The Delicate Connection: IDO (Indolamine 2, 3 dehydrogenase) and Cancer Immunology
Author and Curator: Demet Sag, PhD, CRA, GCP
Table of Contents:
- Abstract
- Dual role for IDO
- Immune System and IDO
- Autoimmune disorders and IDO
- Cancer and Ido
- Clinical Interventions
- Clinical Trials
- Future Actions for Molecular Dx and Targeted Therapies:
- Conclusion
- References
TABLE 1- IDO Clinical Trials
TABLE 2- Kyn induced Genes
TABLE 3 Possible biomarkers and molecular diagnostics targets
TABLE 4: Current Interventions ______________________________________________________________________________________________________________
ABSTRACT:
Overall purpose is to find a method to manipulate IDO for clinical applications, mainly the focus of this review is is cancer prevention and treatment. The first study proving the connection between IDO and immune response came from, a very natural event, a protection of
pregnancy in human. This led to discover that high IDO expression is a common factor in cancer tumors. Thus, attention promoted investigations on IDO’s role in various disease states, immune disorders, transplantation, inflammation, women health, mood disorders.
Many approaches, vaccines and adjuvants are underway to find new immunotherapies by combining the power of
DCs in immune response regulation and specific direction of
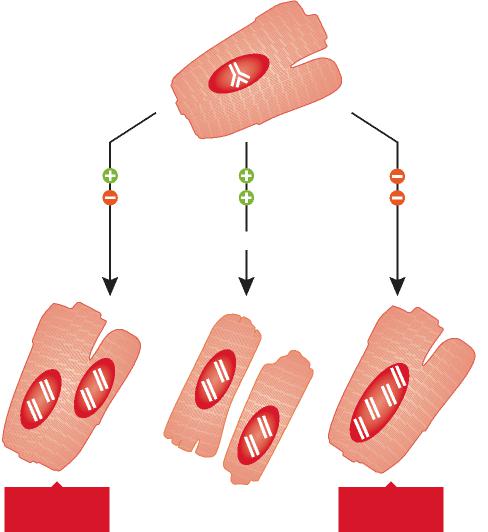
siRNA. As a result, with this unique qualities of IDO, DCs and siRNA, we orchestrated a novel intervention for immunomodulation of IDO by inhibiting with small interference RNA, called siRNA-IDO-DCvax. Proven that our DCvax created a delay and regression of tumor growth without changing the natural structure and characterization of DCs in melanoma and breast cancers in vivo. (** The
shRNA IDO- DCvax is developed by Regen BioPhrama,
San Diego, CA , Thomas Ichim, Ph.D, CSO. and David Koos, CEO)
______________________________________________________________________________________________________________
Double-Edged Sword of IDO: The Good and The Bad for Clinical intervention and Developments
IDO almost has a dual role. There is a positive side of high expression of IDO during pregnancy (29; 28; 114), transplants (115; 116; 117; 118; 119), infectious diseases (96) and but this tolerance is negative during autoimmune-disorders (120; 121; 122), tumors of cancer (123; 124; 117; 121; 125; 126; 127) (127), and mood disorders (46). The increased IDO expression has a double-edged sword in human physiology provides a positive role during protection of fetus and grafts after transplantations but becomes a negative factor during autoimmune disorders, cancer, sepsis and mood disorders.
Prevention of allogeneic fetal rejection is possible by tryptophan metabolism (26) rejecting with lack of IDO but allocating if IDO present (29; 28; 114). These studies lead to find “the natural regulation mechanism” for protecting the transplants from graft versus host disease GVHD (128) and getting rid of tumors.
The plasticity of mammary and uterus during reproduction may hold some more answers to prevent GVHD and tumors of cancer with good understanding of IDO and tryptophan mechanism (129; 130). After allogeneic bone marrow transplants the risk of solid tumor development increased about 80% among 19,229 patients even with a greater risk among patients under 18 years old (117). The adaptation of tolerance against host mechanism is connected to the IDO expression (131). During implantation and early pregnancy IDO has a role by making CD4+CD25+Foxp3+ regulatory T cells (Tregs) and expressing in DCs and MQs (114; 132; 133).
Clonal deletion mechanism prevents mother to react with paternal products since female mice accepted the paternal MHC antigen-expressing tumor graft during pregnancy and rejected three weeks after delivery (134). CTLA-4Ig gene therapy alleviates abortion through regulation of apoptosis and inhibition of spleen lymphocytes (135).
Immune System and IDO DCs are the orchestrator of the immune response (56; 57; 58) with list of functions in uptake, processing, and presentation of antigens; activation of effector cells, such as T-cells and NK-cells; and secretion of cytokines and other immune-modulating molecules to direct the immune response. The differential regulation of IDO in distinct DC subsets is widely studied to delineate and correct immune homeostasis during autoimmunity, infection and cancer and the associated immunological outcomes. Genesis of antigen presenting cells (APCs), eventually the immune system, require migration of monocytes (MOs), which is originated in bone marrow. Then, these MOs move from bloodstream to other tissues to become macrophages and DCs (59; 60).
Initiation of immune response requires APCs to link resting helper T-cell with the matching antigen to protect body. DCs are superior to MQs and MOs in their immune action model. When DCs are first described (61) and classified, their role is determined as a highly potent antigen-presenting cell (APC) subset with 100 to 1000-times more effective than macrophages and B-cells in priming T-cells. Both MQs and monocytes phagocytize the pathogen, and their cell structure contains very large nucleus and many internal vesicles. However, there is a nuance between MQ and DCs, since DCs has a wider capacity of stimulation, because MQs activates only memory T cells, yet DCs can activate both naïve and memory T cells.
DCs are potent activators of T cells and they also have well controlled regulatory roles. DC properties determine the regulation regardless of their origin or the subset of the DCs. DCs reacts after identification of the signals or influencers for their inhibitory, stimulatory or regulatory roles, before they express a complex repertoire of positive and negative cytokines, transmembrane proteins and other molecules. Thus, “two signal theory” gains support with a defined rule. The combination of two signals, their interaction with types of cells and time are critical.
In short, specificity and time are matter for a proper response. When IDO mRNA expression is activated with CTL40 ligand and IFNgamma, IDO results inhibition of T cell production (4). However, if DCs are inhibited by 1MT, an inhibitor of IDO, the response stop but IgG has no affect (10). In addition, if the stimulation is started by a tryptophan metabolite, which is downstream of IDO, such as 3-hydroxyantranilic or quinolinic acids, it only inhibits Th1 but not Th2 subset of T cells (62).
Furthermore, inclusion of signal molecules, such as Fas Ligand, cytochrome c, and pathways also differ in the T cell differentiation mechanisms due to combination, time and specificity of two-signals. The co-culture experiments are great tool to identify specific stimuli in disease specific microenvironment (63; 12; 64) for discovering the mechanism and interactions between molecules in gene regulation, biochemical mechanism and physiological function during cell differentiation.
As a result, the simplest differential cell development from the early development of DCs impact the outcome of the data. For example, collection of MOs from peripheral blood mononuclear cells (PBMCs) with IL4 and GM-CSF leads to immature DCs (iDCs). On next step, treatment of iDCs with tumor necrosis factor (TNF) or other plausible cytokines (TGFb1, IFNgamma, IFNalpha, IFNbeta, IL6 etc.) based on the desired outcome differentiate iDCs into mature DCs (mDCs). DCs live only up to a week but MOs and generated MQs can live up to a month in the given tissue. B cells inhibit T cell dependent immune responses in tumors (65).
AutoImmune Disorders:

The balance of IDO expression becomes necessary to prevent overactive immune response self-destruction, so modulation in tryptophan and NDA metabolisms maybe essential. When splenic IDO-expressing CD11b (+) DCs from tolerized animals applied, they suppressed the development of arthritis, increased the Treg/Th17 cell ratio, and decreased the production of inflammatory cytokines in the spleen (136).
The role of Nicotinamide prevention on type 1 diabetes and ameliorates multiple sclerosis in animal model presented with activities of NDAs stimulating GPCR109a to produce prostaglandins to induce IDO expression, then these PGEs and PGDs converted to the anti-inflammatory prostaglandin, 15d-PGJ(2) (137; 138; 139). Thus, these events promotes endogenous signaling mechanisms involving the GPCRs EP2, EP4, and DP1 along with PPARgamma. (137).
Modulating the immune response at non-canonical at canonocal pathway while keeping the non-canonical Nf-KB intact may help to mend immune disorders. As a result, the targeted blocking in canonical at associated kinase IKKβ and leaving non-canonocal Nf-kB pathway intact, DCs tips the balance towards immune supression. Hence, noncanonical NF-κB pathway for regulatory functions in DCs required effective IDO induction, directly or indirectly by endogenous ligand Kyn and negative regulation of proinflammatory cytokine production. As a result, this may help to treat autoimmune diseases such as rheumatoid arthritis, type 1 diabetes, inflammatory bowel disease, and multiple sclerosis, or allergy or transplant rejection.
While the opposite action needs to be taken during prevention of tumors, that is inhibition of non-canonical pathway. Inflammation induces not only relaxation of veins and lowering blood pressure but also stimulate coagulopathies that worsen the microenvironment and decrease survival rate of patients after radio or chemotherapies.Cancer Generating tumor vaccines and using adjuvants underway (140).
Clinical correlation and genetic responses also compared in several studies to diagnose and target the system for cancer therapies (127; 141; 131). The recent surveys on IDO expression and human cancers showed that IDO targeting is a candidate for cancer therapy since IDO expression recruiting Tregs, downregulates MHC class I and creating negative immune microenvironment for protection of development of tumors (125; 27; 142). Inhibition of IDO expression can make advances in immunotherapy and chemotherapy fields (143; 125; 131; 144).
IDO has a great importance on prevention of cancer development (126). There are many approaches to create the homeostasis of immune response by Immunotherapy. However, given the complexity of immune regulations, immunomodulation is a better approach to correct and relieve the system from the disease. Some of the current IDO targeted immunotherapy or immmunomodulations with RNA technology for cancer prevention (145; 146; 147; 148; 149; 150) or applied on human or animals (75; 151; 12; 115; 152; 9; 125) or chemical, (153; 154) or radiological (155). The targeted cell type in immune system generally DCs, monocytes (94)T cells (110; 156)and neutrophils (146; 157). On this paper, we will concentrate on DCvax on cancer treatments.
IDO and the downstream enzymes in tryptophan pathway produce a series of immunosuppressive tryptophan metabolites that may lead into Tregs proliferation or increase in T cell apoptosis (62; 16; 27; 158), and some can affect NK cell function (159).
The interesting part of the mechanism is even without presence of IDO itself, downstream enzymes of IDO in the kynurenine tryptophan degradation still show immunosuppressive outcome (160; 73) due to not only Kyn but also TGFbeta stimulated long term responses. DC vaccination with IDO plausible (161) due to its power in immune response changes and longevity in the bloodstream for reversing the system for Th17 production (162).
Clinical Interventions are taking advantage of the DC’s central role and combining with enhancing molecules for induction of immunity may overcome tolerogenic DCs in tumors of cancers (163; 164).
The first successful application of DC vaccine used against advanced melanoma after loading DCs with tumor peptides or autologous cell lysate in presence of adjuvants keyhole limpet hematocyanin (KLH) (165). Previous animal and clinical studies show use of DCs against tumors created success (165; 166; 167) as well as some problems due to heterogeneity of DC populations in one study supporting tumor growth rather than diminishing (168).
DC vaccination applied onto over four thousand clinical trial but none of them used siRNA-IDO DC vaccination method. Clinical trials evaluating DCs loaded ex vivo with purified TAAs as an anticancer immunotherapeutic interventions also did not include IDO (Table from (169). This table presented the data from 30 clinical trials, 3 of which discontinued, evaluating DCs loaded ex vivo with TAAs as an anticancer immunotherapy for 12 types of cancer [(AML(1), Breast cancer (4), glioblastoma (1), glioma (2), hepatocellular carcinoma (1), hematological malignancies (1), melanoma (6), neuroblastoma sarcoma (2), NSCLC (1), ovarian cancer (3), pancreatic cancer (3), prostate cancer (10)] at phase I, II or I/II.
Tipping the balance between Treg and Th17 ratio has a therapeutic advantage for restoring the health that is also shown in ovarian cancer by DC vaccination with adjuvants (161). This rebalancing of the immune system towards immunogenicity may restore Treg/Th17 ratio (162; 170) but it is complicated. The stimulation of IL10 and IL12 induce Treg produce less Th17 and inhibiting CTL activation and its function (76; 171; 172) while animals treated with anti-TGFb before vaccination increase the plasma levels of IL-15 for tumor specific T cell survival in vivo (173; 174) ovarian cancer studies after human papilloma virus infection present an increase of IL12 (175).
Opposing signal mechanism downregulates the TGFb to activate CTL and Th1 population with IL12 and IL15 expression (162; 173). The effects of IL17 on antitumor properties observed by unique subset of CD4+ T cells (176) called also CD8+ T cells secrete even more IL17 (177).
Using cytokines as adjuvants during vaccination may improve the efficacy of vaccination since cancer vaccines unlike infections vaccines applied after the infection or disease started against the established adoptive immune response. Adjuvants are used to improve the responses of the given therapies commonly in immunotherapy applications as a combination therapy (178).
Enhancing cancer vaccine efficacy via modulation of the microenvironment is a plausible solution if only know who are the players. Several molecules can be used to initiate and lengthen the activity of intervention to stimulate IDO expression without compromising the mechanism (179). The system is complicated so generally induction is completed ex-vivo stimulation of DCs in cell lysates, whole tumor lysates, to create the microenvironment and natural stimulatory agents. Introduction of molecules as an adjuvants on genetic regulation on modulation of DCs are critical, because order and time of the signals, specific location/ tissue, and heterogeneity of personal needs (174; 138; 180). These studies demonstrated that IL15 with low TGFb stimulates CTL and Th1, whereas elevated TGFb with IL10 increases Th17 and Tregs in cancer microenvironments.

For example Ret-peptide antitumor vaccine contains an extracellular fragment of Ret protein and Th1 polarized immunoregulator CpG oligonucleotide (1826), with 1MT, a potent inhibitor of IDO, brought a powerful as well as specific cellular and humoral immune responses in mice (152).
The main idea of choosing Ret to produce vaccine in ret related carcinomas fall in two criterion, first choosing patients self-antigens for cancer therapy with a non-mutated gene, second, there is no evidence of genetic mutations in Ret amino acids 64-269. Demonstration of proliferating hemangiomas, benign endothelial tumors and often referred as hemangiomas of infancy appearing at head or neck, express IDO and slowly regressed as a result of immune mediated process.
After large scale of genomic analysis show insulin like growth factor 2 as the key regulator of hematoma growth (Ritter et al. 2003). We set out to develop new technology with our previous expertise in immunotherapy and immunomodulation (181; 182; 183; 184), correcting Th17/Th1 ratio (185), and siRNA technology (186; 187). We developed siRNA-IDO-DCvax. Patented two technologies “Immunomodulation using Altered DCs (Patent No: US2006/0165665 A1) and Method of Cancer Treatments using siRNA Silencing (Patent No: US2009/0220582 A1).
In melanoma cancer DCs were preconditioned with whole tumor lysate but in breast cancer model pretreatment completed with tumor cell lysate before siRNA-IDO-DCvax applied. Both of these studies was a success without modifying the autanticity of DCs but decreasing the IDO expression to restore immunegenity by delaying tumor growth in breast cancer (147) and in melanoma (188). Thus, our DCvax specifically interfere with Ido without disturbing natural structure and content of the DCs in vivo showed that it is possible to carry on this technology to clinical applications.
Furthermore, our method of intervention is more sophisticated since it has a direct interaction mechanism with ex-vivo DC modulation without creating long term metabolism imbalance in Trp/Kyn metabolite mechanisms since the action is corrective and non-invasive.
There were several reasons.
First, prevention of tumor development studies targeting non-enzymatic pathway initiated by pDCs conditioned with TGFbeta is specific to IDO1 (189).
Second, IDO upregulation in antigen presenting cells allowing metastasis show that most human tumors express IDO at high levels (123; 124).
Third, tolerogenic DCs secretes several molecules some of them are transforming growth factor beta (TGFb), interleukin IL10), human leukocyte antigen G (HLA-G), and leukemia inhibitory factor (LIF), and non-secreted program cell death ligand 1 (PD-1 L) and IDO, indolamine 2.3-dioxygenase, which promote tumor tolerance. Thus, we took advantage of DCs properties and Ido specificity to prevent the tolerogenicity with siRNA-IDO DC vaccine in both melanoma and breast cancer.
Fourth, IDO expression in DCs make them even more potent against tumor antigens and create more T cells against tumors. IDOs are expressed at different levels by both in broad range of tumor cells and many subtypes of DCs including monocyte-derived DCs (10), plasmacytoid DCs (142), CD8a+ DCs (190), IDO compotent DCs (17), IFNgamma-activated DCs used in DC vaccination. These DCs suppress immune responses through several mechanisms for induction of apoptosis towards activated T cells (156) to mediate antigen-specific T cell anergy in vivo (142) and for enhancement of Treg cells production at sites of vaccination with IDO-positive DCs+ in human patients (142; 191; 192; 168; 193; 194). If DCs are preconditioned with tumor lysate with 1MT vaccination they increase DCvax effectiveness unlike DCs originated from “normal”, healthy lysate with 1MT in pancreatic cancer (195). As a result, we concluded that the immunesupressive effect of IDO can be reversed by siRNA because Treg cells enhances DC vaccine-mediated anti-tumor-immunity in cancer patients.
Gene silencing is a promising technology regardless of advantages simplicity for finding gene interaction mechanisms in vitro and disadvantages of the technology is utilizing the system with specificity in vivo (186; 196). siRNA technology is one of the newest solution for the treatment of diseases as human genomics is only producing about 25,000 genes by representing 1% of its genome. Thus, utilizing the RNA open the doors for more comprehensive and less invasive effects on interventions. Thus this technology is still improving and using adjuvants. Silencing of K-Ras inhibit the growth of tumors in human pancreatic cancers (197), silencing of beta-catenin in colon cancers causes tumor regression in mouse models (198), silencing of vascular endothelial growth factor (VGEF) decreased angiogenesis and inhibit tumor growth (199).
Combining siRNA IDO and DCvax from adult stem cell is a novel technology for regression of tumors in melanoma and breast cancers in vivo. Our data showed that IDO-siRNA reduced tumor derived T cell apoptosis and tumor derived inhibition of T cell proliferation. In addition, silencing IDO made DCs more potent against tumors since treated or pretreated animals showed a delay or decreased the tumor growth (188; 147)
Clinical Trials:
First FDA approved DC-based cancer therapies for treatment of hormone-refractory prostate cancer as autologous cellular immunotherapy (163; 164). However, there are many probabilities to iron out for a predictive outcome in patients.
Table 2 demonstrates the current summary of clinical trials report. This table shows 38 total studies specifically Ido related function on cancer (16), eye (3), surgery (2), women health (4), obesity (1), Cardiovascular (2), brain (1), kidney (1), bladder (1), sepsis shock (1), transplant (1), nervous system and behavioral studies (4), HIV (1) (Table 4). Among these only 22 of which active, recruiting or not yet started to recruit, and 17 completed and one terminated.
Most of these studies concentrated on cancer by the industry, Teva GTC ( Phase I traumatic brain injury) Astra Zeneca (Phase IV on efficacy of CRESTOR 5mg for cardiovascular health concern), Incyte corporation (Phase II ovarian cancer) NewLink Genetics Corporation Phase I breast/lung/melanoma/pancreatic solid tumors that is terminated; Phase II malignant melanoma recruiting, Phase II active, not recruiting metastatic breast cancer, Phase I/II metastatic melanoma, Phase I advanced malignancies) , HIV (Phase IV enrolling by invitation supported by Salix Corp-UC, San Francisco and HIV/AIDS Research Programs).
Many studies based on chemotherapy but there are few that use biological methods completed study with IDO vaccine peptide vaccination for Stage III-IV non-small-cell lung cancer patients (NCT01219348), observational study on effect of biological therapy on biomarkers in patients with untreated hepatitis C, metastasis melanoma, or Crohn disease by IFNalpha and chemical (ribavirin, ticilimumab (NCT00897312), polymorphisms of patients after 1MT drug application in treating patients with metastatic or unmovable refractory solid tumors by surgery (NCT00758537), IDO expression analysis on MSCs (NCT01668576), and not yet recruiting intervention with adenovirus-p53 transduced dendric cell vaccine , 1MT , radiation, Carbon C 11 aplha-methyltryptophan- (NCT01302821).
Among the registered clinical trials some of them are not interventional but observational and evaluation studies on Trp/Kyn ratio (NCT01042847), Kyn/Trp ratio (NCT01219348), Kyn levels (NCT00897312, NCT00573300), RT-PCR analysis for Kyn metabolism (NCT00573300, NCT00684736, NCT00758537), and intrinsic IDO expression of mesenchymal stem cells in lung transplant with percent inhibition of CD4+ and CD8+ T cell proliferation toward donor cells (NCT01668576), determining polymorphisms (NCT00426894). These clinical trials/studies are immensely valuable to understand the mechanism and route of intervention development with the data collected from human populations
Future Actions for Molecular Dx and Targeted Therapies:

Viable tumor environment. Tumor survival is dependent upon an exquisite interplay between the critical functions of stromal development and angiogenesis, local immune suppression and tumor tolerance, and paradoxical inflammation. TEMs: TIE-2 expressing monocytes; “M2” TAMs: tolerogenic tumor-associated macrophages; MDSCs: myeloid-derived suppressor cells; pDCs: plasmacytoid dendritic cells; co-stim.: co-stimulation; IDO: indoleamine 2,3-dioxygenase; VEGF: vascular endothelial growth factor; EGF: epidermal growth factor; MMP: matrix metaloprotease; IL: interleukin; TGF-β: transforming growth factor-beta; TLRs: toll-like receptors. (reference: http://www.hindawi.com/journals/cdi/2012/937253/fig1/)
Current survival or response rate is around 40 to 50 % range. By using specific cell type, selected inhibition/activation sequence based on patient’s genomic profile may improve the efficacy of clinical interventions on cancer treatments. Targeted therapies for specific gene regulation through signal transduction is necessary but there are few studies with genomics based approach.
On the other hand, there are surveys, observational or evaluations (listed in clinical trials section) registered with www.clinicaltrials.gov that will provide a valuable short-list of molecules. Preventing stimulation of Ido1 as well as Tgfb-1gene expression by modulating receptor mediated phosphorylation between TGFb/SMAD either at Mad-Homology 1 (MH1) or Mad-Homology 1 (MH2) domains maybe possible (79; 82; 80). Within Smads are the conserved Mad-Homology 1 (MH1) domain, which is a DNA binding module contains tightly bound Zinc atom.
Smad MH2 domain is well conserved and one the most diverse protein-signal interacting molecule during signal transduction due to two important Serine residues located extreme distal C-termini at Ser-Val-Ser in Smad 2 or at pSer-X-PSer in RSmads (80). Kyn activated orphan G protein–coupled receptor, GPR35 with unknown function with a distinct expression pattern that collides with IDO sites since its expression at high levels of the immune system and the gut (63) (200; 63).
The first study to connect IDO with cancer shows that group (75). The directly targeting to regulate IDO expression is another method through modulating ISREs in its promoter with RNA-peptide combination technology. Indirectly, IDO can be regulated through Bin1 gene expression control over IDO since Bin1 is a negative regulator of IDO and prevents IDO expression. IDO is under negative genetic control of Bin1, BAR adapter–encoding gene Bin1 (also known as Amphiphysin2). Bin1 functions in cancer suppression since attenuation of Bin1 observed in many human malignancies (141; 201; 202; 203; 204; 205; 206) . Null Bin-/- mice showed that when there is lack of Bin1, upregulation of IDO through STAT1- and NF-kB-dependent expression of IDO makes tumor cells to escape from T cell–dependent antitumor immunity.
This pathway lies in non-enzymatic signal transducer function of IDO after stimulation of DCs by TGFb1. The detail study on Bin1 gene by alternative spicing also provided that Bin1 is a tumor suppressor. Its activities also depends on these spliced outcome, such as Exon 10, in muscle, in turn Exon 13 in mice has importance in role for regulating growth when Bin1 is deleted or mutated C2C12 myoblasts interrupted due to its missing Myc, cyclinD1, or growth factor inhibiting genes like p21WAF1 (207; 208).
On the other hand alternative spliced Exon12A contributing brain cell differentiation (209; 210). Myc as a target at the junction between IDO gene interaction and Trp metabolism. Bin1 interacts with Myc either early-dependent on Myc or late-independent on Myc, when Myc is not present. This gene regulation also interfered by the long term signaling mechanism related to Kynurenine (Kyn) acting as an endogenous ligand to AHR in Trp metabolite and TGFb1 and/or IFNalpha and IFNbeta up regulation of DCs to induce IDO in noncanonical pathway for NF-kB and myc gene activations (73; 74). Hence, Trp/Kyn, Kyn/Trp, Th1/Th17 ratios are important to be observed in patients peripheral blood. These direct and indirect gene interactions place Bin1 to function in cell differentiation (211; 212; 205).

Table 3 contains the microarray analysis for Kyn affect showed that there are 25 genes affected by Kyn, two of which are upregulated and 23 of them downregulated (100). This list of genes and additional knowledge based on studies creating the diagnostics panel with these genes as a biomarker may help to analyze the outcomes of given interventions and therapies. Some of these molecules are great candidate to seek as an adjuvant or co-stimulation agents. These are myc, NfKB at IKKA, C2CD2, CREB3L2, GPR115, IL2, IL8, IL6, and IL1B, mir-376 RNA, NFKB3, TGFb, RelA, and SH3RF1. In addition, Lip, Fox3P, CTLA-4, Bin1, and IMPACT should be monitored.
In addition, Table 4 presents the other possible mechanisms. The highlights of possible target/biomarkers are specific TLRs, conserved sequences of IDO across its homologous structures, CCR6, CCR5, RORgammat, ISREs of IDO, Jak, STAT, IRFs, MH1 and MH2 domains of Smads. Endothelial cell coagulation activation mechanism and pDC maturation or immigration from lymph nodes to bloodstream should marry to control not only IDO expression but also genesis of preferred DC subsets. Stromal mesenchymal cells are also activated by these modulation at vascular system and interferes with metastasis of cancer. First, thrombin (human factor II) is a well regulated protein in coagulation hemostasis has a role in cell differentiation and angiogenesis.
Protein kinase activated receptors (PARs), type of GPCRs, moderate the actions. Second, during hematopoietic response endothelial cells produce hematopoietic growth factors (213; 214). Third, components of bone marrow stroma cells include monocytes, adipocytes, and mesenchymal stem cells (215). As a result, addressing this issue will prevent occurrence of coagulapathologies, namely DIC, bleeding, thrombosis, so that patients may also improve response rate towards therapies. Personal genomic profiles are powerful tool to improve efficacy in immunotherapies since there is an influence of age (young vs. adult), state of immune system (innate vs. adopted or acquired immunity). Table 5 includes some of the current studies directly with IDO and indirectly effecting its mechanisms via gene therapy, DNA vaccine, gene silencing and adjuvant applications as an intervention method to prevent various cancer types.
CONCLUSION
IDO has a confined function in immune system through complex interactions to maintain hemostasis of immune responses. The genesis of IDO stem from duplication of bacterial IDO-like genes. Inhibition of microbial infection and invasion by depleting tryptophan limits and kills the invader but during starvation of trp the host may pass the twilight zone since trp required by host’s T cells. Thus, the host cells in these small pockets adopt to new microenvironment with depleted trp and oxygen poor conditions. Hence, the cell metabolism differentiate to generate new cellular structure like nodules and tumors under the protection of constitutively expressed IDO in tumors, DCs and inhibited T cell proliferation.
On the other hand, having a dichotomy in IDO function can be a potential limiting factor that means is that IDOs impact on biological system could be variable based on several issues such as target cells, IDO’s capacity, pathologic state of the disease and conditions of the microenvironment. Thus, close monitoring is necessary to analyze the outcome to prevent conspiracies since previous studies generated paradoxical results.
Current therapies through chemotherapies, radiotherapies are costly and effectiveness shown that the clinical interventions require immunotherapies as well as coagulation and vascular biology manipulations for a higher efficacy and survival rate in cancer patients. Our siRNA and DC technologies based on stem cell modulation will provide at least prevention of cancer development and hopefully prevention in cancer.
11. References
1. Biochemistry of tryptophan in health and disease. BenderDA. 1983, Mol Aspects Med , pp. 6:101–197.
2. Molecular insights into substrate recognition and catalysis by indolamine 2,3-dioxygenase. Forouhar, F., Anderson, R., Mowat, C.F, et al. 2006, PNAS, pp. vol. 104, no:2, 473-478.
3. Importance of the Two Interferon-stimulated Response Element. Konan KV, Taylor, MW. 1996, J. Biol. Chem.-, pp. 19140-5.
4. Induction of indolamine 2,3 dioxygenase: A mechanism of the anti-tumor activity of interferon gamma. Ozaki, Y., Edelstein, M.P., Duch, D.S. 1998, PNAS USA., pp. vol:85, 1242-1246.
5. Localization of the human indoleamine 2,3-dioxygenase (IDO) gene to the pericentromeric region of human chromosome . Burkin, D. J., Kimbro, K. S., Barr, B. L., Jones, C., Taylor, M. W., Gupta, S. L. 1993, Genomics , pp. 17: 262-263.
6. Localization of indoleamine 2,3-dioxygenase gene (INDO) to chromosome 8p12-p11 by fluorescent in situ hybridization. Najfeld, V., Menninger, J., Muhleman, D., Comings, D. E., Gupta, S. L. 1993, Cytogenet. Cell Genet. , pp. 64: 231-232.
7. Molecular cloning, sequencing and expression of human interferon-gamma-inducible indoleamine 2,3-dioxygenase cDNA. Dai, W., Gupta, S. L. 1990, Biochem. Biophys. Res. Commun. , pp. 168: 1-8.
8. Gene structure of human indoleamine 2,3-dioxygenase. Kadoya, A., Tone, S., Maeda, H., Minatogawa, Y., Kido, R. 1992, Biochem. Biophys. Res. Commun. , pp. 189: 530-536.
9. A gene atlas of th emouse and human protein-encoding transcriptomes. Andrew I. Su, Tim Wiltshire, Serge Batalov , Hilmar Lapp , Keith A. Ching , David Block, Jie Zhang , Richard Soden , Mimi Hayakawa , Gabriel Kreiman , Michael P. Cooke , John R. Walker , and John B. Hogenesch. 2004, PNAS, pp. vol. 101, no. 166062-6067 (http://dx.doi.org:/10.1073/pnas.0400782101).
10. Indoleamine 2,3-dioxygenase production by human dendritic cells results in the inhibition of T cell proliferation. Hwu P, Du MX, Lapointe R, Do M, Taylor MW, Young HA. 2000, J. Immunol, pp. 164:3596–3599.
11. Inhibition of T cell proliferation by acrophage tryptophan catabolism. Munn, D.H. et al. 1999, J. Exp. Med., p. 189:1363.
12. HeLa cells cocultured with peripheral blood lymphocytes acquire an immuno-inhibitory phenotype through up-regulation of indoleamine 2,3-dioxygenase activity. Logan, G. J., Smyth, C. M. F., Earl, J. W., Zaikina, I., Rowe, P. B., Smythe, J. A., Alexander, I. E. 2002, Immunology, pp. 105:478-487.
13. Indoleamine 2,3-Dioxygenase – Is It an Immun Suppressor? Soliman H, Mediaville-Varela M, Antonia S. 2010, Cancer J. , pp. 16:354-359.
14. Targeting the immunoregulatory indoleamine 2,3-dioxygenase pathway in immunotherapy. Johnson BA, III, Baban B, Mellor AL. 2009, Immunotherapy. , pp. 645–661.
15. Indoleamine 2,3-dioxygenase and regulation of T cell immunity. AL., Mellor. 2005, Biochem Biophys Res Commun. , pp. 338(1):20–24.
16. Modulation of tryptophan catabolism by regulatory T cells. Fallarino, F., Grohmann, U., Hwang, K. W., Orabona, C., Vacca, C., Bianchi, R., Belladonna, M. L., Fioretti, M. C., Alegre, M.-L., Puccetti, P. 2003, Nature Immun., pp. 4: 1206-1212.
17. CTLA-4-Ig regulates tryptophan catabolism in vivo. Grohmann, U., Orabona, C., Fallarino, F., Vacca, C., Calcinaro, F., Falorni, A., Candeloro, P., Belladonna, M. L., Bianchi, R., Fioretti, M. C., Puccetti, P. 2002, Nature Immun. , pp. 3: 1097-1101.
18. Reverse signaling through GITR ligand enables dexamethasone to activate IDO in allergy. Grohmann, U., Volpi, C., Fallarino, F., Bozza, S., Bianchi, R., Vacca, C., Orabona, C., Belladonna, M. L., Ayroldi, E., Nocentini, G., Boon, L., Bistoni, F., Fioretti, M. C., Romani, L., Riccardi, C., Puccetti, P. 2007, Nature Med., pp. 13:579-586.
19. Cells expressing indoleamine 2,3-dioxygenase inhibit T cell responses. Mellor, A. L., Keskin, D. B., Johnson, T., Chandler, P., Munn, D. H. 2002, J. Immun. , pp. 168: 3771-3776.
20. Chon, SY, Hassanain, HH, Piine, R., and Gupta, SL. 1995, J. Interferon Cytokine Res. , pp. 15, 517-526.
21. Levy, ED, KEsler, DS, Pine, R., Reich, N, and Darnell, JE.Jr et al. 1988, Genes Dev, pp. 2,383-393.
22. Benoist, C. and Manthis, D. 1990, Annu. Rev of Immunol., pp. 8, 681-715.
23. Dorn, A, Durand, B., Marling, C., Meur, M.L., Beoist, C., and Mathis, D. 1987, PNAS USA, pp. 34, 6249-6253.
24. Konan, K.V. Ph.D. Thesis. Transcriptional Regulation of the Indolamine 2,3-oxygenase Gene. s.l. : Indiana University, Bloominigton, 1995.
25. Tryptophan pyrrolase of rabbit intestine: D- and L–tryptophan cleaving enzyme or enzymes. Yamamoto, S., and Hayashi, O. 1967, J Biol Chem, pp. 242: 5260-5266.
26. Prevention of allogeneic fetal rejection by tryptophan catabolism. Munn, DH, Zhou M, Attwood JT, Bondarev I, Conway SJ, Marshall B, Brown C, Mellor AL. 1998, Science, pp. 281:1191–3.
27. Evidence for a tumoral immune resistance mechanismbased on tryptophan degradation by indoleamine 2,3-dioxygenase. Uyttenhove, C. et al. 2003, Nature Med. 9, pp. 1269–1274 .
28. Pregnancy: success and failure within the Th1/Th2/Th3 paradigm. Raghupathy, R. 2001., Seminars in Immunology, pp. Volume 13, Issue 4, Pages 219–227.
29. Why is the fetal allograft not rejected? Davies, C. J. March 2007 , J ANIM SCI , pp. vol. 85 no. 13 suppl E32-E35 .
30. Exploring the mechanism of tryptoophan 2,3-dioxygenase. Thackray, S., Mowat, C.G., Chapman, K. 2008, Biochem. Society Transaction., pp. 36, 1120-1123.
31. The new life of a centenarian: signalling functions of NAD(P). Berger F, Ramírez-Hernández MH, Ziegler M. 2004, Trends Biochem Sci , pp. 29:111–118 .
32. Biochemistry of tryptophan in health and disease. DA, Bender. 1983, Mol Aspects Med, pp. 6:101–197. 33. Poliovirus induces indoleamine-2,3-dioxygenase and quinolinic acid synthesis in macaque brain. Heyes MP, Saito K, Jacobowitz D, Markey SP, Takikawa O, Vickers JH. 1992, FASEB J., pp. 6:2977–2989.
34. Dramatic changes in oxidative tryptophan metabolism along the kynurenine pathway in experimental cerebral and noncerebral malaria. . Sanni LA, Thomas SR, Tattam BN, Moore DE, Chaudhri G, Stocker R, Hunt NH. 1998, Am J Pathol, pp. 152:611–619.
35. Induction of pulmonary indoleamine 2,3-dioxygenase by intraperitoneal injection of bacterial lipopolysaccharide. . Yoshida R, Hayaishi O. 1978, Proc Natl Acad Sci USA , pp. 75:3998–4000.
36. Induction of indoleamine 2,3-dioxygenase in mouse lung during virus infection. Yoshida R, Urade Y, Tokuda M, Hayaishi O. 1979, Proc Natl Acad Sci USA , pp. 76:4084–4086.
37. Induction of pulmonary indoleamine 2,3-dioxygenase by intraperitoneal injection of bacterial lipopolysaccharide. Yoshida R, Hayaishi. 1978, PNAS USA, pp. 3998-4000.
38. Sequence of human 2,3-dioxygenase (TDO2): presence of a glucorticoid response-like element composed of a GTT repeat and intronic CCCCT repeat. Comings DE, Muhleman D, Dietz G, Sherman M, Forest. 1995, Genomics, pp. 29:390-396165.
39. Studies on the biosynthesis of Nicotinamide adenine inucleotide. II.Arole of picolinic carboxylase in the Biosynthesisofnicotinamideadeninedinucleotidefromtryptophan in mammals. Ikeda M, Tsuji H, Nakamura S, Ichiyama A, Nishizuka Y, HayaishiO. 1965, J. Biol. Chem. , pp. 240: 1395-1401.
40. The Secret Life of NAD+: An Old Metabolite Controlling New Metabolic Signaling Pathways. Houtkooper R.H., Carles Cantó C. , Wanders, R.J. and Auwerx, J. 2010, Endocrine Reviews , pp. vol. 31 no. 2 194-223, http://dx.doi.org:/10.1210/er.2009-0026.
41. Stimulation of Nicotinamide adenine dinucleotide biosynthetic pathways delays axonal degeneration after axotomy. Sasaki Y, Araki T, Milbrandt J. 2006, J Neurosci , pp. 26: 8484–8491.
42. European Nicotinamide Diabetes Intervention Trial (ENDIT): a randomised controlled trial of intervention before the onset of type 1 diabetes. Gale EA, Bingley PJ, Emmett CL, CollierT. 2004, Lancet., pp. 363:925–931.
43. Safety of high-dose nicotinamide: a review. Knip M, Douek IF, Moore WP, Gillmor HA, McLean AE, Bingley PJ, Gale EA. 2000, Diabetologia, pp. 43:1337–1345.
44. Large supplements of nicotinic acid and nicotinamide increase tissue NAD and poly(ADP-ribose) levels but do not affect diethylnitrosamine-induced altered hepatic foci in Fischer-344 rats. JacksonTM, Rawling JM, Roebuck BD, Kirkland JB. 1995, J Nutr , p. 125:1455.
45. Characterization and evolution of vertebrate indelamine 2,3-dihydrogenases IDOs from monotremes and marsupials. Yuasa, HJ, Ball, HJ, Ho, YF, Austin, CJ, et al. 2009, Comp. Biochem. Physiol. B. Biochem.. Mol. Biol., pp. 153 (2): 137-144.
46. Novel tryptophan catabolic enzyme IDO2 is the preferred biochemical target of the antitumor indolamine 2,3-dihydrogenase inhibitor compound D-1 methyl-tryptophan. Metz, R., Duhadaway, JB, Kamasani, U, Laury-Kleintop, L., Muller, AJ, Prendergast, GC. 2007, Cancer Res., pp. 67 (15): 7082-7087.
47. Total synthesis of exiguamines A and B inspired by catechollamine chemistry. Sofiyev, V, Lumb, JP, Volgraf, M., Trauner, D. 2012, Chemistry., pp. 18 (16): 4999-5005.
48. Molecular evolution of bacterial indolamine 2,3-dioxygenase. Yuasa, H J, Ushigoe, A, Ball, HJ. 2011, Gene., pp. 484 (1) : 22-31.
49. Infectious tolerance and the long-term acceptance of transplant tissue. Waldman, H., Adams, E., Fairchild, P., and Cobbold, S. 2006, J. Immunol., pp. 212:301-313.
50. Molecular evolution and characterizationof fungal indolamine 2,3-dioxygenases. Yuasa, HJ and Ball, HJ. 2012, J. Mol. Eval., pp. 72 (2): 160-168.
51. convergent evolution. The gene structure of Sulculus 41 kDa myoglobin is homologous with tht of human indolamine dioxygenase. Suzuki, T, Imai, K. 1996, Biochim. Biophys. Acta., pp. 1308(1):41-48.
52. Evolutionof myoglobin. Suzuki, T., Imai, K. 1998, Cell Mol Life Sci, pp. 54(9):979-1004.
53. A myoglobin evolved from indolamine 2,3-dioxygenase, trtptophan-degrading enzyme. Suzuki, T., Kawamichi, H., Imai, K. 1998, Comp Biochem Phisiol. Mol. Biol., pp. 121(2):117-128.
54. Do molluscs possess indolamine 2,3-dioxygenase? Yuasa, HJ and Suzuki, T. 2005, Comp. Biochem. Physiol. B. Biochem. Mol. Biol. , pp. (3) 445-454.
55. Comparison studies of the indolamine dioxygenase-like myoglobin from the abalone Sulculus diversicolor. Suzuki, T., Imai, K. 1997, Comp. Biohem. Phsiol B Biochem Mol Biol, pp. 117 (4)599-604.
56. Orchestration of the immune response by dendritic cells. Buckwalter MR, Albert ML. 2009, Curr Biol., pp. 19(9):355–361.
57. Dendritic cells and the control of immunity. Banchereau J, Steinman RM. 1998, Nature., pp. 245–52.
58. IDO expression by dendritic cells: tolerance and tryptophan catabolism. . Munn DH, Mellor AL. 2004, Nat Rev Immunol. , pp. 762–74.
59. Monocyte and Macrophage. Gordon, S. and Taylor, P.R. 2005, NATURE REVIEWS | IMMUNOLOGY , pp. vol:5, 953-964.
60. Blood monocytes consist of two principal subsets with distinct migratory properties. Geissmann F, Jung S, Littman DR. 2003, Immunity. , pp. 19:71–82.
61. Identification of a novel cell type in peripheral lymphoid organs of mice. I Morphology, quantitation, tissue distribution. . Steinman RM, Cohn ZA. 1973, J Exp Med., pp. 137(5):1142–1162.
62. T cell apoptosis by tryptophan catabolism. Fallarino F, Grohmann U, Vacca C, Bianchi R, Orabona C, Spreca A, Fioretti MC, Puccetti P. 2002, Cell Death Differ , pp. 9:1069–1077.
63. Kynurenine is a novel endothelium derived relaxing factor produced during inflammation. Wang, et al. 2010, Nat. Med., pp. 16(3): 279-285.
64. Activation of the noncanonical NF-kB pathway by HIV controls a Dendritic cell immunoregulatory phenotype. Manches, O. Fernandez, V.M.,, Plumas, J., Chaperot, L., and Bhardwaj, N. 2012, PNAS, pp. vol: 109, 14122-14127.
65. B cells inhibit induction of T cell-dependent tumor immunity. Qin, Z., Richter, G., Schuler, T., Ibe, S., Cao, X, Blakenstein, T. 1998, Nat. Med, p. 4:627.
66. Different partners, Opposite Outcmes: A new perspective of immunobiology of Indolamine 2,3 dioxygenase. Orabona, C., Pallotta, M.T., Grohman, U. 2012, Molecular Medicine., pp. 18:834-842.
67. Indolamine 2,3-dioxygenase: From catalyst to signaling function. Fallarino, F., Grohman, U., and Puccetti, P. 2012, Eurepean J. of Immunol. , pp. 42:1932-1937.
68. IDO: more than an enzyme. Chen, W. 2011, Nature Immonology, pp. 809-811.
69. Indolamine2,3-dehydrogenase in lung dendritic cells promotes Th2 responses and allergic inflammation. Xu, H., Oriss, T.B., Fei, M., Henry, A.C., Melgert, B.N., Chen, L., Mellor, A.L. 2008, PNAS USA, pp. 105: 6690-6695.
70. The immunoregulatory enzyme IDO paradoxically drives B-cellmediated autoimmunity. Scott, G.N., DuHadaway, J., Pigott, E., Ridge, N., Prendergast, G.C., Muller, A.J., Mandik-Nayak, L. 2009, J. Immunol., pp. 182:7509-7517.
71. Tryptophan deprivation sensitizes activated T cells to apoptosis prior to cell division. Lee GK, Park HJ, Macleod M, Chandler P, Munn DH, Mellor AL. 2002, Immunology , pp. 107:452–460.
72. Enzymology of NAD+ homeostasis in man. . Magni G, Amici A, Emanuelli M, Orsomando G, Raffaelli N, Ruggieri S. 2004, Cell Mol Life Sci , pp. 61:19–34.
73. Kynurenine pathway enzymes in dendritic cells initiate tolerogenesis in the absence of functional IDO. . Belladonna ML, Grohmann U, Guidetti P, Volpi C, Bianchi R, Fioretti MC, Schwarcz R, Fallarino F, Puccetti P. 2006, J Immunol. , pp. ;177:130–7.
74. An indogenous tumour promoting ligand of the human aryl hydrocarbon receptor. Opitz, et. al. 2011, pp. http://dx.doi.org:/10.1038/nature10491.
75. Inhibition of indoleamine 2,3-dioxygenase, animmunoregulatorytarget of the cancer suppression gene Bin1, potentiates cancer chemotherapy. Muller, A. J. et al. 2005, Nature Med. , pp. 11, 312–319 .
76. TGF-b; a master of all T cell trades. Li, M.O., Fravell, R.A. 2008, Cell. , pp. 134: 392-404.
77. Palotta, M.T. et al. 2011, Nat. Immunol., pp. 12:870-878. 78. Chen, W. et al. 2003, J. Exp. Immunol., p. 198: 1875.
79. Smads: transcriptional activators of TGF-beta responses. . Derynck R, Zhang Y, Feng XH. 1998, Cell , pp. 95 (6): 737–40.
http://dx.doi.org:/10.1016/S0092-8674(00)81696-7. PMID 9865691.
80. Smad transcription factors. Massagué J, Seoane J, Wotton D. 2005, Genes Dev, pp. 19 (23): 2783–810.
http://dx.doi.org:/10.1101/gad.1350705. PMID .
81. A structural basis for mutational inactivation of the tumour suppressor Smad4. Shi Y, Hata A, Lo RS, Massagué J, Pavletich NP. 1997, Nature., pp. 388 (6637): 87–93. http://dx.doi.org:/10.1038/40431. PMID 9214508.
82. Promoting bone morphogenetic protein signaling through negative regulation of inhibitory Smads. Itoh F, Asao H, Sugamura K, Heldin CH, ten Dijke P, Itoh S. 2001, EMBO J., pp. 20 (15): 4132– http://dx.doi.org:/10.1093/emboj/20.15.4132. PMC 149146. PMID 11483516.
83. SMAD_Signaling_Network. http://www.sabiosciences.com. [Online] 2013. http://www.sabiosciences.com/pathway.php?sn=SMAD_Signaling_Network.
84. Immune inhibitory receptors. Revetch, J.V., and Lanier, L.L. 2000, Science., pp. 290:84-89.
85. Soc3 drives proteasomal degradation of indolamine 2,3-dioxygenase (IDO) and antagonizes IDO-dependent tolerogenesis. Orabona, C., Pallotta, M., Volpi, C., et al. 2008, PNAS USA, pp. 105: 20828-20833.
86. Cutting edge; silencing supressor of cytokine signaling3 expression in dendritic cells turns CD28-Ig from immune adjuvant to supressant. Orabona, C.,, Belladonna, M.L., et all. 2005, J. Immunol., pp. 174: 6582-6586.
87. Molecular signatures of T-cell inhibition in HIV-1 infection. Larsson, M., Shankar. E.M, Che, K.F., Ellegard, R., Barathan, M., Velu, V., and Kamarulzaman, A. 2013, Retrovirology, p. 10:31.
88. TGF-beta and CD4+CD25+ regulatory cells. Huber, S. and Schramn, C. 2006, Front. Bioscie., pp. 11:1014-1023.
89. Immune Escape as a fundemental trait of cancer; focus on IDO. Prendergast, G.C. 2008, Oncogene., pp. 27, 3889-3900.
90. Il-6 inhibits the tolerogenic functionof CD8+ dendritic cells expressing indolamine 2,3-dioxygenase. Grohman, U., Fallarino, F., et al. 2001, J. Immunol., pp. 167:708-714.
91. Avoiding horror autotoxicus: Th eimportance of dentritic cells in peripheral T cell tolerance. Steinman, R.M., and Nussenzweig, M.C. 2002, PNAS, pp. no:1, 351-358.
92. Dendritic-cell function in Toll-like receptor- and MyD88-knockout mice . Kaisho, T., Akira, S. 2001, Trends Immunol , pp. 22,78-83.
93. Innate sensing of self and non-self RNAs by Toll-like receptors. Sioud, M. 2006., Trends Mol Med., pp. 12:67–76.
94. Impaired expression of indoleamine 2, 3-dioxygenase in monocyte-derived dendritic cells in response to Toll-like receptor-7/8 ligands. Furset, G., Fløisand, Y. and Sioud, M. 2008, Immunology., pp. 123(2): 263–271, http://dx.doi.org:/10.1111/j.1365-2567.2007.02695.x.
95. Toll-;ike receptor 9 mediated induction of the immunorepressor pathway of tryptophan metabolism. Fallarino, F., and Puccetti, P. 2006, Eur. J. of Imm., pp. 36:8-11.
96. Toll-like receptors and host defense against microbial pathogens: bringing specificity to the innate immune system. . Netea MG, der Graaf C, Van der Meer JWM, Kullberg BJ. 2004, J Leukoc Biol. , pp. 75:749–55.
97. Species-specific recognition of single-stranded RNA via toll-like receptor 7 and 8. . Heil F, Hemmi H, Hochrein H, et al. 2004, Science. , pp. 303:1526–9.
98. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. . Diebold SS, Kaisho T, Hemmi H, Akira S, Reis e Sousa C. 2004., Science. , pp. 303:1529–31.
99. The role of CpG motifs in innate immunity. Krieg, A.M. 2000., Curr Opin Immunol., pp. 12:35–43.
100. Anendogenous tumour-promoting ligand of the human aryl hydrocarbon receptor. Opitz, C.A., Litzenburger, U.M., Sahm, F., Ott,M., Tritschler, I., Trump, S. 2011, Nature, pp. vol 478; 197-203.
101. Impaired impression of Indolamine 2,3-deoxygenase in monocyte derived DCs in response to TLR-7/8. Furset, G., Floisand, Y., Sioud, M. 2007, Immunology, pp. 263-271.
102. Activationof the noncanonical NF-kB pathway by HIV controls a Dendritic cell immunoregulatory phenotype. Manches, O. Fernandez, V.M.,, Plumas, J., Chaperot, L., and Bhardwaj, N. 2012, PNAS, pp. vol: 109, 14122-14127.
103. Regulation of dendritic cell numbers and maturation by lipopolysaccharide in vivo . de Smedt, T., Pajak, B., Muraille, E., Lespagnard, L., Heinen, E., De Baetselier, P., Urbain, J., Leo, O., Moser, M. 1996, J. Exp. Med., pp. 184,1413-1424.
104. Subsets of dendritic cell precursors express different Toll-like receptors and respond to different microbial antigens . Kadowaki, N., Ho, S., Antonenko, S., de Waal Malefyt, R., Kastelein, R. A., Bazan, F., Liu, Y-J. 2001, J. Exp. Med., pp. 194,863-869 .
105. TRAF6 is a critical factor for dendritic cell maturation and development . Kobayashi, T., Walsh, P. T., Walsh, M. C., Speirs, K. M., Chiffoleau, E., King, C. G., Hancock, W. W., Caamano, J. H., Hunter, C. A., Scott, P., Turka, L. A., Choi, Y. 2003, Immunity , pp. 19,353-363 .
106. Activation of interferon regulatory factor-3 via toll-like receptor 3 and immunomodulatory functions detected in A549 lung epithelial cells exposed to misplaced U1-snRNA. Sadik CD, Bachmann M, Pfeilschifter J, Mühl H. 2009, Nucleic Acids Res. , pp. 37(15):5041-56. http://dx.doi.org:/10.1093/nar/gkp525. Epub 2009 Jun 18.
107. Triggering of the dsRNA sensors TLR3, MDA5, and RIG-I induces CD55 expression in synovial fibroblasts. Karpus ON, Heutinck KM, Wijnker PJ, Tak PP, Hamann J. 2012, PLoS One., p. 7(5):e35606. http://dx.doi.org:/10.1371/journal.pone.0035606. Epub 2012 May 10.
108. The structure of the TLR5-flagellin complex: a new mode of pathogen detection, conserved receptor dimerization for signaling. Lu J, Sun PD. 2012, Sci Signal., p. 5(216):pe11. http://dx.doi.org:/10.1126/scisignal.2002963.
109. Flagellin/Toll-like receptor 5 response was specifically attenuated by keratan sulfate disaccharide via decreased EGFR phosphorylation in normal human bronchial epithelial cells. Shirato K, Gao C, Ota F, Angata T, Shogomori H, Ohtsubo K, Yoshida K, Lepenies B, Taniguchi N. 2013, Biochem Biophys Res Commun., pp. doi:pii: S0006-291X(13)00779-1. http://dx.doi.org:/10.1016/j.bbrc.2013.05.009. [Epub ahead of print].
110. Differential induction of interleukin-10 and interleukin-12 in dendritic cells by microbial Toll-like receptor activators and skewing of T-cell cytokine profiles Infect. Qi, H., Denning, T. L., Soong, L. 2003, Immun. , pp. 71,3337-3342 .
111. Activation of Toll-like receptor 2 on human dendritic cells triggers induction of IL-12, but not IL-10 . Thoma-Uszynski, S., Kiertscher, S. M., Ochoa, M. T., Bouis, D. A., Norgard, M. V., Miyake, K., Godowski, P. J., Roth, M. D., Modlin, R. L. 2000, J. Immunol. , pp. 165,3804-3810.
112. Toll-like receptor 2 (TLR2) and TLR4 differentially activate human dendritic cells . Re, F., Strominger, J. L. 2001, J. Biol. Chem. , pp. 276,37692-37699.
113. Pasare, C., Medzhitov, R. (2003) Toll pathway-dependent blockade of CD4+CD25+ T cell-mediated suppression by dendritic cells. Pasare, C., Medzhitov, R. 2003, Science , pp. 299,1033-1036 .
114. What is the role of regulatory T cells in the success of implantation and early pregnancy? Saito, S., Shima, T., Nakashima, A., Shiozaki, A., Ito, M., Sasaki, Y. 2007, J Assist Reprod Genet, pp. 24: 379-386.
115. Sleeping Beauty-based gene therapy with indoleamine 2,3-dioxygenase inhibits lung allograft fibrosis. Liu H, Liu L, Fletcher BS, Visner GA. 2006, FASEB J, pp. 20:2384-2386.
116. Indoleamine 2,3-dioxygenase expression in transplanted NOD Islets prolongs graft survival after adoptive transfer of diabetogenic splenocytes. Alexander AM, Crawford M, Bertera S, et al. 2002, Diabetes. , pp. 51(2):356–365.
117. Solid Cancers after Bone Marrow Transplantatioin. Curtis, R.E., Rowlings, P.A., Deeg, J., Schirer, D.A. et al. 1997, The New England Journal of Medicine., pp. 336, No: 13: 897-904.
118. More ADO about IDO; GVHD (commentary). Curti, A., Trabanelli, S., Lemoli, M. 2008, Blood, p. 2950.
119. Jasperson, et al, . 2008, Blood, p. 3257.
120. Tolerance, DCs and tryptophan: much ado about IDO. Grohmann U, Fallarino F, Puccetti P. 2003, Trends Immunol, pp. 24:242-248.
121. Evidence for a tumoral immune resistance mechanism based on tryptophan degradation by indoleamine 2,3-dioxygenase. Uyttenhove C, Pilotte L, Théate I, Stroobant V, Colau D, Parmentier N, et al. 2003, Nat Med , pp. 9:1269–74.
122. Indoleamine 2,3-dioxygenase is a critical regulator of acute graft-versus-host disease lethality. Lisa K. Jasperson, Christoph Bucher, Angela Panoskaltsis-Mortari, Patricia A. Taylor, Andrew L. Mellor, David H. Munn, and Bruce R. Blazar. 2008., Blood., pp. 111:3257-3265.
123. The metabolism of tryptophan. 2. The metabolism of tryptophan in patients suffering from cancer of the bladder. . Boyland, E. & Willliams, D.C. 1956, Biochem. J., pp. 64, 578−582 .
124. Tryptophan metabolism in carcinoma of the breast. . Rose, D. 1967, Lancet , pp. 1, 239−241.
125. Inhibitors of indoleamine-2,3-dioxygenase for cancer therapy: can we see the wood for the trees? . Löb S, Königsrainer A, Rammensee HG, Opelz G, Terness P. 2009;, Nat Rev Cancer , pp. 9:445–52. http://dx.doi.org:/10.1158/1078-0432.CCR-11-1331.
126. The hallmarks of cancer. . Hanahan, D. & Weinberg, R.A. 2000., Cell., pp. 100, 57−70.
127. Indoleamine 2,3-Dioxygenase Expression in Human Cancers: Clinical and Immunologic Perspectives. Godin-Ethier, J., Hanafi,L.A., Piccirillo,C.A. and Lapointe, R. 2011, Clin Cancer Res, pp. 17; 6985, http://dx.doi.org:/10.1158/1078-0432.CCR-11-1331.
128. Dendritic cell modification as a route to inhibiting corneal graft rejection by the indirect pathway of allorecognition. Khan A, Fu H, Tan LA, Harper JE, Beutelspacher SC, Larkin DF, Lombardi G, McClure MO, George AJ. 2013, Eur J Immunol., pp. 43(3):734-46. http://dx.doi.org:/10.1002/eji.201242914. Epub 2013 Jan 18.
129. Possible role of the ‘IDO-AhR axis’ in maternal-foetal tolerance. . Hao K, Zhou Q, Chen W, Jia W, Zheng J, Kang J, Wang K, Duan T. 2013, Cell Biol Int., pp. 37(2):105-8. http://dx.doi.org:/10.1002/cbin.10023. Epub 2013 Jan 2.
130. Implication of indolamine 2,3 dioxygenase in the tolerance toward fetuses, tumors, and allografts. . Dürr S, Kindler V. 2013, J Leukoc Biol. , pp. 93(5):681-7.
http://dx.doi.org:/10.1189/jlb.0712347. Epub 2013 Jan 16.
131. Evidence for a tumoral immune resistance mechanism based on tryptophan degradation by indoleamine 2,3-dioxygenase. Uyttenhove C, Pilotte L, Théate I, Stroobant V, Colau D, Parmentier N, et al. 2003, Nat Med, pp. 9:1269–74.
132. NAturally arising CD4+ regulatory T cells for immunologic self-tolerance and negative control of immune responses. Sagaguchi, S. 2004, Annu. Rev. of Immunol., pp. 22: 531-562.
133. Regulatory T cells in transplantation tolerance. Wood, K.J., zZSakaguchi, S.,. 2003, Nat. Rev. Immunol., pp. 3; 199-210.
134. The cell awareness of paternal alloantigens during pregnancy. Tafuri, A., Alferink, J., Hammerling, G.J., Arnold, B. 1995, Science, pp. 270; 630-3.
135. Adenovirus mediated CTLA4Ig transgene therapy alleviates abortion by inhibiting spleen lymphocyte proliferation and regulating apoptosis in the feto-placental unit. Li W, Li B, Li S. 2013, J Reprod Immunol. , pp. 97(2):167-74.
136. A distinct tolerogenic subset of splenic IDO(+)CD11b(+) dendritic cells from orally tolerized mice is responsible for induction of systemic immune tolerance and suppression of collagen-induced arthritis. Park MJ, Park KS, Park HS, Cho ML, Hwang SY, Min SY, Park MK, Park SH, Kim HY. 2012, Cell Immunol. , pp. 278(1-2):45-54. http://dx.doi.org:/10.1016/j.cellimm.2012.06.009. Epub 2012 Jul 10.
137. Pharmacological targeting of IDO-mediated tolerance for treating autoimmune disease. Penberthy, W.T. 2007, Curr. Drug Metab., pp. 8:(3):245-266.
138. Indoleamine 2,3-dioxygenase expression in transplanted NOD Islets prolongs graft survival after adoptive transfer of diabetogenic splenocytes. Alexander AM, Crawford M, Bertera S, et al. 2002, Diabetes. , pp. 51(2):356–365.
139. Heme oxygenase-1 plays an important protective role in experimental autoimmune encephalomyelitis. . Liu Y, Zhu B, Luo L, Li P, Paty DW, Cynader MS. 2001., NeuroReport. , pp. 12(9):1841–1845.
140. Tumor vaccines in 2010: need for integration. Koos, D., Josephs, SF, Alexandrescu, DT et al. 2010, Cell Immunol, pp. 263: 138-147.
141. BIN1 is a novel MYC-interacting protein with features of a tumor suppressor. . Sakamuro, D., Elliott, K., Wechsler-Reya, R. & Prendergast, G.C. 1996, Nat. Genet. , pp. 14, 69−77.
142. Expression of Indolamine 2,3-dioxygenase by plasmacytoid dendritic cells in tumor draining nodes. Munn, S.H., Sharma, M.D., Hou, D., Baban, B. et al. 2004, J. Clin. Invest. , pp. 114: 280-290.
143. Indoleamine 2,3-Dioxygenase Expression in Human Cancers: Clinical and Immunologic Perspectives. Jessica Godin-Ethier, Laïla-Aïcha Hanafi, Ciriaco A. Piccirillo, and Réjean Lapointe. 2011 , Clin Cancer Res, pp. 17; 6985, http://dx.doi.org:/10.1158/1078-0432.CCR-11-1331.
144. Potential regulatory function of human dendritic cells expressing indoleamine 2,3-dioxygenase. . Munn, D.H. et al. 2002, Science 297, 1867−1870, pp. 297, 1867−1870 .
145. An HDAC inhibitor enhances cancer therapeutic efficiency of RNA polymerase III promoter-driven IDO shRNA. Yen MC, Weng TY, Chen YL, Lin CC, Chen CY, Wang CY, Chao HL, Chen CS, Lai MD. 2013, Cancer Gene Ther. , p. http://dx.doi.org:/10.1038/cgt.2013.27. [Epub ahead of print].
146. Systemic delivery of Salmonella typhimurium transformed with IDO shRNA enhances intratumoral vector colonization and suppresses tumor growth. Blache CA, Manuel ER, Kaltcheva TI, Wong AN, Ellenhorn JD, Blazar BR, Diamond DJ. 2012, Cancer Res. , pp. 72(24):6447-56.
http://dx.doi.org:/ZZ1158/0008-5472.CAN-12-0193. Epub 2012 Oct 22.
147. Silencing IDO in dendritic cells: a novel approach to enhance cancer immunotherapy in a murine breast cancer model. Zheng X, Koropatnick J, Chen D, Velenosi T, Ling H, Zhang X, Jiang N, Navarro B, Ichim TE, Urquhart B, Min W. 2013, Int J Cancer., pp.132(4):967-77. http://dx.doi.org:/10.1002/ijc.27710. Epub 2012 Jul 20.
148. Immunosuppressive CD14+HLA-DRlow/neg IDO+ myeloid cells in patients following allogeneic hematopoietic stem cell transplantation. Mougiakakos D, Jitschin R, von Bahr L, Poschke I, Gary R, Sundberg B, Gerbitz A, Ljungman P, Le Blanc K. 2013, Leukemia. , pp. 27(2):377-88.
http://dx.doi.org:/10.1038/leu.2012.215. Epub 2012 Jul 25.
149. Upregulated expression of indoleamine 2, 3-dioxygenase in primary breast cancer correlates with increase of infiltrated regulatory T cells in situ and lymph node metastasis. Yu J, Sun J, Wang SE, Li H, Cao S, Cong Y, Liu J, Ren X. 2011, Clin Dev Immunol. , p. 11:469135.
http://dx.doi.org:/10.1155/2011/469135. Epub 2011 Oct 24.
150. Skin delivery of short hairpin RNA of indoleamine 2,3 dioxygenase induces antitumor immunity against orthotopic and metastatic liver cancer. Huang TT, Yen MC, Lin CC, Weng TY, Chen YL, Lin CM, Lai MD. 2011, Cancer Sci. , pp. 102(12):2214-20. http://dx.doi.org:/10.1111/j.1349-7006.2011.02094.x.
151. Indoleamine 2,3-dioxygenase expression in transplanted NOD Islets prolongs graft survival after adoptive transfer of diabetogenic splenocytes. . Alexander AM, Crawford M, Bertera S, et al. 2002, Diabetes. , pp. 51(2):356–365.
152. Prevention of Spontaneous Tumor Development in a ret Transgenic Mouse Model by Ret Peptide Vaccination with Indoleamine 2,3-Dioxygenase Inhibitor 1-Methyl Tryptophan. Zeng, J., Cai, S., Yi, Y., et al. 2009, Cancer Res., pp. 69: 3963-3970, http://dx.doi.org:/10.1158/0008-5472.CAN-08-2476.
153. Medicinal electronomics bricolage design of hypoxia-targeting antineoplastic drugs and invention of boron tracedrugs as innovative future-architectural drugs. Hori H, Uto Y, Nakata E. 2010, Anticancer Res. , pp. 30(9):3233-42.
154. Synthesis of 4-cyano and 4-nitrophenyl 1,6-dithio-D-manno-, L-ido- and D-glucoseptanosides possessing antithrombotic activity. Bozó E, Gáti T, Demeter A, Kuszmann J. 2002, Carbohydr Res. , pp. 3;337(15):1351-65.
155. Radiopharmaceuticals XXVII. 18F-labeled 2-deoxy-2-fluoro-d-glucose as a radiopharmaceutical for measuring regional myocardial glucose metabolism in vivo: tissue distribution and imaging studies in animals. Gallagher BM, Ansari A, Atkins H, Casella V, Christman DR, Fowler JS, Ido T, MacGregor RR, Som P, Wan CN, Wolf AP, Kuhl DE, Reivich M. 1977, J Nucl Med. , pp. 18(10):990-6.
156. Tryptophan deprivation sensitizes activated T cells to apoptosis prior to cell division. Lee GK, Park HJ, Macleod M, Chandler P, Munn DH, Mellor AL. 2002, Immunology, pp. 107:452–460.
157. Induction of indoleamine 2,3-dioxygenase by uropathogenic bacteria attenuates innate responses to epithelial infection. Loughman JA, Hunstad DA. 2012 , J Infect Dis. , pp. 205(12):1830-9. http://dx.doi.org:/10.1093/infdis/jis280.
158. Inhibition of allogeneic T cell proliferation by indoleamine 2,3-dioxygenase-expressing dendritic cells: mediation of suppression by tryptophan metabolites. . Terness, P., et al. 2002, J. Exp. Med.196:447–457., pp. 196:447–457.
159. The tryptophan catabolite L-kynurenine inhibits the surface expression of NKp46- and NKG2D-activating receptors and regulates NK-cell function. . Chiesa, M.D., et al. 2006, Blood. , pp. 108:4118–4125.38.
160. Differential effects of the tryptophan metabolite 3-hydroxyanthranilic acid on the proliferation of human CD8+ T cells induced by TCR triggering or homeostatic cytokines. Weber, W.P., et al. 2006, Eur. J. Immunol. , pp. 36:296-304.
161. Dendritic cell vaccination against ovarian cancer–tipping the Treg/TH17 balance to therapeutic advantage? Cannon MJ, Goyne H, Stone PJ, Chiriva-Internati M. 2011, Expert Opin Biol Ther. , pp. 11(4):441-5. http://dx.doi.org:/10.1517/14712598.2011.554812.
162. Phenotype, distribution, generation, and functional and clinical relevance of Th17 cells in the human tumor environments. . Kryczek I, Banerjee M, Cheng P, et al. 2009, Blood., pp. 114:1141–1149.
163. The use of dendritic cells in cancer immunitherapy. Schuler, G., Schuker-Turner, B., Steinman, RM, 2003, Curr. Opin. Immunol., pp. 15: 138-147.
164. Clinical applications of dentritic cell vaccines. Morse, MA, Lyerly, HK. 2000, Curr. Opin. Mol Ther., pp. 2:20-28.
165. Vaccination of melanoma patients with peptide or tumor lysate-pulsed dendritic cells. Nestle, FO, Alijagic, S., Gillet, M. et al. 1998, Nat. Med., pp. 4: 328-332.
166. Dentritic cell based tumor vaccination in prostate and renal cell cancer: a systamatic review. Draube, A., Klein-Gonzales, Matheus, S et al. 2011, Plos One, p. 6:e1881.
167. [Online] http://www.fda.gov/BiologicsBloodVaccines/CellularGeneTherapy-Products/ApprovedProducts/ucm210215.htm.
168. Dendritic cell based antitumor vaccination: impact of functional indolamine 2,3-dioxygenase expression. Wobster, m., Voigt, H., Houben, R. et al. 2007, Cancer Immunol Immunother, pp. 56:1017-1024. 169. [Online] oncoimmunology.2012 October1; 1(17):1111-1134, http://dx.doi.org:/10.4161/onci.21494.
170. Interleukins 1beta and 6 but not transforming growth factor-beta are essential for the differentiation of interleukin 17-producing human T helper cells. Acosta-Rodriguez EV, Napolitani G, Lanzavecchia A, Sallusto F. 2007 , Nat Immunol. , pp. 8(9):942-9.
171. IFNgamma promotes generationof Il-10 secreting CD4+ T cells that suppress generationof CD8responses in an antigen-experienced host. Liu, X.S., Leerberg, J., MacDonald, K., Leggatt, G.R., Frazer, I.H. 2009, J. Immunol., pp. 183: 51-58.
172. Antigen, in the presence of TGF-beta, induces up-regulationof FoxP3gfp+ in CD4+ TCR transgenic T cells that mediate linked supressionof CD8+ T cell responses. . Kapp, J.A., Honjo, K., Kapp, L.M., Goldsmith, K., Bucy, R.P. 2007, J. Immunol., pp. 179: 2105-2114.
173. Opposing effects of TGF-beta and IL-15 cytokines control the number of short lived effecctor CD8+ T cells. Sanjabi, S, Mosaheb, M.M., Flavell, R.A. 2009, Immunity., pp. 31; 131-144.
174. Synergestic enhancement of CD8+ T cell mediated tumor vaccines efficacy by an anti-tumor forming growth factor-beta monoclonal antibody. . Terabe, M., Ambrosino, E., Takaku, S. et al. 2009, Clin. Cancer Res., pp. 15; 6560-9.
175. IL-12 enhances CTL synapse formationand induces self-reactivity. Markinewicz, MA, Wise, EL, Buchwald, ZS et al. 2009, J. Immunol., pp. 182: 1351-1362.
176. Tumor specific Th17-polarized cells eradicate large established melanoma. Muranski, P., Boni, A., Antony, PA, et al. 2008, Blood, pp. 112; 362-373.
177. Type17 CD8+ T cells dispplay enhanced antitumor immunity. Hinrichs, C.S., Kaiser, A., Paulos, C.M., et al. 2008, Blood., pp. 112:362-373.
178. Marying Immunotherapy with Chemotherapy: Why Say IDO? Muller, AJ, and Prendergrast, GC. 2005, Cancer Research, pp. 65: 8065-8068.
179. Enhancing Cancer Vaccine efficacy via Modulationof the Tumor Environment. Disis, ML. 2009, Clin Cancer Res, pp. 15: 6476-6478.
180. Systemic inhibition of transforming growth factor beta 1 in glioma bearing mice improves the therapeutic efficacy of glioma-associated antigen peptide vaccines. Ueda, R., Fujita, M., Zhu, X., et al. 2009, Clin. Cancer res., pp. 15: 6551-9.
181. Immune modulation by silencing IL-12 productionin dendritic cells using smal interfering RNA. Hill, JA, Ichim, TE, Kusznieruk, KP, et al. 2003, J. Immunol, pp. 171:809-813.
182. Immune modulation and tolerance induction by RelB-silenced dentritic cells through RNA interference. Li, M. Zang, X, Zheng, X, et al. 2007, J. Immunol, pp. 178: 5480-7.
183. RNAi mediated CD40-CD54 interruption promotes tolerance in autoimmune arthritis. . Zheng, X., Suzuki, M., Zhang, X., et al. 2010, Arthritis Res. Ther., p. 12:R13.
184. Dendritic cells genetically engineered to express Fas ligand induce donor-specific hyporesponsiveness and prolong allograft survival. Min, WP. Gorczynki, R., huang, XY et al. 2000, J. Immunol., pp. 164:161-167.
185. LF15-0195 generates tolerogenic dendritic cells by supressionof NF-kappaB signaling through inhibitionof IKK activity. . Yang, J., Bernier, SM, Ichim, TE, et al. 2003, J Leukoc. Biol., pp. 74: 438-447.
186. RNA interfrence: A potent tool for gene specific therapeutics. . Ichim, TE, Li, M., Qian, H., Popov, HI, Rycerz, K., Zheng, X., White, D., Zhong, R., and Min, WP. 2004, Am. J. Transplant, pp. 4:1227-1236.
187. A novel in vivo siRNA delivery system specifically targeting dendritic cells and silencing CD40 genes for immunomodulation. Zheng, X., Vladau, C., Zhang, X. et al. 2009, Blood, pp. 113:2646-2654.
188. Reinstalling Antitumor Immunity by Inhibiting Tumor derived ImmunoSupressive Molecule IDO through RNA interference. Zheng, X et al. 2006, Int. Journal of Immunology., pp. 177:5639-5646.
189. Roles of TGFbeta in metastasis. Padua, D., Massague, J. 2009, Cell Res., pp. 19;89-102.
190. Functional expression of indolamine2,3-dioxygenase by murine CDalpha+dendritic cells. Fallarino, F., Vacca, C, Orabona, C et al. 2002, Int Immunol., pp. 14:65-8.
191. Indolamine2,3-dioxygenase controls conversion of Fox3+ Tregs to TH17-like cells in tumor draining lymph nodes. Sharma, MD, Hou, DY, Liu, Y et al. 2009, Blood, pp.113: 6102-11.
192. IDO upregulates regulatory T cells via tryptoophan catabolite and supresses encephalitogenic T cell responses in experimental autoimmune encephalomyelitis. Yan, Y, Zhang, GX, Gran, B et al. 2010, J Immunol, pp. 185; 5953-61.
193. IDO activates regulatory T cells and blocks their conversion into Th-17-like T cells. Baban, B, Chandler, PR, Sharma, MD et al. 2009, J Immunol, pp. 183; 2475-83.
194. Enhancement of vaccine-mediated antitumor immunity in cancer patients after depletionof regulatory T cells. Dannull, J., Farrand, KJ, Mathews, SA, et al. 2005, J Clin Invest, pp. 115: 3623-33.
195. 1-MT enhances potency of tumor cell lysate pulled dentritic cells against pancreatic adenocarcinoma by downregulating percentage of Tregs. Li, Y, Xu, J, Zhou, H. et al. 2010, J Huazhong Univ Sci Technol Med Sci , pp. 30: 344-8.
196. siRNA mediated antitumorigenesis for drug target validation and therapeutics. Lu, PY, Xie, FY and Woodle, MC. 2003, Curr Opin Mol. Ther., pp. 5:225-234.
197. Stable supression of tumorigenicity by virus-mediated RNA interference. Brumellkamp, TR, Bernards, R, Agami, R. 2002, Cancer Cell, pp. 2; 243-247.
198. Small interferring RNAs directed against beta-catenin inhibit the in vitro and in vivo growth of colon cancer cells. Verma, UN, Surabhi, RM, Schmaltieg, A., Becerra, C., Gaynor, RB. 2003, Clin. Cancer. Res., pp. 9:1291-1300.
199. siRNA mediated inhibition of vascular endothelial growth factor severely limits tumor resistance to antiangiogeneic thromboposdin-1 and slows tumor vascularization and growth. Filleur, S., Courtin, A, Ait-Si-Ali, S., Guglielmi, J., Merel, C., Harel-Bellan, A., CLezardin, P., and Cabon, F. 2003, Cancer Res, pp. 63; 3919-3922.
200. Kynurenic acid as a ligand for orphan G protein-coupled receptor GPR35. . Wang, J., et al. 2006, J. Biol.Chem. , pp. 281:22021–22028. 201. Bin1 functionally interacts with Myc in cells and inhibits cell proliferation by multiple mechanisms. Elliott, K. et al. 1999, Oncogene , pp. 18, 3564−3573 .
202. Mechanism for elimination of a tumor suppressor: aberrant splicing of a brain-specific exon causes loss of function of Bin1 in melanoma. . Ge, K. et al. 1999, Proc. Natl. Acad. Sci. USA, pp. 96, 9689−9694.
203. Losses of the tumor suppressor Bin1 in breast carcinoma are frequent and reflect deficits in a programmed cell death capacity. Ge, K. et al. 2000, Int. J. Cancer , pp. 85, 376−383.
204. Loss of heterozygosity and tumor suppressor activity of Bin1 in prostate carcinoma. Ge, K. et al. 2000, Int. J. Cancer , pp. 86, 155−161.
205. Expression of a MYCN-interacting isoform of the tumor suppressor BIN1 is reduced in neuroblastomas with unfavorable biological features. . Tajiri, T. et al. 2003, Clin. Cancer Res., pp. 9, 3345−3355.
206. Targeted deletion of the suppressor gene Bin1/Amphiphysin2 enhances the malignant character of transformed cells. Muller, A.J., DuHadaway, J.B., Donover, P.S., Sutanto-Ward, E. & Prendergast, G.C. 2004, Cancer Biol. Ther. , p. 3.
207. Interactions of myogenic factors and the retinoblastoma protein mediates muscle commitment and cell differentiation. Gu, WJ., Scheniider,W., Condrolli,G., Kaushal,, S, Mahdavi,V., Nadal-Gnard, B. 1993, Cell, pp. 72; 309-324.
208. Structural analysis of the human BIN1 gene: evidence of tissue-specific transcriptional regualtion and alternate splicing. Wechsler-Reya, R, Sakamuro, J., Zhang, J., DuHadaway, J., and Predengast. 1998, J of Biol Chem.
209. A role for th ePutative Tuimor Supressor Bin1 in Muscle Differentiation. Wechsler-Reya, R., Elliott, KJ, Prendergast, GC. 1998, Molecular and Cellular Biology, p. 18 (1) :566.
210. The putative tumor repressor BIN1 is a short lived nuclear phosphoprotein whose localization is altered in malignant cells. Wechsler-Reya, R., Elliot, K., Herlyn, M., Prendergast, GC. 1997, Cancer Res, pp. 57: 3258-3263.
211. Transformation selective apoptosis by farnesyltransferase inhibitors requires Bin1. DuHadaway, J.B. et al. 2003, Oncogene, pp. 22, 3578−3588 (2003).
212. The c-Myc-interacting adapter protein Bin1 activates a caspase-independent cell death program. Elliott, K., Ge, K., Du, W. & Prendergast, G.C. 2000., Oncogene , pp. 19, 4669−4684.
213. Growth stimulation of human bone marrow cells in agar culture by vascular cells. Knudtzon, S., and Mortensen, BT. 1975, Blood, pp. 46 (6) 937-943.
214. Exogenous endothelial cells as accelerators of hematopoietic reconstitution. Mizer, C., Ichim, TE, Alexandrescu, DT, DAsanu, CA, Ramos, F., Turner, A., Woods, EJ, Bogon, V., Murphy, MP, Koos, D., and Patel, A. 2013, J. Translational Medicine, p. 10: 231.
215. Dissecting the bone marrow microenvironment . Torok-Storb, B. et al. 1999, Annals of New York Academy of Science, pp. 872: 164-170. 217. Yuasa, XX and Ball YY. 2011.
218. Possible role of the ‘IDO-AhR axis’ in maternal-foetal tolerance. Hao K, Zhou Q, Chen W, Jia W, Zheng J, Kang J, Wang K, Duan T. 2013, Cell Biol Int. , pp. 37(2):105-8. http://dx.doi.org:/10.1002/cbin.10023.
219. Toll pathway-dependent blockade of CD4+CD25+ T cell-mediated suppression by dendritic cells. Pasare, C., Medzhitov, R. 2003, Science , pp. 299,1033-1036 .
220. Activation of Toll-like receptor 2 on human dendritic cells triggers induction of IL-12, but not IL-10. Thoma-Uszynski, S., Kiertscher, S. M., Ochoa, M. T., Bouis, D. A., Norgard, M. V., Miyake, K., Godowski, P. J., Roth, M. D., Modlin, R. L. 2000, J. Immunol. , pp. 165,3804-3810.
Like this:
Like Loading...
Read Full Post »