cancer cells achieve one of the most common phenotypes, namely, the “Warburg effect,” i.e., elevated glycolysis in the presence of oxygen, is still a topic of hypothesis, unless the involvement of glycolysis genes is considered.
The Warburg effect has been observed in differentiating cancer cells (e.g., cells that undergo epithelial-to-mesenchymal and mesenchymal-to-amoeboid transition), cells resistant to anoikis, and cells which interact with the stromal components of the metastatic niche. The epithelial-to-mesenchymal transition is involved in the resistance to chemotherapy in gastrointestinal cancer cells.
Cancer metastasis can be regarded as an integrated “escape program” triggered by redox changes. These alterations might be associated with avoiding oxidative stress in the niche of the tumor cells, or presumably with the response to treatments aimed at genetic targets, such as chemotherapy and radiation.
The introduction of induced pluripotent stem (iPS) cell genes was necessary for inducing the expression of immature status-related proteins in gastrointestinal cancer cells, and that the induced pluripotent cancer (iPC) cells were distinct from natural cancer cells with regard to their sensitivity to differentiation inducing treatment. For the complete eradication of cancer, however, future efforts should be directed toward improving translational research.
Cancer metabolism.
Glycolysis is elevated in tumors, but a pyruvate kinase (PK) “bottleneck” interrupts phosphoenol pyruvate (PEP) to pyruvate conversion. Thus, alanine following muscle proteolysis transaminates to pyruvate, feeding lactate dehydrogenase, converting pyruvate to lactate, (Warburg effect) and NAD+ required for glycolysis. Cytosolic malate dehydrogenase also provides NAD+ (in OAA to MAL direction). Malate moves through the shuttle giving back OAA in the mitochondria. Below the PK-bottleneck, pyruvate dehydrogenase (PDH) is phosphorylated (second bottleneck). However, citrate condensation increases: acetyl-CoA, will thus come from fatty acids b-oxydation and lipolysis, while OAA sources are via PEP carboxy kinase, and malate dehydrogenase, (pyruvate carboxylase is inactive). Citrate quits the mitochondria, (note interrupted Krebs cycle). In the cytosol, ATP citrate lyase cleaves citrate into acetyl CoA and OAA.
Acetyl CoA will make fatty acids-triglycerides. Above all, OAA pushes transaminases in a direction usually associated to gluconeogenesis! This consumes protein stores, providing alanine (ALA); like glutamine, it is essential for tumors. The transaminases output is aspartate (ASP) it joins with ASP from the shuttle and feeds ASP transcarbamylase, starting pyrimidine synthesis. ASP in not processed by argininosuccinate synthetase, which is blocked, interrupting the urea cycle.
Arginine gives ornithine via arginase, ornithine is decarboxylated into putrescine by ornithine decarboxylase. Putrescine and SAM form polyamines (spermine spermidine) via SAM decarboxylase. The other product 5-methylthioadenosine provides adenine. Arginine deprivation should affect tumors. The SAM destruction impairs methylations, particularly of PP2A, removing the “signaling kinase brake”, PP2A also fails to dephosphorylate PK and PDH, forming the “bottlenecks”.
Insulin or IGF actions boost the cellular influx of glucose and glycolysis. However, if the signaling pathway gets out of control, the tyrosine kinase phosphorylations may lead to a parallel PK blockade explaining the tumor bottleneck at the end of glycolysis. Since an activation of enyme kinases may indeed block essential enzymes (PK, PDH and others); in principle, the inactivation of phosphatases may also keep these enzymes in a phosphorylated form and lead to a similar bottleneck and we do know that oncogenes bind and affect PP2A phosphatase. In sum, a perturbed MAP kinase pathway, elicits metabolic features that would give to tumor cells their metabolic advantage.
Warburg effect and the prognostic value of stromal caveolin-1 as a marker of a lethal tumor microenvironment
Cancer cells show a broad spectrum of bioenergetic states, with some cells using aerobic glycolysis while others rely on oxidative phosphorylation as their main source of energy. In addition, there is mounting evidence that metabolic coupling occurs in aggressive tumors, between epithelial cancer cells and the stromal compartment, and between well-oxygenated and hypoxic compartments. We recently showed that oxidative stress in the tumor stroma, due to aerobic glycolysis and mitochondrial dysfunction, is important for cancer cell mutagenesis and tumor progression. More specifically, increased autophagy/mitophagy in the tumor stroma drives a form of parasitic epithelial-stromal metabolic coupling. These findings explain why it is effective to treat tumors with either inducers or inhibitors of autophagy, as both would disrupt this energetic coupling. We also discuss evidence that glutamine addiction in cancer cells produces ammonia via oxidative mitochondrial metabolism.
Ammonia production in cancer cells, in turn, could then help maintain autophagy in the tumor stromal compartment. In this vicious cycle, the initial glutamine provided to cancer cells would be produced by autophagy in the tumor stroma. Thus, we believe that parasitic epithelial-stromal metabolic coupling has important implications for cancer diagnosis and therapy, for example, in designing novel metabolic imaging techniques and establishing new targeted therapies. In direct support of this notion, we identified a loss of stromal caveolin-1 as a marker of oxidative stress, hypoxia, and autophagy in the tumor microenvironment, explaining its powerful predictive value. Loss of stromal caveolin-1 in breast cancers is associated with early tumor recurrence, metastasis, and drug resistance, leading to poor clinical outcome.
The conventional ‘Warburg effect’ versus oxidative mitochondrial metabolism
Warburg’s original work indicated that while glucose uptake and lactate production are greatly elevated, a cancer cell’s rate of mitochondrial respiration is similar to that of normal cells. He, however, described it as a ‘respiratory impairment’ due to the fact that, in cancer cells, mitochondrial respiration is smaller, relative to their glycolytic power, but not smaller relative to normal cells. He recognized that oxygen consumption is not diminished in tumor cells, but that respiration is disturbed because glycolysis persists in the presence of oxygen. Unfortunately, the perception of his original findings was simplified over the years, and most subsequent papers validated that cancer cells undergo aerobic glycolysis and produce lactate, but did not measure mitochondrial respiration, and just presumed decreased tricarboxylic acid (TCA) cycle activity and reduced oxidative phosphorylation [1,2]. It is indeed well documented that, as a consequence of intra-tumoral hypoxia, the hypoxia-inducible factor (HIF)1α pathway is activated in many tumors cells, resulting in the direct up-regulation of lactate dehydrogenase (LDH) and increased glucose consumption.
It is now clear that cancer cells utilize both glycolysis and oxidative phosphorylation to satisfy their metabolic needs. Experimental assessments of ATP production in cancer cells have demonstrated that oxidative pathways play a signifi cant role in energy generation, and may be responsible for about 50 to 80% of the ATP generated. several studies now clearly indicate that mitochondrial activity and oxidative phosphorylation support tumor growth. Loss-of-function mutations in the TCA cycle gene IDH1 (isocitrate dehydrogenase 1) are found in about 70% of gliomas, but, interestingly, correlate with a better prognosis and improved survival, suggesting that severely decreased activity in one of the TCA cycle enzymes does not favor tumor aggressiveness. The mitochondrial protein p32 was shown to maintain high levels of oxidative phosphorylation in human cancer cells and to sustain tumorigenicity in vivo. In addition, STAT3 is known to enhance tumor growth and to predict poor prognosis in human cancers. Interestingly, a pool of STAT3 localizes to the mitochondria, to sustain high levels of mitochondrial respiration and to augment transformation by oncogenic Ras. Similarly, the mitochondrial transcription factor A (TFAM), which is required for mitochondrial DNA replication and oxidative phosphorylation, is also required for K-Ras induced lung tumorigenesis.
There is also evidence that pro-oncogenic molecules regulate mitochondrial function. Cyclin D1 inhibits mitochondrial function in breast cancer cells. Overexpression of cyclin D1 is observed in about 50% of invasive breast cancers and is associated with a good clinical outcome, indicating that inhibition of mitochondrial activity correlates with favorable prognosis. Importantly, it was shown that the oncogene c-Myc stimulates mitochondrial biogenesis, and enhances glutamine metabolism by regulating the expression of mitochondrial glutaminase, the first enzyme in the glutamine utilization pathway. Glutamine is an essential metabolic fuel that is converted to alpha-ketoglutarate and serves as a substrate for the TCA cycle or for glutathione synthesis, to promote energy production and cellular biosynthesis, and to protect against oxidative stress. Interestingly, pharmacological targeting of mitochondrial glutaminase inhibits cancer cell transforming activity, suggesting that glutamine metabolism and its role in fueling and replenishing the TCA cycle are required for neoplastic transformation.
Reverse Warburg Effect.
It is increasingly apparent that the tumor microenvironment regulates neoplastic growth and progression. Activation of the stroma is a critical step required for tumor formation. Among the stromal players, cancer associated fi broblasts (CAFs) have recently taken center stage [25]. CAFs are activated, contractile fibroblasts that display features of myo-fibroblasts, express muscle specific actin, and show an increased ability to secrete and remodel the extracellular matrix. They are not just neutral spectators, but actively support malignant transformation and metastasis, as compared to normal resting fibroblasts.
Importantly, the tumor stroma dictates clinical outcome and constitutes a source of potential biomarkers. Expression profiling has identified a cancer-associated stromal signature that predicts good and poor clinical prognosis in breast cancer patients, independently of other factors.
A loss of caveolin-1 (Cav-1) in the stromal compartment is a novel biomarker for predicting poor clinical outcome in all of the most common subtypes of human breast cancer, including the more lethal triple negative subtype. A loss of stromal Cav-1 predicts early tumor recurrence, lymph node metastasis, tamoxifen-resistance, and poor survival.
Overall, breast cancer patients with a loss of stromal Cav-1show a 20% 5-year survival rate, compared to the 80% 5-year survival of patients with high stromal Cav-1 expression. In triple negative patients, the 5-year survival rate is 75.5% for high stromal Cav-1 versus 9.4% for absent stromal Cav-1. A loss of stromal Cav-1 also predicts progression to invasive disease in ductal carcinoma in situ patients, suggesting that a loss of Cav-1 regulates tumor progression. Similarly, a loss of stromal Cav-1 is associated with advanced disease and metastasis, as well as a high Gleason score, in prostate cancer patients.
The autophagic tumor stroma model of cancer metabolism.
Cancer cells induce oxidative stress in adjacent cancer-associated fibroblasts (CAFs). This activates reactive oxygen species (ROS) production and autophagy. ROS production in CAFs, via the bystander eff ect, serves to induce random mutagenesis in epithelial cancer cells, leading to double-strand DNA breaks and aneuploidy. Cancer cells mount an anti-oxidant defense and upregulate molecules that protect them against ROS and autophagy, preventing them from undergoing apoptosis. So, stromal fibroblasts conveniently feed and mutagenize cancer cells, while protecting them against death. See the text for more details. A+, autophagy positive; A-, autophagy negative; AR, autophagy resistant.
1. Recycled Nutrients
2. Random Mutagenesis
3. Protection Against Apoptosis
The clinical use of PET is well established in Hodgkin’s lymphomas which are composed of less than 10% tumor cells, the rest being stromal and inflammatory cells. Yet, Hodgkin’s lymphomas are very PET avid tumors, suggesting that 2-deoxy-glucose uptake may be associated with the tumor stroma. That the fibrotic component may be glucose avid is further supported by the notion that PET is clinically used to assess the therapeutic response in gastrointestinal stromal tumors (GIST), which are a subset of tumors of mesenchymal origin.
The reverse Warburg effect can be described as ‘metabolic coupling’ between supporting glycolytic stromal cells and oxidative tumor cells. Metabolic cooperativity between adjacent cell-compartments is observed in several normal physiological settings.
The reverse Warburg effect.
Via oxidative stress, cancer cells activate two major transcription factors in adjacent stromal fibroblasts (hypoxia-inducible factor (HIF)1α and NFκB).
This leads to the onset of both autophagy and mitophagy, as well as aerobic glycolysis, which then produces recycled nutrients (such as lactate, ketones, and glutamine).
These high-energy chemical building blocks can then be transferred and used as fuel in the tricarboxylic acid cycle (TCA) in adjacent cancer cells.
The outcome is high ATP production in cancer cells, and protection against cell death. ROS, reactive oxygen species.
The methylation hypothesis and the role of PP2A phosphatase
Diethanolamine decreased choline derivatives and methyl donors in the liver, like seen in a choline deficient diet. Such conditions trigger tumors in mice, particularly in the B6C3F1 strain. Again, the historical perspective recalled by Newberne’s comment brings us back to insulin. Indeed, after the discovery of insulin in 1922, Banting and Best were able to keep alive for several months depancreatized dogs, treated with pure insulin. However, these dogs developed a fatty liver and died. Unlike pure insulin, the total pancreatic extract contained a substance that prevented fatty liver: a lipotropic substance identified later as being choline. Like other lipotropes, (methionine, folate, B12) choline supports transmethylation reactions, of a variety of substrates, that would change their cellular fate, or action, after methylation. In the particular case concerned here, the removal of triglycerides from the liver, as very low-density lipoprotein particles (VLDL), requires the synthesis of lecithin, which might decrease if choline and S-adenosyl methionine (SAM) are missing. Hence, a choline deficient diet decreases the removal of triglycerides from the liver; a fatty liver and tumors may then form. In sum, we have seen that pathways exemplified by the insulin-tyrosine kinase signaling pathway, which control anabolic processes, mitosis, growth and cell death, are at each step targets for oncogenes; we now find that insulin may also provoke fatty liver and cancer, when choline is not associated to insulin.
We know that after the tyrosine kinase reaction, serine-threonine kinases take over along the signaling route. It is thus highly probable that serine-threonine phosphatases will counteract the kinases and limit the intensity of the insulin or insulin like signals. One of the phosphatases involved is PP2A, itself the target of DNA viral oncogenes (Polyoma or SV40 antigens react with PP2A subunits and cause tumors). We found a possible link between the PP2A phosphatase brake and choline. the catalytic C subunit of PP2A is associated to a structural subunit A. When C receives a methyl, the dimer recruits a regulatory subunit B. The trimer then targets specific proteins that are dephosphorylated. choline, via SAM, methylates PP2A, which is targeted toward the serine-threonine kinases that are counteracted along the insulin-signaling pathway.
The choline dependent methylation of PP2A is the brake, the “antidote”, which limits “the poison” resulting from an excess of insulin signaling. Moreover, it seems that choline deficiency is involved in the L to M2 transition of PK isoenzymes. The negative regulation of Ras/MAP kinase signals mediated by PP2A phosphatase seems to be complex. The serine-threonine phosphatase does more than simply counteracting kinases; it binds to the intermediate Shc protein on the signaling cascade, which is inhibited. The targeting of PP2A towards proteins of the signaling pathway depends of the assembly of the different holoenzymes.
The relative decrease of methylated PP2A in the cytosol, not only cancels the brake over the signaling kinases, but also favors the inactivation of PK and PDH, which remain phosphorylated, contributing to the metabolic anomaly of tumor cells. In order to prevent tumors, one should then favor the methylation route rather than the phosphorylation route for choline metabolism. This would decrease triglycerides, promote the methylation of PP2A and keep it in the cytosol, reestablishing the brake over signaling kinases. Moreover, PK, and PDH would become active after the phosphatase action. One would also gain to inhibit their kinases as recently done with dichloroacetate for PDH kinase. The nuclear or cytosolic targeting of PP2A isoforms is a hypothesis also inspired by several works.
Hypoxic adaptations in the presence of oxygen
Through different biochemical and biophysical pathways, which are characteristic to cancer cells, tumor cells adopt this phenotype, i.e., high glycolysis and decreased respiration, in the presence of oxygen. It has been shown that although the induction of hypoxia and cellular proliferation engage entirely different cellular pathways, they often coexist during tumor growth. The ability of cells to grow during hypoxia results, in part, from the crosstalk between hypoxia-inducible factors (Hifs) and the proto-oncogene c-Myc. These genes partially regulate the development of complex adaptations of tumor cells growing in low O2, and contribute to fine tuning the adaptive responses of cells to hypoxic environments.
Hypoxic conditions seem to trigger back the expression of the fetal gene packet via HIF1-Von-Hippel signals. The mechanism would depend of a double switch since not all fetal genes become active after hypoxia. First, the histones have to be in an acetylated form, opening the way to transcription factors, this depends either of histone eacetylase (HDAC) inhibition or of histone acetyltransferase (HAT) activation, and represents the main switch
Growth hormone-IGF actions, the control of asymmetrical mitosis
When IGF – Growth hormone operate, the fatty acid source of acetyl CoA takes over. Indeed, GH stimulates a triglyceride lipase in adipocytes, increasing the release of fatty acids and their b oxidation. In parallel, GH would close the glycolytic source of acetyl CoA, perhaps inhibiting the hexokinase interaction with the mitochondrial ANT site. This effect, which renders apoptosis possible, does not occur in tumor cells. GH mobilizes the fatty acid source of acetyl CoA from adipocytes, which should help the formation of ketone bodies. Since citrate synthase activity is elevated in tumors, ketone bodies do not form. This result silences several genes like PETEN, P53, or methylase inhibitory genes. It is probable that the IGFBP gene gets silent as well.
Uncoupling Proteins in Cancer
Uncoupling proteins (UCPs) are a family of inner mitochondrial membrane proteins whose function is to allow the re-entry of protons to the mitochondrial matrix, by dissipating the proton gradient and, subsequently, decreasing membrane potential and production of reactive oxygen species (ROS). Due to their pivotal role in the intersection between energy efficiency and oxidative stress, UCPs are being investigated for a potential role in cancer.
Mitochondria have been shown to be key players in numerous cellular events tightly related with the biology of cancer. Although energy production relies on the glycolytic pathway in cancer cells, these organelles also participate in many other processes essential for cell survival and proliferation such as ROS production, apoptotic and necrotic cell death, modulation of oxygen concentration, calcium and iron homeostasis, and certain metabolic and biosynthetic pathways. Many of these mitochondrial-dependent processes are altered in cancer cells, leading to a phenotype characterized, among others, by higher oxidative stress, inhibition of apoptosis, enhanced cell proliferation, chemoresistance, induction of angiogenic genes and aggressive fatty acid oxidation. Uncoupling proteins, a family of inner mitochondrial membrane proteins specialized in energy-dissipation, has aroused enormous interest in cancer due to their relevant impact on such processes and their potential for the development of novel therapeutic strategies.
Briefly, oxidation of reduced nutrient molecules, such as carbohydrates, lipids, and proteins, through cellular metabolism yields electrons in the form of reduced hydrogen carriers NADH+ and FADH2. These reduced cofactors donate electrons to a series of protein complexes embedded in the inner mitochondrial membrane known as the electron transport chain (ETC). These complexes use the energy released from electron transport for active pumping of protons across the inner membrane, generating an electrochemical gradient. Mitochondria orchestrate conversions between different forms of energy, coupling aerobic respiration to phosphorylation.
Conversion of metabolic fuel into ATP is not a fully efficient process. Some of the energy of the electrochemical gradient is not coupled to ATP production due to a phenomenon known as proton leak, which consists of the return of protons to the mitochondrial matrix through alternative pathways that bypass ATP synthase. Although this apparently futile cycle of protons is physiologically important, accounting for 20-25% of basal metabolic rate, its function is still a subject of debate. Several different functions have been suggested for proton leak, including thermogenesis, regulation of energy metabolism, and control of body weight and attenuation of reactive oxygen species (ROS) production. Although a part of the proton leak may be attributed to biophysical properties of the inner membrane, such as protein/lipid interfaces, the bulk of the proton conductance is linked to the action of a family of mitochondrial proteins termed uncoupling proteins.
Mitochondria are the major sources of reactive oxygen species (ROS). Aerobic respiration involves the complete reduction of oxygen to water, which is catalysed by complex IV (or cytochrome c oxidase). Nevertheless, during the transfer of electrons along the electron transport complexes, single electrons sometimes escape and result in a single electron reduction of molecular oxygen to form a superoxide anion, which, in turn is the precursor of other ROS.
One of the most interesting functions attributed to UCPs is their ability to decrease the formation of mitochondrial ROS. Mitochondria are the main source of ROS in cells. Superoxide formation is strongly activated under resting (state 4) conditions when the membrane potential is high and the rate of electron transport is limited by lack of ADP and Pi. Thus, there is a well established strong positive correlation between membrane potential and ROS production.
A small increase in membrane potential gives rise to a large stimulation of ROS production, whereas a small decrease in membrane potential (10 mV) is able to inhibit ROS production by 70% . Therefore, mild uncoupling, i.e., a small decrease in membrane potential, has been suggested to have a natural antioxidant effect.
Consistent with such a proposal, the inhibition of UCPs by GDP in mitochondria has been shown to increase membrane potential and mitochondrial ROS production. The loss of UCP2 or UCP3 in knockouts yielded increased ROS production concurrent with elevated membrane potential specifically in those tissues normally expressing the missing protein.
The hypothesis of UCPs as an antioxidant defense has been strongly supported by the fact that these proteins have been shown to be activated by ROS or by-products of lipid peroxidation, showing that UCPs would form part of a negative feed-back mechanism aimed to mitigate excessive ROS production and oxidative damage.
ROS and Cancer
ROS are thought to play multiple roles in tumor initiation, progression and maintenance, eliciting cellular responses that range from proliferation to cell death. In normal cells, ROS play crucial roles in several biological mechanisms including phagocytosis, proliferation, apoptosis, detoxification and other biochemical reactions. Low levels of ROS regulate cellular signaling and play an important role in normal cell proliferation. During initiation of cancer, ROS may cause DNA damage and mutagenesis, while ROS acting as second messengers stimulate proliferation and inhibit apoptosis, conferring growth advantage to established cancer cells. Cancer cells have been found to have increased ROS levels.
One of the functional roles of these elevated ROS levels during tumor progression is constant activation of transcription factors such as NF-kappaB and AP-1 which induce genes that promote proliferation and inhibit apoptosis. In addition, oxidative stress can induce DNA damage which leads to genomic instability and the acquisition of new mutations, which may contribute to cancer progression.
Role of ROS in control of proliferation and apoptosis
ROS are also essential mediators of apoptosis which eliminates cancer and other cells that threaten our health [81–86]. Many chemotherapeutic drugs and radiotherapy are aimed at increasing ROS levels to promote apoptosis by stimulating pro-apoptotic singaling molecules such as ASK1, JNK and p38. Because of the pivotal role of ROS in triggering apoptosis, antioxidants can inhibit this protective mechanism by depleting ROS. Thus, antioxidant mechanisms are thought to interfere with the therapeutic activity of anticancer drugs that kill advanced stage cancer cells by apoptosis.
Effect of uncoupling proteins on proliferation and apoptosis in relation to ROS levels
Uncoupling-to-survive hypothesis (proposed by Brand)
- the ability of UCP2 to increase lifespan is mediated by decreased ROS production and oxidative stress.
- the ability of mild uncoupling to avoid ROS formation, gives a reasonable argument to hypothesize about a role for UCPs in cancer prevention
Consistently, Derdák et al. showed that Ucp2−/− mice treated with the carcinogen azoxymethane were found to develop more aberrant crypt foci and colon tumours than Ucp2+/+ in relation with increased oxidative stress and enhanced NF-kappaB activation.
Roles of UCPs in Cancer Progression
The growth of a tumor from a single genetically altered cell is a stepwise progression requiring the alterations of several genes which contribute to the acquisition of a malignant phenotype. Such genetic alterations are positively selected when in the tumor, they confer a proliferative, survival or treatment resistance advantage for the host cell. In addition, several mutations, such as those silencing tumour suppressor genes, trigger the probability of accumulating new mutations, so the process of malignant transformation is progressively self-accelerated.
Considering the ability of UCPs to modulate mutagenic ROS, as well as mitochondrial bioenergetics and membrane potential, both involved in regulation of cell survival, an interesting question is whether UCPs can be involved in the progression of cancer.
Increased uncoupled respiration may be a mechanism to lower cellular oxygen concentration and, thus, alter molecular pathways of oxygen sensing such as those regulated by hypoxia-inducible factor (HIF). In normoxia, the alpha subunit of HIF-1 is a target for prolyl hydroxylase, which makes HIF-1alpha a target for degradation by the proteasome. During hypoxia, prolyl hydroxylase is inhibited since it requires oxygen as a cosubstrate. Thus, hypoxia allows HIF to accumulate and translocate into the nucleus for induction of target genes regulating glycolysis, angiogenesis and hematopoiesis. By this mechanism, UCPs activity may contribute to increase the expression of genes related to the formation of blood vessels, and thus promote tumor growth.
Roles of UCPs in Cancer Energy Metabolism
Lynen and colleagues proposed that the root of the Warburg effect is not in the inability of mitochondria to carry out respiration, but rather would rely on their incapacity to synthesize ATP in response to membrane potential.
The ability of UCPs to uncouple ATP synthesis from respiration and the fact that UCP2 is overexpressed in several chemoresistant cancer cell lines and primary human colon cancers have lead to speculate about the existence of a link between UCPs and the Warburg effect. As mentioned above, uncoupling induced by overexpression of UCP2 has been shown to prevent ROS formation, and, in turn, increase apoptotic threshold in cancer cells, providing a pro-survival advantage and a resistance mechanism to cope with ROS-inducing chemo-therapeutic agents.
Mitochondrial Krebs cycle is one of the sources for these anabolic precursors. The export of these metabolites to cytoplasm for anabolic purposes involves the replenishment of the cycle intermediates by anaplerotic substrates such as pyruvate and glutamate. Thus, glycolysis-derived pyruvate, as well as alpha-ketoglutarate derived from glutaminolysis, may be necessary to sustain anaplerotic reactions. At the same time, to keep Krebs cycle functional, the reduced cofactors NADH and FADH2 would have to be re-oxidized, a function which relies on the mitochondrial respiratory chain. Once again, uncoupling may be crucial for cancer cell mitochondrial metabolism, allowing Krebs cycle to be kept functional to meet the vigorous biosynthetic demand of cancer cells.
Several cancer cells resistant to chemotherapeutics and radiation often exhibit higher rates of fatty acid oxidation and it has been observed that inhibition of fatty acid oxidation potentiates apoptotic death induced by chemotherapeutic agents. These findings are in agreement with the proposed need of fatty acid for the activity of UCPs, suggesting that the lack of these potential substrates or activators would decrease uncoupling activity, subsequently increasing membrane potential, ROS production and therefore lowering apoptotic threshold.
Roles of UCPs in Cancer Cachexia
Cachexia is a wasting syndrome characterized by weakness, weight and fat loss, and muscle atrophy which is often seen in patients with advanced cancer or AIDS. Cachexia has been suggested to be responsible for at least 20 % of cancer deaths and also plays an important part in the compromised immunity leading to death from infection. The imbalance between energy intake and energy expenditure underlying cachexia cannot be reversed nutritionally.
Alterations leading to high energy expenditure, such as excessive proton leak or mitochondrial uncoupling, are likely mechanisms underlying cachexia. In fact, increased expression of UCP1 in BAT and UCP2 and UCP3 in skeletal muscle have been shown in several murine models of cancer cachexia
Roles of UCPs in Chemoresistance
Cancer cells acquire drug resistance as a result of selection pressure dictated by unfavorable microenvironments. Although mild uncoupling may clearly be useful under normal conditions or under severe or chronic metabolic stress such as hypoxia or anoxia, it may be a mechanism to elude oxidative stress-induced apoptosis in advanced cancer cells. Several anti-cancer treatments are based on promotion of ROS formation, to induce cell growth arrest and apoptosis. Thus, increased UCP levels in cancer cells, rather than a marker of oxidative stress, may be a mechanisms conferring anti-apoptotic advantages to the malignant cell, increasing their ability to survive in adverse microenvironments, radiotherapy and chemotherapy. UCPs appear to play a permissive role in tumor cell survival and growth.
Expression of UCPs promote bioenergetics adaptation and cell survival. UCPs appear to be critical to determine the sensitivity of cancer cells to several chemotherapeutic agents and radiotherapy, interfering with the activation of mitochondria driven apoptosis.
From a therapeutic viewpoint, inhibition of glycolysis in UCP2 expressing tumours or specific inhibition of UCP2 are, respectively, attractive strategies to target the specific metabolic signature of cancer cells.
Hypoxia-inducible factor-1 in tumour angiogenesis
HIF-b subunits, is a heterodimeric transcriptional activator. In response to
hypoxia,
- stimulation of growth factors, and
- activation of oncogenes as well as carcinogens,
HIF-1a is overexpressed and/or activated and targets those genes which are required for angiogenesis, metabolic adaptation to low oxygen and promotes survival.
Several dozens of putative direct HIF-1 target genes have been identified on the basis of one or more cis-acting hypoxia-response elements that contain an HIF-1 binding site. Activation of HIF-1 in combination with activated signaling pathways and regulators is implicated in tumour progression and prognosis.
In order for a macroscopic tumour to grow, adequate oxygen delivery must be effected via tumor angiogenesis that results from an increased synthesis of angiogenic factors and a decreased synthesis of anti-angiogenic factors. The metabolic adaptation of tumor cells to reduced oxygen availability by increasing glucose transport and glycolysis to promote survival are important consequences in response to hypoxia.
Hypoxia and HIF-1
Hypoxia is one of the major drivers to tumour progression as hypoxic areas form in human tumours when the growth of tumour cells in a given area outstrips local neovascularization, thereby creating areas of inadequate perfusion. Although several transcriptional factors have been reported to be involved
in the response to hypoxic stress such as AP-1, NF-kB and HIF-1, HIF-1 is the most potent inducer of the expression of genes such as those encoding for glycolytic enzymes, VEGF and erythropoietin.
HIF-a subunit exists as at least three isoforms, HIF-1a, HIF-2a and HIF-3a. HIF-1a and HIF-2a can form heterodimers with HIF-b. Although HIF-b subunits are constitutive nuclear proteins, both HIF-1a andHIF-2a subunits are strongly induced by hypoxia in a similar manner. HIF-1a is up-regulated in hypoxic tumour cells and activates the transcription of target genes by binding to cis-acting enhancers, hypoxic responsive element (HRE) close to the promoters of these genes with a result of tumour cellular adaptation to hypoxia and tumour angiogenesis, and promotion of further growth of the primary tumour. Studies have shown HIF-1a to be over-expressed by both tumour cells and such stromal cells as macrophages in many forms of human malignancy.
Regulation of HIF-1
The first regulator of HIF-1 is oxygen. HIF-1α appears to be the HIF-1 subunit regulated by hypoxia. The oxygen sensors in the HIF-1α pathway are two kinds of oxygen dependent hydroxylases. One is prolyl hydroxylase which could hydroxylize the proline residues 402 and 564 at the oxygen dependent domain (ODD) of HIF-1 in the presence of oxygen and iron with a result of HIF-α degradation. The other is hydroxylation of Asn803 at the C-terminal transactivation domain (TAD-C) by FIH-1, which could inhibit the interaction of HIF-1α with co-activator p300 with a subsequent inhibition of HIF-1α transactivity. The hydroxylation of proline 564 at ODD of HIF-1α under normoxia was shown using a novel hydroxylation-specific antibody to detect hydroxylized HIF-1α.
Oncogene comes as the second regulator. Many oncogenes have effects on HIF-1α. Among them, some function in regulation of HIF-1α protein stability or degradation, others play roles in several activated signaling pathways. Tumor suppressor genes as p53 and von Hippel-Lindau (VHL) influence the levels and functions of HIF-1. The wild type (wt) form of p53 protein was involved in inhibiting HIF-1 activity by targeting the HIF-1a subunit for Mdm2-mediated ubiquitination and proteasomal degradation, and in inducing inhibitors of angiogenesis such as thrombospondin-1, while loss of wt p53 (by gene deletion or mutation) could enhance HIF-1α accumulation in hypoxia.
The third regulator is a battery of growth factors and cytokines from stromal and parenchymal cells such as
- EGF,
- transforming growth factor-α,
- insulin-like growth factors 1 and 2,
- heregulin, and interleukin-1b
via autocrine and paracrine pathways. These regulators not only induce the expression of HIF-1α protein, HIF-1 DNA binding activity and transactivity, but also make HIF-1 target gene expression under normoxia or hypoxia.
The fourth one is a group of reactive oxygen species (ROS) resulting from carcinogens such as Vanadate and Cr (VI) or stimulation of cytokines such as angiotensin and TNFa. However, it seems controversial when it comes to the production of ROS under hypoxia and their individual role in regulation of HIF-1a. It is well known that ROS plays an important role in carcinogenesis induced by a variety of carcinogens.
Signaling Pathways Involved in Regulation of HIF-1α
HIF-1 is a phosphorylated protein and its phosphorylation is involved in HIF-1a subunit expression and/or stabilization as well as in the regulation of HIF-1 transcriptional activity. Three signaling pathways involved in the regulation of HIF-1α have been reported to date.
- The PI-3k pathway has been mainly and frequently implicated in regulation of HIF-1α protein expression and stability.
- Akt is also activated by hypoxia. Activated Akt initiates two different pathways in regulation of HIF-1α. The function of these two pathways appears to show consistent impact on HIF-1α activation.
- Signal transduction pathway in HIF-1α regulation.Oncogenes, growth factors and hypoxia have been documented to regulate HIF-1α protein and increase its transactivity. GSK and mTOR were two target events of Akt and could contribute to decreasing HIF-1α degradation and increasing HIF-1α protein synthesis. Activated ERK1/2 could mainly up-regulate.
HIF-1a, Angiogenesis and Tumour Prognosis
Hypoxia, oncogenes and a variety of growth factors and cytokines increase HIF-1α stability and/or synthesis and transactivation to initiate tumour angiogenesis, metabolic adaptation to hypoxic situation and promote cell survival or anti-apoptosis resulting from a consequence of more than sixty putative direct HIF-1 target gene expressions.
The crucial role of HIF-1 in tumour angiogenesis has sparked scientists and clinical researchers to try their best to understand the whole diagram of HIF-1 so as to find out novel approaches to inhibit HIF-1 overexpression. Indeed, the combination of anti-angiogenic agent and inhibitor of HIF-1 might be particularly efficacious, as the angiogenesis inhibitor would cut off the tumour’s blood supply and HIF-1 inhibitor would reduce the ability of tumour adaptation to hypoxia and suppress the proliferation and promote apoptosis. Screens for small-molecule inhibitors of HIF-1 are underway and several agents that inhibit HIF-1, angiogenesis and xenograft growth have been identified.
Hypoxia, autophagy, and mitophagy in the tumor stroma
Metabolomic profiling reveals that Cav-1(-/-) null mammary fat pads display a highly catabolic metabolism, with the increased release of several metabolites, such as amino acids, ribose and nucleotides, and a shift towards gluconeogenesis, as well as mitochondrial dysfunction. These changes are consistent with increased autophagy, mitophagy and aerobic glycolysis, all processes that are induced by oxidative stress. Autophagy or ‘self-eating’ is the process by which cells degrade their own cellular components to survive during starvation or to eliminate damaged organelles after oxidative stress. Mitophagy, or mitochondrial-autophagy, is particularly important to remove damaged ROS-generating mitochondria.
An autophagy/mitophagy program is also triggered by hypoxia. Hypoxia is a common feature of solid tumors, and promotes cancer progression, invasion and metastasis. Interestingly, via induction of autophagy, hypoxia is sufficient to induce a dramatic loss of Cav-1 in fibroblasts. The hypoxia-induced loss of Cav-1 can be inhibited by the autophagy inhibitor chloroquine, or by pharmacological inhibition of HIF1α. Conversely, small interfering RNA-mediated Cav-1 knock-down is sufficient to induce pseudo-hypoxia, with HIF1α and NFκB activation, and to promote autophagy/mitophagy, as well as a loss of mitochondrial membrane potential in stromal cells. These results indicate that a loss of stromal Cav-1 is a marker of hypoxia and oxidative stress.
In a co-culture model, autophagy in cancer-associated fibroblasts was shown to promote tumor cell survival via the induction of the pro-autophagic HIF1α and NFκB pathways in the tumor stromal microenvironment. Finally, the mitophagy marker Bnip3L is selectively upregulated in the stroma of human breast cancers lacking Cav-1, but is notably absent from the adjacent breast cancer epithelial cells.
Metabolome profiling of several types of human cancer tissues versus corresponding normal tissues have consistently shown that cancer tissues are highly catabolic, with the significant accumulation of many amino acids and TCA cycle metabolites. The levels of reduced glutathione were decreased in primary and metastatic prostate cancers compared to benign adjacent prostate tissue, suggesting that aggressive disease is associated with increased oxidative stress. Also, these data show that the tumor microenvironment has increased oxidative-stress-induced autophagy and increased catabolism.
Taken together, all these findings suggest an integrated model whereby
A loss of stromal Cav-1 induces autophagy/mitophagy in the tumor stroma, via oxidative stress.
This creates a catabolic micro-environment with the local accumulation of chemical building blocks and recycled nutrients (such as amino acids and nucleotides), directly feeding cancer cells to sustain their survival and growth.
This novel idea is termed the ‘autophagic tumor stroma model of cancer’ .
This new paradigm may explain the ‘autophagy paradox’, which is based on the fact that both the systemic inhibition and systemic stimulation of autophagy prevent tumor formation.
What is presented suggests that vectorial energy transfer from the tumor stroma to cancer cells directly sustains tumor growth, and that interruption of such metabolic coupling will block tumor growth. Autophagy inhibitors (such as chloroquine) functionally block the catabolic transfer of metabolites from the stroma to the tumor, inducing cancer cell starvation and death. Conversely, autophagy inducers (such as rapamycin) promote autophagy in tumor cells and induce cell death. Thus, both inhibitors and inducers of autophagy will have a similar effect by severing the metabolic coupling of the stroma and tumor cells, resulting in tumor growth inhibition (cutting ‘off ’ the fuel supply).
This model may also explain why enthusiasm for antiangiogenic therapy has been dampened. In most cases, the clinical benefits are short term, and more importantly, new data suggest an unexpected link between anti-angiogenic treatments and metastasis. In pre-clinical models, anti-vascular endothelial growth factor (anti-VEGF) drugs (sunitinib and anti-VEGFR2 blocking antibodies) were shown to inhibit localized tumor formation, but potently induced relapse and metastasis. Thus, by inducing hypoxia in the tumor microenvironment, antiangiogenic drugs may create a more favorable metastatic niche.
Glutamine, glutaminolysis.
In direct support that cancer cells use mitochondrial oxidative metabolism, many investigators have shown that cancer cells are ‘addicted’ to glutamine. Glutamine is a non-essential amino acid that is metabolized to glutamate and enters the TCA cycle as α-ketoglutarate, resulting in high ATP generation via oxidative phosphorylation. Recent studies also show that ammonia is a by-product of glutaminolysis. In addition, ammonia can act as a diffusible inducer of autophagy. Given these observations, glutamine addiction in cancer cells provides another mechanism for driving and maintaining autophagy in the tumor micro-environment .
In support of this idea, a loss of Cav-1 in the stroma is sufficient to drive autophagy, resulting in increased glutamine production in the tumor micro-environment. Thus, this concept defines a new vicious cycle in which autophagy in the tumor stroma transfers glutamine to cancer cells, and the by-product of this metabolism, ammonia, maintains autophagic glutamine production. This model fits well with the ‘autophagic tumor stroma model of cancer metabolism’, in which energy rich recycled nutrients (lactate, ketones, and glutamine) fuel oxidative mitochondrial metabolism in cancer cells.
Glutamine utilization in cancer cells and the tumor stroma. Oxidative mitochondrial metabolism of glutamine in cancer cells produces ammonia. Ammonia production is sufficient to induce autophagy. Thus, autophagy in cancer-associated fibroblasts provides cancer cells with an abundant source of glutamine. In turn, the ammonia produced maintains the autophagic phenotype of the adjacent stromal fibroblasts.
Lessons from other paradigms
an infectious parasitic cancer cell that metastasizes and captures mitochondrial DNA from host cells
Cancer cells behave like ‘parasites’, by inducing oxidative stress in normal host fibroblasts, resulting in the production of recycled nutrients via autophagy.
This is exactly the same mechanism by which infectious parasites (such as malaria) obtain nutrients and are propagated by inducing oxidative stress and autophagy in host cells. In this regard, malaria is an ‘intracellular’ parasite, while cancer cells may be thought of as ‘extracellular’ parasites. This explains why chloroquine is both an effective antimalarial drug and an effective anti-tumor agent, as it functions as an autophagy inhibitor, cutting off the ‘fuel supply’ in both disease states.
Human cancer cells can ‘steal’ live mitochondria or mitochondrial DNA from adjacent mesenchymal stem cells in culture, which then rescues aerobic glycolysis in these cancer cells. This is known as mitochondrial transfer. Interestingly, metastatic breast cancer cells show the up-regulation of numerous mitochondrial proteins, specifically associated with oxidative phosphorylation, as seen by unbiased proteomic analysis.
Thus, increased mitochondrial oxidative metabolism may be a key driver of tumor cell metastasis. In further support of this argument, treatment of MCF7 cancer cells with lactate is indeed sufficient to induce mitochondrial biogenesis in these cells.
To determine if these findings may be clinically relevant, a lactate-induced gene signature was recently generated using MCF7 cells. This gene signature shows that lactate induces ‘stemness’ in cancer cells, and this lactate induced gene signature predicts poor clinical outcome (including tumor recurrence and metastasis) in breast cancer patients.
REFERENCES
Li W and Zhao Y. “Warburg Effect” and Mitochondrial Metabolism in Skin Cancer. Epidermal Pigmentation, Nucleotide Excision Repair and Risk of Skin Cancer. J Carcinogene Mutagene 2012; S4:002 doi:10.4172/2157-2518.
Seyfried TN, Shelton LM. Cancer as a metabolic disease. Nutrition & Metabolism 2010; 7:7(22 pg). doi:10.1186/1743-7075-7-7
Israël M, Schwartz L. The metabolic advantage of tumor cells. Molecular Cancer 2011, 10:70-82. http://www.molecular-cancer.com/content/10/1/70
Valle A, Oliver J, Roca P. Role of Uncoupling Proteins in Cancer. Cancers 2010, 2, 567-591; doi:10.3390/cancers2020567
Ishii H, Doki Y, Mori M. Perspective beyond Cancer Genomics: Bioenergetics of Cancer Stem Cells. Yonsei Med J 2010; 51(5):617-621. DOI 10.3349/ymj. 2010.51.5.617 pISSN: 0513-5796, eISSN: 1976-2437
Sotgia F, Martinez-Outschoorn, Pavlides S,Howell A . Understanding the Warburg effect and the prognostic value of stromal caveolin-1 as a marker of a lethal tumor microenvironment. Breast Cancer Research 2011, 13:213-26. http://breast-cancer-research.com/content/13/4/213
Yong-Hong Shi, Wei-Gang Fang. Hypoxia-inducible factor-1 in tumour angiogenesis. World J Gastroenterol 2004; 10(8): 1082-1087. http:// wjgnet.com /1007-9327/10/1082.asp
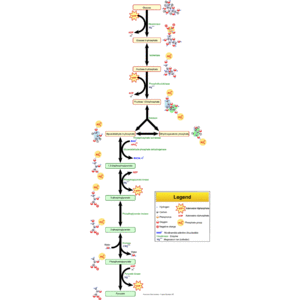
English: Glycolysis pathway overview. (Photo credit: Wikipedia)
Normal
0
false
false
false
EN-US
X-NONE
X-NONE
/* Style Definitions */
table.MsoNormalTable
{mso-style-name:”Table Normal”;
mso-tstyle-rowband-size:0;
mso-tstyle-colband-size:0;
mso-style-noshow:yes;
mso-style-priority:99;
mso-style-parent:””;
mso-padding-alt:0in 5.4pt 0in 5.4pt;
mso-para-margin-top:0in;
mso-para-margin-right:0in;
mso-para-margin-bottom:10.0pt;
mso-para-margin-left:0in;
line-height:115%;
mso-pagination:widow-orphan;
font-size:11.0pt;
font-family:”Calibri”,”sans-serif”;
mso-ascii-font-family:Calibri;
mso-ascii-theme-font:minor-latin;
mso-hansi-font-family:Calibri;
mso-hansi-theme-font:minor-latin;}
Warburg Effect, glycolysis, pyruvate kinase, PKM2, PIM2, mtDNA, complex I, NSCLC
Ribeirão Preto Area, Brazil|Government Relations
Current- University, USP-FMRP Physiology – Biochemistry
Previous- Imperial Cancer Research Fund Lab, Instituto de Investigaciones Bioquimicas, Luta armada e CIA
Education – FMRP-USP
While exile interrupted ny graduation in Medicine, started a port grade at Leloir´s Instituto Investigaciones bioquímicas Buenos Aires Argentina Jan 1970 – jan 1972. PhD from 1972- jan 1976 FMRP-USP . Post Doc ICRF London England 1979 -1980
“Those…..whose acquitance with scientific research is derived chiefly from its practical results easily develop a completely false notion of the { scientific } mentality.”
My goal and previous experience besides laboratory work was dedicated to solve this apparent conflict.
When Pasteur did his work studying the chemistry outside yeast-cells, he was able to perceive that in anaerobiosis yeast cells were able to convert sugar at a great velocity to its end products. While the same yeast cells, therefore with the same genome did it at a very slow speed in aerobiosis. Warburg tested tumor cells for the same and found that while normal cells from the organ in which he has found tumors presented a similar metabolic regulatory response to anaerobic/aerobic transition tumor cells did not. Tumos cells continued to display strong acidification (producing great amounts of lactate in the culture media) in aerobiosis.
Tumor cells displayed a failure in this regulatory mechanism that, he O. Warburg, named Pasteur effect. He also noticed that this defect continues from old cancer cells to newer ones. Therefore, for him, it is a genetic defect. Furthermore, as mitochondria generates newer mitochondria in the cells, the genetic component is in the mitochondria that were the site of core metabolism in aerobiosis.
Larry Bernstein
Part of what you describe is in Warburg biography by Hans Krebs (out of print). He also refers to a Meyerhof Quotient to express the degree of metabolic anaerobiosis. I don’t recall a reference to a mitochondrial “genetic” defect. That is farsighted. He did conclude that once the cancer cells were truly anaplastic, metastatic behavior was irreversible. It was only later that another Nobelist who described fatty acid synthesis, I think, concluded that the synthetic process tied up the same aerobic pathway so that anaerobic glycolysis became essential for the cells energetics. You can correct me if I’m in error. Where does the substrate come from? Lean body mass breaks down to provide gluconeogenic precursors. Points of concern are – deamination of branch chain AAs, the splitting of a 6 carbon sugar into 2 3-carbon chains, and the conversion of pyruvate to lactate with the reverse reaction blocked. There is also evidence that there is an impairment in the TCA cycle at the point of fumarase. Then there is never any consideration of the flow of substrates back and forth across the mitochondrial membrane (malate, aspartate), and the redox potentials.
Jose: For reasons independent of my will, the copy of Science,123(3191):309-14,1956 translation of O Warburg original article is not in my hands. Some that do not find any alternative form to respond to my critics concerning molecular biology distortion of biochemistry left on purpose, almost all of my older reprints on the rain. Anyway, O. Warburg refers to mitochondria using the expression of “grana” or “grains” if my recollections are correct. The original work of O Warburg is one of 1924 – Biochemisch ZeiTschrift.,152:51-60.1924. “Weberssert method zur messing der atmung un glycolyse, drese zeitschr.” Other aspects of your interesting comment I will try to comment latter.
By Jose Eduardo de Salles Roselino
Larry Bernstein, not as a correction but , following my line of reasoning about carbono fluxes…It is almost impossible to figure out what was really inside the mind-set of a scientific researcher at the time he has performed his work. Anyway, by the knowledge available at his time, we may conjecture about, even when we acknowledge that, what he may have had in mind at the start could be quite different from what he have latter published from his results.
In case, we try to present the ideas behind Warburg´s works we may take into account the following pre-existing knowledge: Lavoisier (done by 1779-1784) measured very carefully the amount of heat released by respiration and chemical oxidative processes. Reached the conclusion that respiration was slower but essentially similar to carbon combustion by chemical oxidative processes. By early XIX caloric values for gram of sugar, lipids and proteins where made clear. T Schwann recognized that yeast cells convert sugar in ethanol plus a volatile acid (carbon dioxide). Pasteur-effect was seen just as a change in the velocity of product production from a same sugar and/or a decrease in sugar concentration in the growth medium. This is a change in carbon flux velocity in the oxidative process calorimetrically measured by Lavoisier.
In Germany, however, the preponderance of organic chemical point of view has great scientists as Berzelius, Liebig etc. to consider that yeast where similar to inorganic catalysts. The oxidative process was caused by oxygen only. You may amuse yourself reading, how they attack T. Schwann proposal in Annalen, 29: 100 (1839). When O Warburg paid tribute to L. Pasteur, we can see clearly on which side he was. Furthermore, his apparatus, in which I was introduced to this line of research, a rather modern version at that time (1960s) that look like a very futuristic robotic version of the original with a pair of blinking red and green round lights as two eyes on a metallic head, offers a very important clue about the change in carbon flow.
Inside a Warburg glass flask, kept at very finely controlled temperature to avoid changes in pressure derived from changes in the temperature, you would certainly find a central well in which the volatile acid must be trapped in order to determine a pressure reduction only due to the decrease in oxygen pressure. Inside this central well, you could determine one product (carbon dioxide) and in the outside of it where tissues or cells where in the media you could also determine the other end product of the oxidative combustion of sugar (lactate). This has provide him with the result that indicate not only a change in the velocity as originally found by Pasteur in yeast cells but in the organic flow of sugar carbons.
comment – E-mail: vbungau11@yahoo.com
-About carcinogenesis- Electronegativity is a nucleus of an atom’s ability to attract and maintain a cloud of electrons. Copper atom electronegativity is higher than the Iron atom electronegativity. Atom with lower electronegativity (Iron),remove the atom with higher electronegativity (Copper) of combinations. This means that,in conditions of acidosis, we have cytochrome oxidase with iron (red, neoplastic), instead of cytochrome oxidase with copper (green, normal).I think this is the key to carcinogenesis. Siincerely, Dr. Viorel Bungau
Read Full Post »