Inadequacy of EHRs
Larry H. Bernstein, MD, FCAP, Curator
LPBI
EHRs need better workflows, less ‘chartjunk’
By Marla Durben Hirsch
Electronic health records currently handle data poorly and should be enhanced to better collect, display and use it to support clinical care, according to a new study published in JMIR Medical Informatics.
The authors, from Beth Israel Deaconess Medical Center and elsewhere, state that the next generation of EHRs need to improve workflow, clinical decision-making and clinical notes. They decry some of the problems with existing EHRs, including data that is not displayed well, under-networked, underutilized and wasted. The lack of available data causes errors, creates inefficiencies and increases costs. Data is also “thoughtlessly carried forward or copied and pasted into the current note” creating “chartjunk,” the researchers say.
They suggest ways that future EHRs can be improved, including:
- Integrating bedside and telemetry monitoring systems with EHRs to provide data analytics that could support real time clinical assessments
- Formulating notes in real-time using structured data and natural language processing on the free text being entered
- Formulating treatment plans using information in the EHR plus a review of population data bases to identify similar patients, their treatments and outcomes
- Creating a more “intelligent” design that capitalizes on the note writing process as well as the contents of the note.
“We have begun to recognize the power of data in other domains and are beginning to apply it to the clinical space, applying digitization as a necessary but insufficient tool for this purpose,” the researchers say. “The vast amount of information and clinical choices demands that we provide better supports for making decisions and effectively documenting them.”
Many have pointed out the flaws in current EHR design that impede the optimum use of data and hinder workflow. Researchers have suggested that EHRs can be part of a learning health system to better capture and use data to improve clinical practice, create new evidence, educate, and support research efforts.
Disrupting Electronic Health Records Systems: The Next Generation
1Beth Israel Deaconess Medical Center, Division of Pulmonary, Critical Care, and Sleep Medicine, Boston, MA, US; 2Yale University, Yale-New Haven Hospital, Department of Pulmonary and Critical Care, New Haven, CT, US; 3Center for Urban Science and Progress, New York University, New York, NY, US; 4Center for Wireless Health, Departments of Anesthesiology and Neurological Surgery, University of Virginia, Charlottesville, VA, US
*these authors contributed equally
JMIR 23.10.15 Vol 3, No 4 (2015): Oct-Dec
Celi LA, Marshall JD, Lai Y, Stone DJ. Disrupting Electronic Health Records Systems: The Next Generation. JMIR Med Inform 2015;3(4):e34 DOI: 10.2196/medinform.4192 PMID: 26500106
Weed introduced the “Subjective, Objective, Assessment, and Plan” (SOAP) note in the late 1960s []. This note entails a high-level structure that supports the thought process that goes into decision-making: subjective data followed by ostensibly more reliable objective data employed to formulate an assessment and subsequent plan. The flow of information has not fundamentally changed since that time, but the complexities of the information, possible assessments, and therapeutic options certainly have greatly expanded. Clinicians have not heretofore created anything like an optimal data system for medicine [,]. Such a system is essential to streamline workflow and support decision-making rather than adding to the time and frustration of documentation [].
What this optimal data system offers is not a radical departure from the traditional thought processes that go into the production of a thoughtful and useful note. However, in the current early stage digitized medical system, it is still incumbent on the decision maker/note creator to capture the relevant priors, and to some extent, digitally scramble to collect all the necessary updates. The capture of these priors is a particular challenge in an era where care is more frequently turned over among different caregivers than ever before. Finally, based on a familiarity of the disease pathophysiology, the medical literature and evidence-based medicine (EBM) resources, the user is tasked with creating an optimal plan based on that assessment. In this so-called digital age, the amount of memorization, search, and assembly can be minimized and positively supported by a well-engineered system purposefully designed to assist clinicians in note creation and, in the process, decision-making.
Since 2006, use of electronic health records (EHRs) by US physicians increased by over 160% with 78% of office-based physicians and 59% of hospitals having adopted an EHR by 2013 [,]. With implementation of federal incentive programs, a majority of EHRs were required to have some form of built-in clinical decision support tools by the end of 2012 with further requirements mandated as the Affordable Care Act (ACA) rolls out []. These requirements recognize the growing importance of standardization and systematization of clinical decision-making in the context of the rapidly changing, growing, and advancing field of medical knowledge. There are already EHRs and other technologies that exist, and some that are being implemented, that integrate clinical decision support into their functionality, but a more intelligent and supportive system can be designed that capitalizes on the note writing process itself. We should strive to optimize the note creation process as well as the contents of the note in order to best facilitate communication and care coordination. The following sections characterize the elements and functions of this decision support system ().
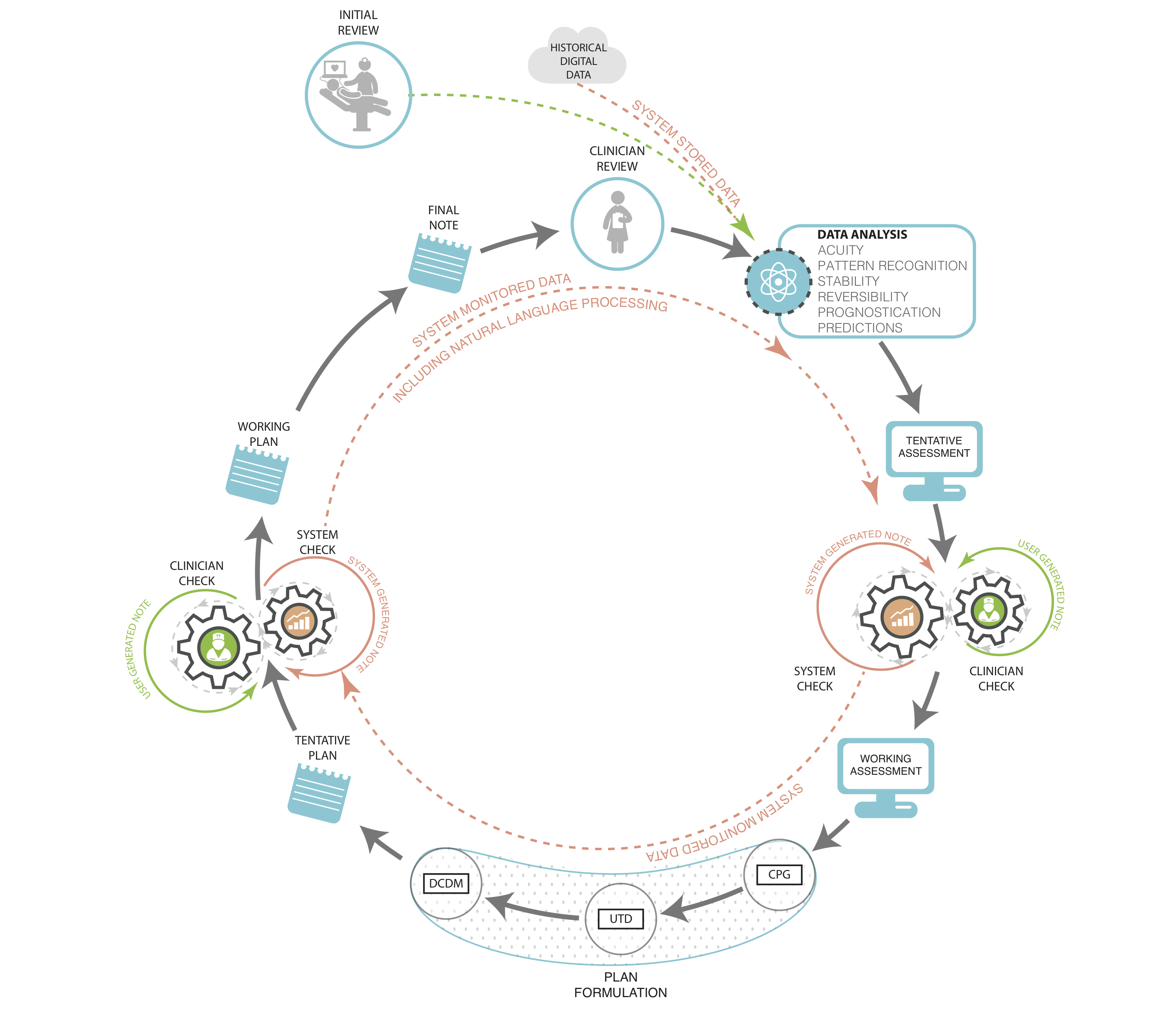
http://medinform.jmir.org/article/viewFile/4192/1/68490
Figure 1. Clinician documentation with fully integrated data systems support. Prior notes and data are input for the following note and decisions. Machine analyzes input and displays suggested diagnoses and problem list, and test and treatment recommendations based on various levels of evidence: CPG – clinical practice guidelines, UTD – Up to Date®, DCDM – Dynamic Clinical Data Mining.View this figure
Incorporating Data
Overwhelmingly, the most important characteristic of the electronic note is its potential for the creation and reception of what we term “bidirectional data streams” to inform both decision-making and research. By bidirectional data exchange, we mean that electronic notes have the potential to provide data streams to the entirety of the EHR database and vice versa. The data from the note can be recorded, stored, accessed, retrieved, and mined for a variety of real-time and future uses. This process should be an automatic and intrinsic property of clinical information systems. The incoming data stream is currently produced by the data that is slated for import into the note according to the software requirements of the application and the locally available interfaces []. The provision of information from the note to the system has both short- and long-term benefits: in the short term, this information provides essential elements for functions such as benchmarking and quality reporting; and in the long term, the information provides the afferent arm of the learning system that will identify individualized best practices that can be applied to individual patients in future formulations of plans.
Current patient data should include all the electronically interfaced elements that are available and pertinent. In addition to the usual elements that may be imported into notes (eg, laboratory results and current medications), the data should include the immediate prior diagnoses and treatment items, so far as available (especially an issue for the first note in a care sequence such as in the ICU), the active problem list, as well as other updates such as imaging, other kinds of testing, and consultant input. Patient input data should be included after verification (eg, updated reviews of systems, allergies, actual medications being taken, past medical history, family history, substance use, social/travel history, and medical diary that may include data from medical devices). These data priors provide a starting point that is particularly critical for those note writers who are not especially (or at all) familiar with the patient. They represent historical (and yet dynamic) evidence intended to inform decision-making rather than “text” to be thoughtlessly carried forward or copied and pasted into the current note.
Although the amount and types of data collected are extremely important, how it is used and displayed are paramount. Many historical elements of note writing are inexcusably costly in terms of clinician time and effort when viewed at a level throughout the entire health care system. Redundant items such as laboratory results and copy-and-pasted nursing flow sheet data introduce a variety of “chartjunk” that clutters documentation and makes the identification of truly important information more difficult and potentially even introduces errors that are then propagated throughout the chart [,]. Electronic systems are poised to automatically capture the salient components of care so far as these values are interfaced into the system and can even generate an active problem list for the providers. With significant amounts of free text and “unstructured data” being entered, EHRs will need to incorporate more sophisticated processes such as natural language processing and machine learning to provide accurate interpretation of text entered by a variety of different users, from different sources, and in different formats, and then translated into structured data that can be analyzed by the system.
Optimally, a fully functional EHR would be able to provide useful predictive data analytics including the identification of patterns that characterize a patient’s normal physiologic state (thereby enabling detection of significant change from that state), as well as mapping of the predicted clinical trajectory, such as prognosis of patients with sepsis under a number of different clinical scenarios, and with the ability to suggest potential interventions to improve morbidity or mortality []. Genomic and other “-omic” information will eventually be useful in categorizing certain findings on the basis of individual susceptibilities to various clinical problems such as sepsis, auto-immune disease, and cancer, and in individualizing diagnostic and treatment recommendations. In addition, an embedded data analytic function will be able to recognize a constellation of relatively subtle changes that are difficult or impossible to detect, especially in the presence of chronic co-morbidities (eg, changes consistent with pulmonary embolism, which can be a subtle and difficult diagnosis in the presence of long standing heart and/or lung disease) [,].
The data presentation section must be thoughtfully displayed so that the user is not overwhelmed, but is still aware of what elements are available, and directed to those aspects that are most important. The user then has the tools at hand to construct the truly cognitive sections of the note: the assessment and plan. Data should be displayed in a fashion that efficiently and effectively provides a maximally informationally rich and minimally distracting graphic display. The fundamental principle should result in a thoughtfully planned data display created on the ethos of “just enough and no more,” as well as the incorporation of clinical elements such as severity, acuity, stability, and reversibility. In addition to the now classic teachings of Edward Tufte in this regard, a number of new data artists have entered the field []. There is room for much innovation and improvement in this area, as medicine transitions from paper to a digital format that provides enormous potential and capability for new types of displays.
Integrating the Monitors
Bedside and telemetry monitoring systems have become an element of the clinical information system but they do not yet interact with the EHR in a bidirectional fashion to provide decision support. In addition to the raw data elements, the monitors can provide data analytics that could support real-time clinical assessment as well as material for predictive purposes apart from the traditional noisy alarms [,]. It may be less apparent how the reverse stream (EHR to bedside monitor) would work, but the EHR can set the context for the interpretation of raw physiologic signals based on previously digitally captured vital signs, patient co-morbidities and current medications, as well as the acute clinical context.
In addition, the display could provide an indication of whether technically ”out of normal range” vital signs (or labs in the emergency screen described below) are actually “abnormal” for this particular patient. For example, a particular type of laboratory value for a patient may have been chronically out of normal range and not represent a change requiring acute investigation and/or treatment. This might be accomplished by displaying these types of ”normally abnormal” values in purple or green rather than red font for abnormal, or via some other designating graphic. The purple font (or whatever display mode was utilized) would designate the value as technically abnormal, but perhaps notcontextually abnormal. Such designations are particularly important for caregivers who are not familiar with the patient.
It also might be desirable to use a combination of accumulated historical data from the monitor and the EHR to formulate personalized alarm limits for each patient. Such personalized alarm limits would provide a smarter range of acceptable values for each patient and perhaps also act to reduce the unacceptable number of false positive alarms that currently plague bedside caregivers (and patients) []. These alarm limits would be dynamically based on the input data and subject to reformulation as circumstances changed. We realize that any venture into alarm settings becomes a regulatory and potentially medico-legal issue, but these intimidating factors should not be allowed to grind potentially beneficial innovations to a halt. For example, “hard” limits could be built into the alarm machine so that the custom alarm limits could not fall outside certain designated values.
Supporting the Formulation of the Assessment
Building on both prior and new, interfaced and manually entered data as described above, the next framework element would consist of the formulation of the note in real time. This would consist of structured data so far as available and feasible, but is more likely to require real-time natural language processing performed on the free text being entered. Different takes on this kind of templated structure have already been introduced into several electronic systems. These include note templates created for specific purposes such as end-of-life discussions, or documentation of cardiopulmonary arrest. The very nature of these note types provides a robust context for the content. We also recognize that these shorter and more directed types of notes are not likely to require the kind of extensive clinical decision support (CDS) from which an admission or daily progress note may benefit.
Until the developers of EHRs find a way to fit structured data selection seamlessly and transparently into workflow, we will have to do the best we can with the free text that we have available. While this is a bit clunky in terms of data utilization purposes, perhaps it is not totally undesirable, as free text inserts a needed narrative element into the otherwise storyless EHR environment. Medical care can be described as an ongoing story and free text conveys this story in a much more effective and interesting fashion than do selected structured data bits. Furthermore, stories tend to be more distinctive than lists of structured data entries, which sometimes seem to vary remarkably little from patient to patient. But to extract the necessary information, the computer still needs a processed interpretation of that text. More complex systems are being developed and actively researched to act more analogously to our own ”human” form of clinical problem solving [], but until these systems are integrated into existing EHRs, clinicians may be able to help by being trained to minimize the potential confusion engendered by reducing completely unconstrained free text entries and/or utilizing some degree of standardization within the use of free text terminologies and contextual modifiers.
Employing the prior data (eg, diagnoses X, Y, Z from the previous note) and new data inputs (eg, laboratory results, imaging reports, and consultants’ recommendations) in conjunction with the assessment being entered, the system would have the capability to check for inconsistencies and omissions based on analysis of both prior and new entries. For example, a patient in the ICU has increasing temperature and heart rate, and decreasing oxygen saturation. These continuous variables are referenced against other patient features and risk factors to suggest the possibility that the patient has developed a pulmonary embolism or an infectious ventilator-associated complication. The system then displays these possible diagnoses within the working assessment screen with hyperlinks to the patient’s flow sheets and other data supporting the suggested problems (). The formulation of the assessment is clearly not as potentially evidence-based as that of the plan; however, there should still be dynamic, automatic and rapid searches performed for pertinent supporting material in the formulation of the assessment. These would include the medical literature, including textbooks, online databases, and applications such as WebMD. The relevant literature that the system has identified, supporting the associations listed in the assessment and plan, can then be screened by the user for accuracy and pertinence to the specific clinical context. Another potentially useful CDS tool for assessment formulation is a modality we have termed dynamic clinical data mining (DCDM) []. DCDM draws upon the power of large sets of population health data to provide differential diagnoses associated with groupings or constellations of symptoms and findings. Similar to the process just described, the clinician would then have the ability to review and incorporate these suggestions or not.
An optional active search function would also be provided throughout the note creation process for additional flexibility—clinicians are already using search engines, but doing so sometimes in the absence of specific clinical search algorithms (eg, a generic search engine such Google). This may produce search results that are not always of the highest possible quality [,]. The EHR-embedded search engine would have its algorithm modified to meet the task as Google has done previously for its search engine []. The searchable TRIP database provides a search engine for high-quality clinical evidence, as do the search modalities within Up to Date, Dynamed, BMJ Clinical Evidence, and others [,].
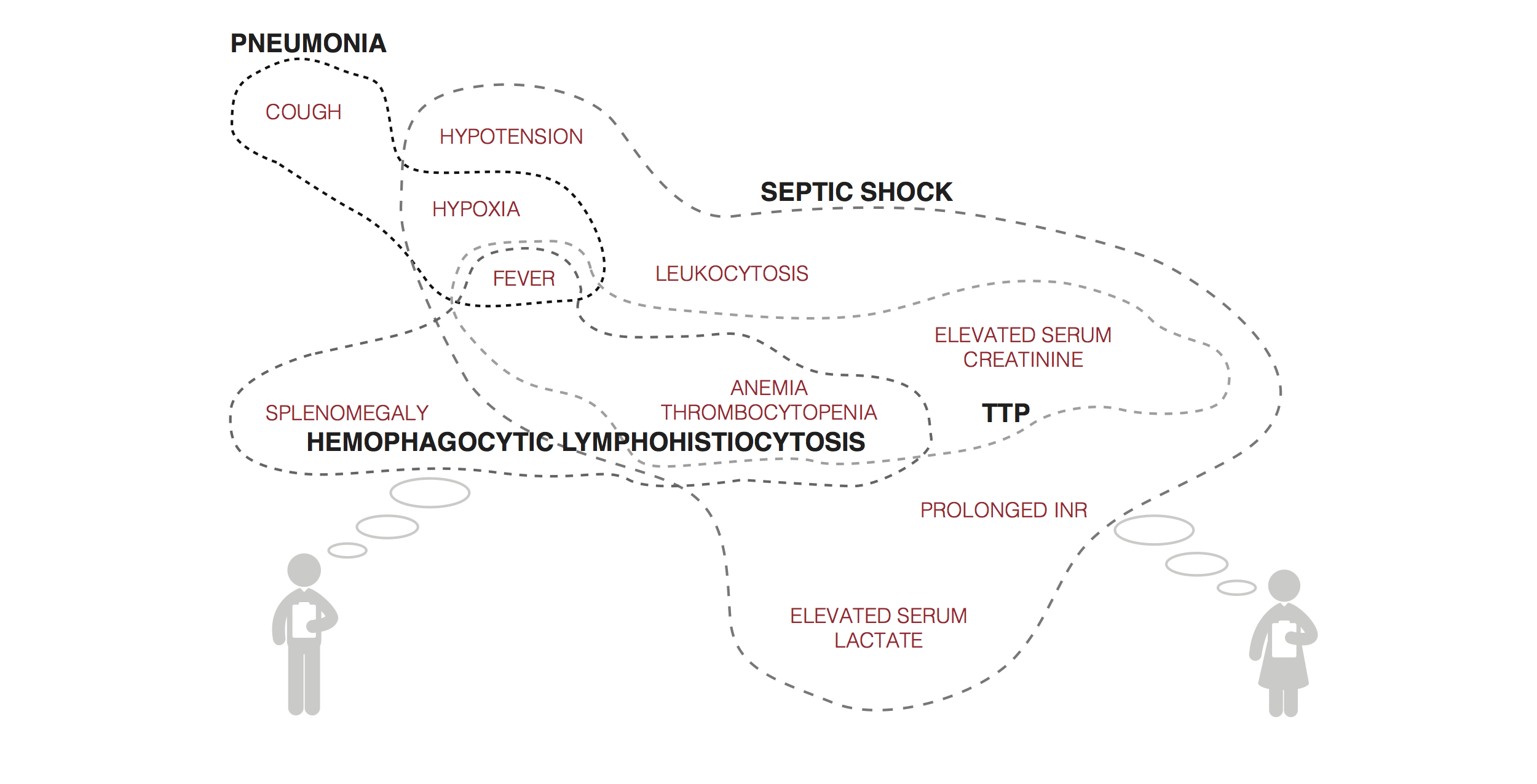
http://medinform.jmir.org/article/viewFile/4192/1/68491
Figure 2. Mock visualization of symptoms, signs, laboratory results, and other data input and systems suggestion for differential diagnoses.
Supporting the Formulation of the Plan
With the assessment formulated, the system would then formulate a proposed plan using EBM inputs and DCDM refinements for issues lying outside EBM knowledge. Decision support for plan formulation would include items such as randomized control trials (RCTs), observational studies, clinical practice guidelines (CPGs), local guidelines, and other relevant elements (eg, Cochrane reviews). The system would provide these supporting modalities in a hierarchical fashion using evidence of the highest quality first before proceeding down the chain to lower quality evidence. Notably, RCT data are not available for the majority of specific clinical questions, or it is not applicable because the results cannot be generalized to the patient at hand due to the study’s inclusion and exclusion criteria []. Sufficiently reliable observational research data also may not be available, although we expect that the holes in the RCT literature will be increasingly filled by observational studies in the near future [,]. In the absence of pertinent evidence-based material, the system would include the functionality which we have termed DCDM, and our Stanford colleagues have termed the “green button” [,]. This still-theoretical process is described in detail in the references, but in brief, DCDM would utilize a search engine type of approach to examine a population database to identify similar patients on the basis of the information entered in the EHR. The prior treatments and outcomes of these historical patients would then be analyzed to present options for the care of the current patient that were, to a large degree, based on prior data. The efficacy of DCDM would depend on, among other factors, the availability of a sufficiently large population EHR database, or an open repository that would allow for the sharing of patient data between EHRs. This possibility is quickly becoming a reality with the advent of large, deidentified clinical databases such as that being created by the Patient Centered Outcomes Research Institute [].
The tentative plan could then be modified by the user on the basis of her or his clinical “wetware” analysis. The electronic workflow could be designed in a number of ways that were modifiable per user choice/customization. For example, the user could first create the assessment and plan which would then be subject to comment and modification by the automatic system. This modification might include suggestions such as adding entirely new items, as well as the editing of entered items. In contrast, as described, the system could formulate an original assessment and plan that was subject to final editing by the user. In either case, the user would determine the final output, but the system would record both system and final user outputs for possible reporting purposes (eg, consistency with best practices). Another design approach might be to display the user entry in toto on the left half of a computer screen and a system-formulated assessment () and plan on the right side for comparison. Links would be provided throughout the system formulation so that the user could drill into EHR-provided suggestions for validation and further investigation and learning. In either type of workflow, the system would comparatively evaluate the final entered plan for consistency, completeness, and conformity with current best practices. The system could display the specific items that came under question and why. Users may proceed to adopt or not, with the option to justify their decision. Data reporting analytics could be formulated on the basis of compliance with EBM care. Such analytics should be done and interpreted with the knowledge that EBM itself is a moving target and many clinical situations do not lend themselves to resolution with the current tools supplied by EBM.
Since not all notes call for this kind of extensive decision support, the CDS material could be displayed in a separate columnar window adjacent to the main part of the screen where the note contents were displayed so that workflow is not affected. Another possibility would be an “opt-out” button by which the user would choose not to utilize these system resources. This would be analogous but functionally opposite to the “green button” opt-in option suggested by Longhurst et al, and perhaps be designated the “orange button” to clearly make this distinction []. Later, the system would make a determination as to whether this lack of EBM utilization was justified, and provide a reminder if the care was determined to be outside the bounds of current best practices. While the goal is to keep the user on the EBM track as much as feasible, the system has to “realize” that real care will still extend outside those bounds for some time, and that some notes and decisions simply do not require such machine support.
There are clearly still many details to be worked out regarding the creation and use of a fully integrated bidirectional EHR. There currently are smaller systems that use some components of what we propose. For example, a large Boston hospital uses a program called QPID which culls all previously collected patient data and uses a Google-like search to identify specific details of relevant prior medical history which is then displayed in a user-friendly fashion to assist the clinician in making real-time decisions on admission []. Another organization, the American Society of Clinical Oncology, has developed a clinical Health IT tool called CancerLinQ which utilizes large clinical databases of cancer patients to trend current practices and compare the specific practices of individual providers with best practice guidelines []. Another hospital system is using many of the components discussed in a new, internally developed platform called Fluence that allows aggregation of patient information, and applies already known clinical practice guidelines to patients’ problem lists to assist practitioners in making evidenced-based decisions []. All of these efforts reflect inadequacies in current EHRs and are important pieces in the process of selectively and wisely incorporating these technologies into EHRs, but doing so universally will be a much larger endeavor.
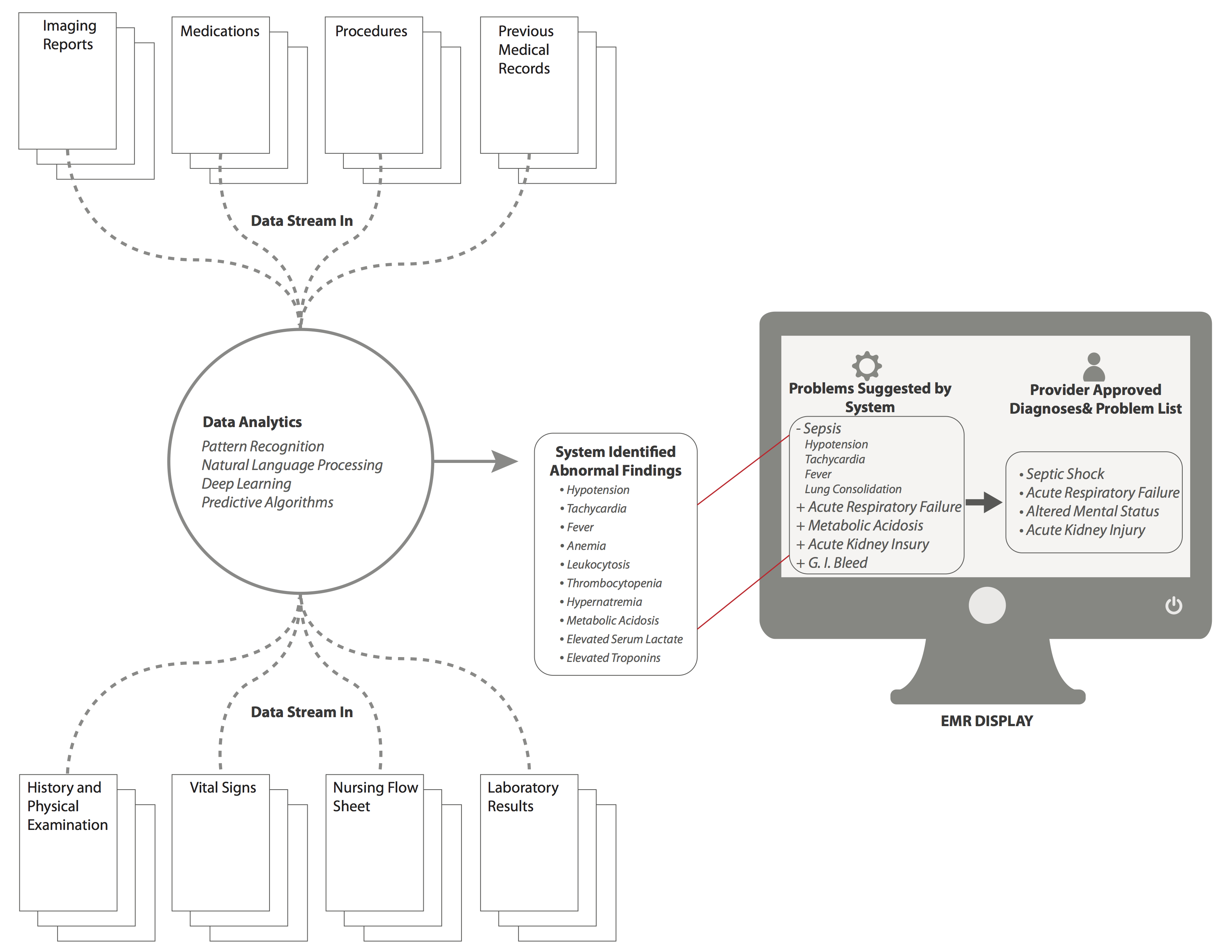

http://medinform.jmir.org/article/viewFile/4192/1/68492
Figure 3. Mock screenshot for the “Assessment and Plan” screen with background data analytics. Based on background analytics that are being run by the system at all times, a series of “problems” are identified and suggested by the system, which are then displayed in the EMR in the box on the left. The clinician can then select problems that are suggested, or input new problems that are then displayed in the the box on the right of the EMR screen, and will now be apart of ongoing analytics for future assessment. View this figure
Conclusions
Medicine has finally entered an era in which clinical digitization implementations and data analytic systems are converging. We have begun to recognize the power of data in other domains and are beginning to apply it to the clinical space, applying digitization as a necessary but insufficient tool for this purpose (personal communication from Peter Szolovits, The Unreasonable Effectiveness of Clinical Data. Challenges in Big Data for Data Mining, Machine Learning and Statistics Conference, March 2014). The vast amount of information and clinical choices demands that we provide better supports for making decisions and effectively documenting them. The Institute of Medicine demands a “learning health care system” where analysis of patient data is a key element in continuously improving clinical outcomes []. This is also an age of increasing medical complexity bound up in increasing financial and time constraints. The latter dictate that medical practice should become more standardized and evidence-based in order to optimize outcomes at the lowest cost. Current EHRs, mostly implemented over the past decade, are a first step in the digitization process, but do not support decision-making or streamline the workflow to the extent to which they are capable. In response, we propose a series of information system enhancements that we hope can be seized, improved upon, and incorporated into the next generation of EHRs.
There is already government support for these advances: The Office of the National Coordinator for Health IT recently outlined their 6-year and 10-year plans to improve EHR and health IT interoperability, so that large-scale realizations of this idea can and will exist. Within 10 years, they envision that we “should have an array of interoperable health IT products and services that allow the health care system to continuously learn and advance the goal of improved health care.” In that, they envision an integrated system across EHRs that will improve not just individual health and population health, but also act as a nationwide repository for searchable and researchable outcomes data []. The first step to achieving that vision is by successfully implementing the ideas and the system outlined above into a more fully functional EHR that better supports both workflow and clinical decision-making. Further, these suggested changes would also contribute to making the note writing process an educational one, thereby justifying the very significant time and effort expended, and would begin to establish a true learning system of health care based on actual workflow practices. Finally, the goal is to keep clinicians firmly in charge of the decision loop in a “human-centered” system in which technology plays an essential but secondary role. As expressed in a recent article on the issue of automating systems []:
In this model (human centered automation)…technology takes over routine functions that a human operator has already mastered, issues alerts when unexpected situations arise, provides fresh information that expands the operator’s perspective and counters the biases that often distort human thinking. The technology becomes the expert’s partner, not the expert’s replacement.
Key Concepts and Terminology
A number of concepts and terms were introduced throughout this paper, and some clarification and elaboration of these follows:
- Affordable Care Act (ACA): Legislation passed in 2010 that constitutes two separate laws including the Patient Protection and Affordable Care Act and the Health Care and Education Reconciliation Act. These two pieces of legislation act together for the expressed goal of expanding health care coverage to low-income Americans through expansion of Medicaid and other federal assistance programs [].
- Clinical Decision Support (CDS) is defined by CMS as “a key functionality of health information technology” that encompasses a variety of tools including computerized alerts and reminders, clinical guidelines, condition-specific order sets, documentations templates, diagnostic support, and other tools that “when used effectively, increases quality of care, enhances health outcomes, helps to avoid errors and adverse events, improves efficiency, reduces costs, and boosts provider and patient satisfaction” [].
- Cognitive Computing is defined as “the simulation of human thought processes in a computerize model…involving self learning systems that use data mining, pattern recognition and natural language processing to mimic the way the human brain works” []. Defined by IBM as computer systems that “are trained using artificial intelligence and machine learning algorithms to sense, predict, infer and, in some ways, think” [].
- Deep learning is a form of machine learning (a more specific subgroup of cognitive computing) that utilizes multiple levels of data to make hierarchical connections and recognize more complex patterns to be able to infer higher level concepts from lower levels of input and previously inferred concepts []. demonstrates how this concept relates to patients illustrating the system recognizing patterns of signs and symptoms experienced by a patient, and then inferring a diagnosis (higher level concept) from those lower level inputs. The next level concept would be recognizing response to treatment for proposed diagnosis, and offering either alternative diagnoses, or change in therapy, with the system adapting as the patient’s course progresses.
- Dynamic clinical data mining (DCDM): First, data mining is defined as the “process of discovering patterns, automatically or semi-automatically, in large quantities of data” []. DCDM describes the process of mining and interpreting the data from large patient databases that contain prior and concurrent patient information including diagnoses, treatments, and outcomes so as to make real-time treatment decisions [].
- Natural Language Processing (NLP) is a process based on machine learning, or deep learning, that enables computers to analyze and interpret unstructured human language input to recognize and even act upon meaningful patterns [,].
References
- Weed LL. Medical records, patient care, and medical education. Ir J Med Sci 1964 Jun;462:271-282. [Medline]
- Celi L, Csete M, Stone D. Optimal data systems: the future of clinical predictions and decision support. Curr Opin Crit Care 2014 Oct;20(5):573-580. [CrossRef] [Medline]
- Cook DA, Sorensen KJ, Hersh W, Berger RA, Wilkinson JM. Features of effective medical knowledge resources to support point of care learning: a focus group study. PLoS One 2013 Nov;8(11):e80318 [FREE Full text] [CrossRef] [Medline]
- Cook DA, Sorensen KJ, Wilkinson JM, Berger RA. Barriers and decisions when answering clinical questions at the point of care: a grounded theory study. JAMA Intern Med 2013 Nov 25;173(21):1962-1969. [CrossRef] [Medline]
more ….
The Electronic Health Record: How far we have travelled, and where is journey’s end?
http://pharmaceuticalintelligence.com/2012/09/21/the-electronic-health-record-how-far-we-have-travelled-and-where-is-journeys-end/
A focus of the Accountable Care Act is improved delivery of quality, efficiency and effectiveness to the patients who receive healthcare in US from the providers in a coordinated system. The largest confounder in all of this is the existence of silos that are not readily crossed, handovers, communication lapses, and a heavy paperwork burden. We can add to that a large for profit insurance overhead that is disinterested in the patient-physician encounter. Finally, the knowledge base of medicine has grown sufficiently that physicians are challenged by the amount of data and the presentation in the Medical Record.
I present a review of the problems that have become more urgent to fix in the last decade. The administration and paperwork necessitated by health insurers, HMOs and other parties today may account for 40% of a physician’s practice, and the formation of large physician practice groups and alliances of the hospital and hospital staffed physicians (as well as hospital system alliances) has increased in response to the need to decrease the cost of non-patient care overhead. I discuss some of the points made by two innovators from the healthcare and the communications sectors.
I also call attention to the New York Times front page article calling attention to a sharp rise in inflation-adjusted Medicare payments for emergency-room services since 2006 due to upcoding at the highest level, partly related to the ability to physician ability to overstate the claim for service provided by correctible improvements I discuss below. (NY Times, 9/22/2012). The solution still has another built in step that requires quality control of both the input and the output, achievable today. This also comes at a time that there is a nationwide implementation of ICD-10 to replace ICD-9 coding.

US medical groups’ adoption of EHR (2005) (Photo credit: Wikipedia)
The first finding by Robert S Didner, on “Decision Making in the Clinical Setting”, concludes that the gathering of information has large costs while reimbursements for the activities provided have decreased, detrimental to the outcomes that are measured. He suggests that this data can be gathered and reformatted to improve its value in the clinical setting by leading to decisions with optimal outcomes. He outlines how this can be done.
The second is a discussion by Emergency Medicine physicians, Thomas A Naegele and harry P Wetzler, who have developed a Foresighted Practice Guideline (FPG) (“The Foresighted Practice Guideline Model: A Win-Win Solution”). They focus on collecting data from similar patients, their interventions, and treatments to better understand the value of alternative courses of treatment. Using the FPG model will enable physicians to elevate their practice to a higher level and they will have hard information on what works. These two views are more than 10 years old, and they are complementary.
Didner points out that there is no one sequence of tests and questions that can be optimal for all presenting clusters. Even as data and test results are acquired, the optimal sequence of information gathering is changed, depending on the gathered information. Thus, the dilemma is created of how to collect clinical data. Currently, the way information is requested and presented does not support the way decisions are made. Decisions are made in a “path-dependent” way, which is influenced by the sequence in which the components are considered. Ideally, it would require a separate form for each combination of presenting history and symptoms, prior to ordering tests, which is unmanageable. The blank paper format is no better, as the data is not collected in the way it would be used, and it constitutes separate clusters (vital signs, lab work{also divided into CBC, chemistry panel, microbiology, immunology, blood bank, special tests}]. Improvements have been made in the graphical presentation of a series of tests. Didner presents another means of gathering data in machine manipulable form that improves the expected outcomes. The basis for this model is that at any stage of testing and information gathering there is an expected outcome from the process, coupled with a metric, or hierarchy of values to determine the relative desirability of the possible outcomes.
He creates a value hierarchy:
- Minimize the likelihood that a treatable, life-threatening disorder is not treated.
- Minimize the likelihood that a treatable, permanently-disabling or disfiguring disorder is not treated.
- Minimize the likelihood that a treatable, discomfort causing disorder is not treated.
- Minimize the likelihood that a risky procedure, (treatment or diagnostic procedure) is inappropriately administered.
- Minimize the likelihood that a discomfort causing procedure is inappropriately administered.
- Minimize the likelihood that a costly procedure is inappropriately administered.
- Minimize the time of diagnosing and treating the patient.
- Minimize the cost of diagnosing and treating the patient.
In reference to a way of minimizing the number, time and cost of tests, he determines that the optimum sequence could be found using Claude Shannon’s Information theory. As to a hierarchy of outcome values, he refers to the QALY scale as a starting point. At any point where a determination is made there is disparate information that has to be brought together, such as, weight, blood pressure, cholesterol, etc. He points out, in addition, that the way the clinical information is organized is not opyimal for the way to display information to enhance human cognitive performance in decision support. Furthermore, he looks at the limit of short term memory as 10 chunks of information at any time, and he compares the positions of chess pieces on the board with performance of a grand master, if the pieces are in an order commensurate with a “line of attack”. The information has to be ordered in the way it is to be used! By presenting information used for a particular decision component in a compact space the load on short term memory is reduced, and there is less strain in searching for the relevant information.
He creates a Table to illustrate the point.
Correlation of weight with other cardiac risk factors
Chol 0.759384
HDL -0.53908
LDL 0.177297
bp-syst 0.424728
bp-dia 0.516167
Triglyc 0.637817
The task of the information system designer is to provide or request the right information, in the best form, at each stage of the procedure.
The FPG concept as deployed by Naegele and Wetzler is a model for design of a more effective health record that has already shown substantial proof of concept in the emergency room setting. In principle, every clinical encounter is viewed as a learning experience that requires the collection of data , learning from similar patients, and comparing the value of alternative courses of treatment. The framework for standard data collection is the FPG model. The FPG is distinguished from hindsighted guidelines which are utilized by utilization and peer review organizations. Over time, the data forms patient clusters and enables the physician to function at a higher level.
Hypothesis construction is experiential, and hypothesis generation and testing is required to go from art to science in the complex practice of medicine. In every encounter there are 3 components: patient, process, and outcome. The key to the process is to collect data on patients, processes and outcomes in a standard way. The main problem with a large portion of the chart is that the description is not uniform. This is not fully resolved with good natural language encryption. The standard words and phrases that may be used for a particular complaint or condition constitute a guideline. This type of “guided documentation” is a step in moving toward a guided practice. It enables physicians to gather data on patients, processes and outcomes of care in routine settings, and they can be reviewed and updated. This is a higher level of methodology than basing guidelines on “consensus and opinion”.
When Lee Goldman, et al., created the guideline for classifying chest pain in the emergency room, the characteristics of the chest pain was problematic. In dealing with this he determined that if the chest pain was “stabbing”, or if it radiated to the right foot, heart attack is excluded.
The IOM is intensely committed to practice guidelines for care. The guidelines are the data bases of the science of medical decision making and disposition processing, and are related to process-flow. However, the hindsighted or retrospective approach is diagnosis or procedure oriented. HPGs are the tool used in utilization review. The FPG model focuses on the physician-patient encounter and is problem oriented. We can go back further and remember the contribution by Lawrence Weed to the “structured medical record”.
The physicians today use an FPG framework in looking at a problem or pathology (especially in pathology, which extends the classification by used of biomarker staining). The Standard Patient File Format (SPPF) was developed by Weed and includes: 1. Patient demographics; 2. Front of the chart; 3. Subjective: Objective; Assessment/diagnosis;6. Plan; Back of the chart. The FPG retains the structure of the SPPF All of the words and phrases in the FPG are the data base for the problem or condition. The current construct of the chart is uninviting: nurses notes, medications, lab results, radiology, imaging.
Realtime Clinical Expert Support and Validation System
Gil David and Larry Bernstein have developed, in consultation with Prof. Ronald Coifman, in the Yale University Applied Mathematics Program, a software system that is the equivalent of an intelligent Electronic Health Records Dashboard that provides empirical medical reference and suggests quantitative diagnostics options.
The introduction of a DASHBOARD has allowed a presentation of drug reactions, allergies, primary and secondary diagnoses, and critical information about any patient the care giver needing access to the record. The advantage of this innovation is obvious. The startup problem is what information is presented and how it is displayed, which is a source of variability and a key to its success. It is also imperative that the extraction of data from disparate sources will, in the long run, further improve the diagnostic process. For instance, the finding of both ST depression on EKG coincident with an increase of a cardiac biomarker (troponin). Through the application of geometric clustering analysis the data may interpreted in a more sophisticated fashion in order to create a more reliable and valid knowledge-based opinion. In the hemogram one can view data reflecting the characteristics of a broad spectrum of medical conditions. Characteristics expressed as measurements of size, density, and concentration, resulting in more than a dozen composite variables, including the mean corpuscular volume (MCV), mean corpuscular hemoglobin concentration (MCHC), mean corpuscular hemoglobin (MCH), total white cell count (WBC), total lymphocyte count, neutrophil count (mature granulocyte count and bands), monocytes, eosinophils, basophils, platelet count, and mean platelet volume (MPV), blasts, reticulocytes and platelet clumps, as well as other features of classification. This has been described in a previous post.
It is beyond comprehension that a better construct has not be created for common use.
W Ruts, S De Deyne, E Ameel, W Vanpaemel,T Verbeemen, And G Storms. Dutch norm data for 13 semantic categoriesand 338 exemplars. Behavior Research Methods, Instruments, & Computers 2004; 36 (3): 506–515.
De Deyne, S Verheyen, E Ameel, W Vanpaemel, MJ Dry, WVoorspoels, and G Storms. Exemplar by feature applicability matrices and other Dutch normative data for semantic concepts.
Behavior Research Methods 2008; 40 (4): 1030-1048
Landauer, T. K., Ross, B. H., & Didner, R. S. (1979). Processing visually presented single words: A reaction time analysis [Technical memorandum]. Murray Hill, NJ: Bell Laboratories. Lewandowsky , S. (1991).
Weed L. Automation of the problem oriented medical record. NCHSR Research Digest Series DHEW. 1977;(HRA)77-3177.
Naegele TA. Letter to the Editor. Amer J Crit Care 1993;2(5):433.
The potential contribution of informatics to healthcare is more than currently estimated
http://pharmaceuticalintelligence.com/2013/02/18/the-potential-contribution-of-informatics-to-healthcare-is-more-than-currently-estimated/
The estimate of improved costsavings in healthcare and diagnostic accuracy is extimated to be substantial. I have written about the unused potential that we have not yet seen. In short, there is justification in substantial investment in resources to this, as has been proposed as a critical goal. Does this mean a reduction in staffing? I wouldn’t look at it that way. The two huge benefits that would accrue are:
- workflow efficiency, reducing stress and facilitating decision-making.
- scientifically, primary knowledge-based decision-support by well developed algotithms that have been at the heart of computational-genomics.
Can computers save health care? IU research shows lower costs, better outcomes
Cost per unit of outcome was $189, versus $497 for treatment as usual
Last modified: Monday, February 11, 2013
BLOOMINGTON, Ind. — New research from
Indiana University has found that machine learning — the same computer science discipline that helped create voice recognition systems, self-driving cars and credit card fraud detection systems — can drastically improve both the cost and quality of
health care in the United States.
Physicians using an artificial intelligence framework that predicts future outcomes would have better patient outcomes while significantly lowering health care costs.
Using an artificial intelligence framework combining Markov Decision Processes and Dynamic Decision Networks, IU
School of Informatics and Computing researchers Casey Bennett and Kris Hauser show how simulation modeling that understands and predicts the outcomes of treatment could
- reduce health care costs by over 50 percent while also
- improving patient outcomes by nearly 50 percent.
The work by Hauser, an assistant professor of computer science, and Ph.D. student Bennett improves upon their earlier work that
- showed how machine learning could determine the best treatment at a single point in time for an individual patient.
By using a new framework that employs sequential decision-making, the previous single-decision research
- can be expanded into models that simulate numerous alternative treatment paths out into the future;
- maintain beliefs about patient health status over time even when measurements are unavailable or uncertain; and
- continually plan/re-plan as new information becomes available.
In other words, it can “think like a doctor.” (Perhaps better because of the limitation in the amount of information a bright, competent physician can handle without error!)
“The Markov Decision Processes and Dynamic Decision Networks enable the system to deliberate about the future, considering all the different possible sequences of actions and effects in advance, even in cases where we are unsure of the effects,” Bennett said. Moreover, the approach is non-disease-specific — it could work for any diagnosis or disorder, simply by plugging in the relevant information. (This actually raises the question of what the information input is, and the cost of inputting.)
The new work addresses three vexing issues related to health care in the U.S.:
- rising costs expected to reach 30 percent of the gross domestic product by 2050;
- a quality of care where patients receive correct diagnosis and treatment less than half the time on a first visit;
- and a lag time of 13 to 17 years between research and practice in clinical care.
Framework for Simulating Clinical Decision-Making
“We’re using modern computational approaches to learn from clinical data and develop complex plans through the simulation of numerous, alternative sequential decision paths,” Bennett said. “The framework here easily out-performs the current treatment-as-usual, case-rate/fee-for-service models of health care.” (see the above)
Bennett is also a data architect and research fellow with Centerstone Research Institute, the research arm of Centerstone, the nation’s largest not-for-profit provider of community-based behavioral health care. The two researchers had access to clinical data, demographics and other information on over 6,700 patients who had major clinical depression diagnoses, of which about 65 to 70 percent had co-occurring chronic physical disorders like diabetes, hypertension and cardiovascular disease. Using 500 randomly selected patients from that group for simulations, the two
- compared actual doctor performance and patient outcomes against
- sequential decision-making models
using real patient data.
They found great disparity in the cost per unit of outcome change when the artificial intelligence model’s
- cost of $189 was compared to the treatment-as-usual cost of $497.
- the AI approach obtained a 30 to 35 percent increase in patient outcomes
Bennett said that “tweaking certain model parameters could enhance the outcome advantage to about 50 percent more improvement at about half the cost.”
While most medical decisions are based on case-by-case, experience-based approaches, there is a growing body of evidence that complex treatment decisions might be effectively improved by AI modeling. Hauser said “Modeling lets us see more possibilities out to a further point – because they just don’t have all of that information available to them.” (Even then, the other issue is the processing of the information presented.)
Using the growing availability of electronic health records, health information exchanges, large public biomedical databases and machine learning algorithms, the researchers believe the approach could serve as the basis for personalized treatment through integration of diverse, large-scale data passed along to clinicians at the time of decision-making for each patient. Centerstone alone, Bennett noted, has access to health information on over 1 million patients each year. “Even with the development of
new AI techniques that can approximate or even surpass human decision-making performance, we believe that the most effective long-term path could be combining artificial intelligence with human clinicians,” Bennett said. “Let humans do what they do well, and let machines do what they do well. In the end, we may maximize the potential of both.”
“
Artificial Intelligence Framework for Simulating Clinical Decision-Making: A
Markov Decision Process Approach” was published recently in Artificial Intelligence in Medicine. The research was funded by the Ayers Foundation, the Joe C. Davis Foundation and Indiana University.
For more information or to speak with Hauser or Bennett, please contact Steve Chaplin, IU Communications, at 812-856-1896 or stjchap@iu.edu.
IBM Watson Finally Graduates Medical School
It’s been more than a year since IBM’s Watson computer appeared on Jeopardy and defeated several of the game show’s top champions. Since then the supercomputer has been furiously “studying” the healthcare literature in the hope that it can beat a far more hideous enemy: the 400-plus biomolecular puzzles we collectively refer to as cancer.
Anomaly Based Interpretation of Clinical and Laboratory Syndromic Classes
Larry H Bernstein, MD, Gil David, PhD, Ronald R Coifman, PhD. Program in Applied Mathematics, Yale University, Triplex Medical Science.
Statement of Inferential Second Opinion
Realtime Clinical Expert Support and Validation System
Gil David and Larry Bernstein have developed, in consultation with Prof. Ronald Coifman, in the Yale University Applied Mathematics Program, a software system that is the equivalent of an intelligent Electronic Health Records Dashboard that provides
- empirical medical reference and suggests quantitative diagnostics options.
Background
The current design of the Electronic Medical Record (EMR) is a linear presentation of portions of the record by
- services, by
- diagnostic method, and by
- date, to cite examples.
This allows perusal through a graphical user interface (GUI) that partitions the information or necessary reports in a workstation entered by keying to icons. This requires that the medical practitioner finds
- the history,
- medications,
- laboratory reports,
- cardiac imaging and EKGs, and
- radiology
in different workspaces. The introduction of a DASHBOARD has allowed a presentation of
- drug reactions,
- allergies,
- primary and secondary diagnoses, and
- critical information about any patient the care giver needing access to the record.
The advantage of this innovation is obvious. The startup problem is what information is presented and how it is displayed, which is a source of variability and a key to its success.
Proposal
We are proposing an innovation that supercedes the main design elements of a DASHBOARD and
- utilizes the conjoined syndromic features of the disparate data elements.
So the important determinant of the success of this endeavor is that it facilitates both
- the workflow and
- the decision-making process
- with a reduction of medical error.
This has become extremely important and urgent in the 10 years since the publication “To Err is Human”, and the newly published finding that reduction of error is as elusive as reduction in cost. Whether they are counterproductive when approached in the wrong way may be subject to debate.
We initially confine our approach to laboratory data because it is collected on all patients, ambulatory and acutely ill, because the data is objective and quality controlled, and because
- laboratory combinatorial patterns emerge with the development and course of disease. Continuing work is in progress in extending the capabilities with model data-sets, and sufficient data.
It is true that the extraction of data from disparate sources will, in the long run, further improve this process. For instance, the finding of both ST depression on EKG coincident with an increase of a cardiac biomarker (troponin) above a level determined by a receiver operator curve (ROC) analysis, particularly in the absence of substantially reduced renal function.
The conversion of hematology based data into useful clinical information requires the establishment of problem-solving constructs based on the measured data. Traditionally this has been accomplished by an intuitive interpretation of the data by the individual clinician. Through the application of geometric clustering analysis the data may interpreted in a more sophisticated fashion in order to create a more reliable and valid knowledge-based opinion.
The most commonly ordered test used for managing patients worldwide is the hemogram that often incorporates the review of a peripheral smear. While the hemogram has undergone progressive modification of the measured features over time the subsequent expansion of the panel of tests has provided a window into the cellular changes in the production, release or suppression of the formed elements from the blood-forming organ to the circulation. In the hemogram one can view data reflecting the characteristics of a broad spectrum of medical conditions.
Progressive modification of the measured features of the hemogram has delineated characteristics expressed as measurements of
- size,
- density, and
- concentration,
resulting in more than a dozen composite variables, including the
- mean corpuscular volume (MCV),
- mean corpuscular hemoglobin concentration (MCHC),
- mean corpuscular hemoglobin (MCH),
- total white cell count (WBC),
- total lymphocyte count,
- neutrophil count (mature granulocyte count and bands),
- monocytes,
- eosinophils,
- basophils,
- platelet count, and
- mean platelet volume (MPV),
- blasts,
- reticulocytes and
- platelet clumps,
- perhaps the percent immature neutrophils (not bands)
- as well as other features of classification.
The use of such variables combined with additional clinical information including serum chemistry analysis (such as the Comprehensive Metabolic Profile (CMP)) in conjunction with the clinical history and examination complete the traditional problem-solving construct. The intuitive approach applied by the individual clinician is limited, however,
- by experience,
- memory and
- cognition.
The application of rules-based, automated problem solving may provide a more reliable and valid approach to the classification and interpretation of the data used to determine a knowledge-based clinical opinion.
The classification of the available hematologic data in order to formulate a predictive model may be accomplished through mathematical models that offer a more reliable and valid approach than the intuitive knowledge-based opinion of the individual clinician. The exponential growth of knowledge since the mapping of the human genome has been enabled by parallel advances in applied mathematics that have not been a part of traditional clinical problem solving. In a univariate universe the individual has significant control in visualizing data because unlike data may be identified by methods that rely on distributional assumptions. As the complexity of statistical models has increased, involving the use of several predictors for different clinical classifications, the dependencies have become less clear to the individual. The powerful statistical tools now available are not dependent on distributional assumptions, and allow classification and prediction in a way that cannot be achieved by the individual clinician intuitively. Contemporary statistical modeling has a primary goal of finding an underlying structure in studied data sets.
In the diagnosis of anemia the variables MCV,MCHC and MCH classify the disease process into microcytic, normocytic and macrocytic categories. Further consideration of
proliferation of marrow precursors,
- the domination of a cell line, and
- features of suppression of hematopoiesis
provide a two dimensional model. Several other possible dimensions are created by consideration of
- the maturity of the circulating cells.
The development of an evidence-based inference engine that can substantially interpret the data at hand and convert it in real time to a “knowledge-based opinion” may improve clinical problem solving by incorporating multiple complex clinical features as well as duration of onset into the model.
An example of a difficult area for clinical problem solving is found in the diagnosis of SIRS and associated sepsis. SIRS (and associated sepsis) is a costly diagnosis in hospitalized patients. Failure to diagnose sepsis in a timely manner creates a potential financial and safety hazard. The early diagnosis of SIRS/sepsis is made by the application of defined criteria (temperature, heart rate, respiratory rate and WBC count) by the clinician. The application of those clinical criteria, however, defines the condition after it has developed and has not provided a reliable method for the early diagnosis of SIRS. The early diagnosis of SIRS may possibly be enhanced by the measurement of proteomic biomarkers, including transthyretin, C-reactive protein and procalcitonin. Immature granulocyte (IG) measurement has been proposed as a more readily available indicator of the presence of
- granulocyte precursors (left shift).
The use of such markers, obtained by automated systems in conjunction with innovative statistical modeling, may provide a mechanism to enhance workflow and decision making.
An accurate classification based on the multiplicity of available data can be provided by an innovative system that utilizes the conjoined syndromic features of disparate data elements. Such a system has the potential to facilitate both the workflow and the decision-making process with an anticipated reduction of medical error.
This study is only an extension of our approach to repairing a longstanding problem in the construction of the many-sided electronic medical record (EMR). On the one hand, past history combined with the development of Diagnosis Related Groups (DRGs) in the 1980s have driven the technology development in the direction of “billing capture”, which has been a focus of epidemiological studies in health services research using data mining.
In a classic study carried out at Bell Laboratories, Didner found that information technologies reflect the view of the creators, not the users, and Front-to-Back Design (R Didner) is needed. He expresses the view:
“Pre-printed forms are much more amenable to computer-based storage and processing, and would improve the efficiency with which the insurance carriers process this information. However, pre-printed forms can have a rather severe downside. By providing pre-printed forms that a physician completes
to record the diagnostic questions asked,
- as well as tests, and results,
- the sequence of tests and questions,
might be altered from that which a physician would ordinarily follow. This sequence change could improve outcomes in rare cases, but it is more likely to worsen outcomes. “
Decision Making in the Clinical Setting. Robert S. Didner
A well-documented problem in the medical profession is the level of effort dedicated to administration and paperwork necessitated by health insurers, HMOs and other parties (ref). This effort is currently estimated at 50% of a typical physician’s practice activity. Obviously this contributes to the high cost of medical care. A key element in the cost/effort composition is the retranscription of clinical data after the point at which it is collected. Costs would be reduced, and accuracy improved, if the clinical data could be captured directly at the point it is generated, in a form suitable for transmission to insurers, or machine transformable into other formats. Such data capture, could also be used to improve the form and structure of how this information is viewed by physicians, and form a basis of a more comprehensive database linking clinical protocols to outcomes, that could improve the knowledge of this relationship, hence clinical outcomes.
How we frame our expectations is so important that
- it determines the data we collect to examine the process.
In the absence of data to support an assumed benefit, there is no proof of validity at whatever cost. This has meaning for
- hospital operations, for
- nonhospital laboratory operations, for
- companies in the diagnostic business, and
- for planning of health systems.
In 1983, a vision for creating the EMR was introduced by Lawrence Weed and others. This is expressed by McGowan and Winstead-Fry.
J J McGowan and P Winstead-Fry. Problem Knowledge Couplers: reengineering evidence-based medicine through interdisciplinary development, decision support, and research.
Bull Med Libr Assoc. 1999 October; 87(4): 462–470. PMCID: PMC226622 Copyright notice

Example of Markov Decision Process (MDP) transition automaton (Photo credit: Wikipedia)
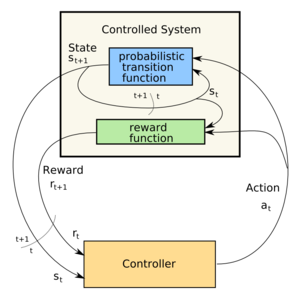
Control loop of a Markov Decision Process (Photo credit: Wikipedia)

English: IBM’s Watson computer, Yorktown Heights, NY (Photo credit: Wikipedia)

English: Increasing decision stakes and systems uncertainties entail new problem solving strategies. Image based on a diagram by Funtowicz, S. and Ravetz, J. (1993) “Science for the post-normal age” Futures 25:735–55 (http://dx.doi.org/10.1016/0016-3287(93)90022-L). (Photo credit: Wikipedia)
Read Full Post »