Accurate Identification and Treatment of Emergent Cardiac Events
Author: Larry H Bernstein, MD, FCAP
In the immediately preceding article, I discussed the difficulties in predicting long-term safety for developing drugs, and the cost of failure in early identification.
It is not the same scale of issue as for the patient emergently presenting to the ED. Despite enormous efforts to reduce the development of and the complications of acute ischemia related cardiac events, the accurate diagnosis of the patient presenting to the emergency room is still, as always, reliant on clinical history, physical examination, effective use of the laboratory, and increasingly helpful imaging technology. The main issue that we have a consensus agreement that PLAQUE RUPTURE is not the only basis for a cardiac ischemic event. The introduction of high sensitivity troponin tests has made it no less difficult after throwing out the receiver-operator characteristic curve (ROC) and assuming that any amount of cardiac troponin released from the heart is pathognomonic of an acute ischemic event. This has resulted in a consensus agreement that
- ctn measurement at a coefficient of variant (CV) measurement in excess of 2 Std dev of the upper limit of normal is a “red flag”
- signaling AMI? or other cardiomyopathic disorder
This is the catch. The ROC curve established AMI in ctn(s) that were accurate for NSTEMI – (and probably not needed with STEMI or new Q-wave, not previously seen) –
- ST-depression
- T-wave inversion
- in the presence of other findings
- suspicious for AMI
Wouldn’t it be nice if it was like seeing a robin on your lawn after a harsh winter? Life isn’t like that. When acute illness hits the patient may well present with ambiguous findings. We are accustomed to relying on
- clinical history
- family history
- co-morbidities, eg., diabetes, obesity, limited activity?, diet?
- stroke and/or peripheral vascular disease
- hypertension and/or renal vascular disease
- aortic atherosclerosis or valvular heart disease
- these are evidence, and they make up syndromic classes
- Electrocardiogram – 12 lead EKG (as above)
- Laboratory tests
- isoenzyme MB of creatine kinase (CK)… which declines after 12-18 hours
- isoenzyme-1 of LD if the time of appearance is > day-1 after initial symptoms (no longer used)
- cardiac troponin cTnI or cTnT
- genome testing
- advanced analysis of EKG
This may result in more consults for cardiologists, but it lays the ground for better evaluation of the patient, in the long run. When you look at the amount of information that has to be presented to the physician, there is serious need for improvement in the electronic medical record to benefit the patient and the caregivers. Recently, we have a publication on a new test that has been evaluated, closely related to the C-reactive protein (CRP), a test that has generated much discussion over the effect of treatment for patients who have elevated CRP in the absence of increased LDL cholesterol, diabetes, or obvious atherosclerotic comorbidities. The serum pentraxin 3 test is related to cell mediated immunity, and an evaluation has been published in the Journal of Investigative Medicine.
Journal of Investigative Medicine Feb 2013; 61 (2): 278–285.
http://dx.doi.org/10.231/JIM.0b013e31827c2971
Serum Pentraxin 3 Levels Are Associated With the Complexity and Severity of Coronary Artery Disease in Patients With Stable Angina Pectoris
Karakas, Mehmet Fatih MD*; Buyukkaya, Eyup MD*; Kurt, Mustafa MD*; et al.
From the Departments of Cardiology and,Clinical Biochemistry, Mustafa Kemal University, Tayfur Ata Sokmen Medical School, Hatay, Turkey.
Reprints: Mehmet Fatih Karakas, MD, Antakya 31005, Turkey. E-mail: mfkarakas@hotmail.com.
Abstract
Background: Atherosclerosis is a complex inflammatory process. Although pentraxin 3 (PTX-3), a newly identified inflammatory marker, was associated with adverse outcomes in stable angina pectoris,
- an association between PTX-3 and the complexity of coronary artery disease (CAD) has not been reported.
The aim of the present study is to assess
- the association between the level of PTX-3 and
- the complexity and severity of CAD assessed with
- SYNTAX and Gensini scores in patients with stable angina pectoris.
Methods: The study population is 2 groups:
- 161 patients with anginal symptoms and evidence of ischemia
- who underwent coronary angiography and
- 50 age- and sex- matched control subjects without evidence of ischemia .
Patients were grouped into 3 groups according to the complexity and severity of coronary lesions
- assessed by the SYNTAX score (30 patients with a SYNTAX score of 0 were excluded).
Serum PTX-3 and high-sensitivity C-reactive protein (hs-CRP) levels were measured in both groups.
Results: The PTX-3 levels demonstrated
- an increase from low to high SYNTAX groups (r = 0.72, P < 0.001).
Whereas the low SYNTAX group had statistically significantly higher PTX-3 levels when compared with the control group (0.50 ± 0.01 vs 0.24 ± 0.01 ng/mL, P < 0.001),
- the hs-CRP levels were not different (0.81 ± 0.42 vs 0.86 ± 0.53 mg/dL, P = 0.96).
- but the intermediate SYNTAX group had higher hs-CRP levels compared with the low SYNTAX group (1.3 ± 0.66 vs 0.86 ± 0.53 mg/dL, P = 0.002).
Serum PTX-3 levels and hs-CRP levels were both correlated with the SYNTAX scores and Gensini scores (for SYNTAX: r = 0.87 [P < 0.001] and r = 0.36 [P = 0.01]; for Gensini: r = 0.75 [P < 0.001] and r = 0.27 [P = 0.002], respectively), and
- according to the results of univariate and multivariate analyses, for “intermediate and high” SYNTAX scores, age, diabetes mellitus, low-density lipoprotein cholesterol, hs-CRP, and PTX-3
- were found to be independent predictors, whereas
- for the presence of “high” SYNTAX score only PTX-3 was found to be an independent predictor.
- The receiver operating characteristic curve analysis further revealed that the PTX-3 level was
- a strong indicator of high SYNTAX score with an area under the curve of 0.91 (95% confidence interval, 0.86–0.96).
Conclusions: Pentraxin 3, a novel inflammatory marker, was more tightly associated with the complexity and severity of CAD than hs-CRP and
- it was found to be an independent predictor for high SYNTAX score.
The association between atherosclerosis and inflammation has been more understood during recent years. Currently, atherosclerosis is considered as a complex inflammatory process in which
- leukocytes and inflammatory markers are involved.1
Several inflammatory markers
- high-sensitivity C-reactive protein (hs-CRP),
- fibrinogen, and
- complement C3…. are associated with cardiovascular events.1–5
Pentraxin 3 (PTX-3), that resembles CRP both in structure and function,1 is produced both by
- hematopoietic cells such as macrophages, dendritic cells, neutrophils, and by
- nonhematopoietic cells such as fibroblasts and vascular endothelial cells.2
Plasma PTX-3 levels may be elevated in patients with
- vasculitis,6
- acute myocardial infarction,7,8 and
- systemic inflammation or sepsis,9
- psoriasis,
- unstable angina pectoris, and
- heart failure.10–13
Dubin et al14 reported that PTX-3 levels are associated with with adverse outcomes in stable angina pectoris (SAP). Despite reports about the association of PTX-3 and coronary artery disease (CAD),
an association between the level of PTX-3 and the complexity and severity of CAD is not established.15,16 Thus, the aim of this study was
- to assess the association between the level of PTX-3 and the complexity and severity of CAD assessed with SYNTAX and Gensini scores in SAP patients.
MATERIALS AND METHODS
Of 211 patients were prospectively recruited, 161 SAP patients with evidence of ischemia (positive treadmill or myocardial perfusion scan) underwent coronary angiography for suspected CAD, and 50 age- and sex- matched outpatient subjects with a negative treadmill or myocardial perfusion scan test were taken as the control group. Patients were excluded if they had
- acute coronary syndrome
- history of previous myocardial infarction;
- coronary artery bypass grafting or percutaneous coronary intervention;
- secondary hypertension (HT);
- renal failure;
- hepatic failure;
- chronic obstructive lung disease and/or
- manifest heart disease, such as
- cardiac failure (left ventricular ejection fraction <50%),
- atrial fibrillation, and
- moderate to severe cardiac valve disease; and
- SYNTAX score of zero
Similarly, patients were excluded with
- infection,
- acute stress, or chronic systemic inflammatory disease and
- those who had been receiving medications affecting the number of leukocytes .
Thirty patients were excluded from the study because the coronary angiograms revealed normal coronary arteries (SYNTAX score of 0). All the participants included in the study were informed about the study, and they voluntarily consented to participate. The Serum PTX-3 level was measured on blood samples collected after 12-hour fast just prior to coronary angiography and kept at −80°C until the assays were performed. PTX3 was measured by enzyme immunoassay (EIA) using quantitative kit (human PTX-3/TSG-14 immunoassay, DPTX30; R&D Systems, Inc, Minneapolis, MN). The intra-assay and interassay coefficients of variation ranged from 3.8% to 4.4% and 4.1% to 6.1%, respectively (minimum detectable concentration, 0.025 ng/mL). High-sensitivity CRP was measured in serum by EIA (Immage hs-CRP EIA Kit; Beckman Coulter Inc, Brea, CA). Transthoracic echocardiography was performed, and biplane Simpson’s ejection fraction (%) was calculated before coronary angiography. Hypertension was defined as having at least 2 blood pressure measurements greater than 140/90 mm Hg or using antihypertensive drugs, whereas diabetes mellitus (DM) was defined as having at least 2 fasting blood sugar measurements greater than 126 mg/dL or using antidiabetic drugs. Smoking was categorized into current smokers and nonsmokers. Nonsmokers included ex-smokers who had quit smoking for at least 6 months before the study. Body mass index (BMI) values were calculated based on the height and weight of each patient. Medications used before the coronary angiography were noted. The study was approved by the local ethics committee.
SYNTAX and Gensini Scores
To grade the complexity of CAD, the SYNTAX score was used. Each coronary lesion with a stenosis diameter of 50% or greater in vessels of 1.5 mm or greater was scored. Parameters used in the SYNTAX scoring are shown in Table 1. The latest online updated version (2.11) was used in the calculation of the SYNTAX scores (www.syntaxscore.com).17 The SYNTAX score was classified as follows:
- low SYNTAX score (≤22),
- intermediate SYNTAX score (23–32)
- high SYNTAX score (≥33).
Table 1 http://images.journals.lww.com/jinvestigativemed/LargeThumb.00042871-201302000-00007.TT1.jpeg
The severity of CAD was determined by the Gensini score, which
- measures the extent of coronary stenosis according to degree and location.18
In the Gensini scoring system,
- larger segments are more heavily weighted ranging from 0.5 to 5.0
- left main coronary artery × 5;
- proximal segment of the left anterior descending coronary artery [LAD] × 2.5;
- proximal segment of the circumflex artery × 2.5;
- midsegment of the LAD × 1.5;
- right coronary artery distal segment of the LAD,
- posterolateral artery, and obtuse marginal artery × 1;
- and others × 0.5.
The narrowing of the coronary artery lumen is rated
- 2 for 0% to 25% stenosis,
- 4 for 26% to 50%,
- 8 for 51% to 75%,
- 16 for 76% to 90%,
- 32 for 91% to 99%,
- 64 for 100%.
The Gensini index is the sum of the total weights for each segment. All angiographic variables of the SYNTAX and Gensini score were computed by
- 2 experienced cardiologists who were blinded to the procedural data and clinical outcomes.
The final decision was reached by consensus when a conflict occurred.The number of diseased vessels with
- greater than 50% luminal stenosis was scored from 1 to 3 (namely, 1-, 2-, or 3-vessel disease), and
- a lesion greater than 50% in the left main coronary artery was regarded as a 2-vessel disease.
Statistical Analyses
Statistical analyses were conducted with SPSS 17 (SPSS Inc, Chicago, IL) software package program.
Continuous variables were expressed as mean ± SD or median ± interquartile range values, whereas categorical variables were presented as percentages.
The differences between normally distributed numeric variables were evaluated by Student t test or 1-way analysis of variance, whereas
- non–normally distributed variables were analyzed by Mann-Whitney U test or Kruskal-Wallis variance analysis as appropriate.
χ2 Test was used for the comparison of categorical variables. Pearson test was used for correlation analysis.
To determine the independent predictors of “intermediate and high” SYNTAX scores and only “high” SYNTAX scores,
- 2 different sets of univariate and multivariate analyses were performed
- (in the first model SYNTAX cutoff was 22, whereas
- in the second model SYNTAX cutoff was 33).
The standardized parameters that were found to have a significance (P < 0.10) in the univariate analysis were evaluated by stepwise logistic regression analysis.
Ninety-five percent confidence interval (CI) and odds ratio (OR) per SD increase were presented together. Interobserver and intraobserver variability for SYNTAX scores
- was done by Bland-Altman analysis.
An exploratory evaluation of additional cut points was performed using the receiver operating characteristic (ROC) curve analysis.
All the P values were 2-sided, and a P < 0.05 was considered as statistically significant.
RESULTS
Baseline Characteristics
In total, 181 patients (50.2 ± 6.5 years, 52.5% were composed of males) were included in the study. Baseline clinical, angiographic, and laboratory characteristics of the patients
relative to SYNTAX score groups are shown in Table 2. Age, sex, HT, DM, BMI, and medication were not different between the groups. Baseline clinical and laboratory characteristics
of patients according to PTX-3 quartiles are shown in Table 3. The Bland-Altman analysis revealed that the degrees of intraobserver and interobserver variability for SYNTAX score
and Gensini score readings were 5% and 6% for SYNTAX and 8% and 9% for Gensini, respectively.
Table 2 http://images.journals.lww.com/jinvestigativemed/Original.00042871-201302000-00007.TT2.jpeg
Table 3 http://images.journals.lww.com/jinvestigativemed/Original.00042871-201302000-00007.TT3.jpeg
The PTX-3 levels demonstrated an increase from the low SYNTAX group to the high SYNTAX group (r = 0.87, P < 0.001).
The low SYNTAX group had statistically significantly higher PTX-3 levels when compared with the control group (0.50 ± 0.01 vs 0.24 ± 0.01 ng/mL, P < 0.001); similarly,
the PTX-3 levels were higher in the high SYNTAX group than in both
- the intermediate SYNTAX group (0.84 ± 0.08 vs 0.55 ± 0.01 ng/mL, P < 0.001) and
- the low SYNTAX group (0.84 ± 0.08 vs 0.50 ± 0.01 ng/mL, P < 0.001).
- there was no difference in levels of PTX-3 between the low and the intermediate SYNTAX group (0.50 ± 0.01 vs 0.55 ± 0.01 ng/mL, P = 0.09).
On the other hand, there was no difference in levels of hs-CRP between the control and the low SYNTAX group (0.81 ± 0.42 vs 0.86 ± 0.53 mg/dL, P = 0.96).
The intermediate SYNTAX group had statistically significantly higher hs-CRP levels
- compared with the low SYNTAX group (1.3 ± 0.66 vs 0.86 ± 0.53 mg/dL, P = 0.002);
- the hs-CRP levels were not different between the high SYNTAX group
- and the intermediate SYNTAX group. (1.3 ± 0.66 vs 1.3 ± 0.43 mg/dL, P = 0.99).
Univariate correlation analysis revealed a positive correlation between serum PTX-3 levels and hs-CRP levels with
- the SYNTAX and Gensini scores
- for SYNTAX: r = 0.87 [P < 0.001] and r = 0.36 [P = 0.01];
- for Gensini: r = 0.75 [P < 0.001] and r = 0.27 [P = 0.002], (Fig. 1).
In addition to that, the Gensini and SYNTAX scores are found to be well correlated with each other (r = 0.80, P < 0.001).
When the SYNTAX score was taken as continuous variable, multivariate linear regression analysis revealed that
- the SYNTAX score was correlated with PTX-3 and hs-CRP (for PTX-3: β = 0.84 [P < 0.001]; hs-CRP: β =0.08 [P = 0.032]).
Figure 1 http://images.journals.lww.com/jinvestigativemed/Original.00042871-201302000-00007.FF1.jpeg
For determining the predictors of intermediate and high SYNTAX scores and only-high SYNTAX scores,
- 2 different sets of univariate and multivariate analyses were performed among the patients who underwent coronary angiography.
For predicting the intermediate and high SYNTAX scores, the SYNTAX score was dichotomized into
- high (score ≥22) and
- low (<22) groups,
whereas for predicting the only-high SYNTAX scores, the SYNTAX score was dichotomized into
- 2 groups with a score of 33 or greater and a score of less than 33.
In the first multivariate analysis (where SYNTAX cutoff was 22), the parameters showing significance in the univariate analysis
- age,
- sex,
- HT,
- DM,
- low-density lipoprotein cholesterol [LDL-C],
- hs-CRP,
- PTX-3
were evaluated by multivariate analysis to determine the
- independent predictors of intermediate and high SYNTAX scores.
In the univariate analysis, higher values of
- age (OR, 1.5 [95% CI, 1.1–2.0]; P = 0.01),
- LDL-C (OR, 1.3 [95% CI, 0.98–1.8]; P = 0.068),
- hs-CRP (OR, 2.6 [95% CI, 1.8–3.8]; P < 0.001), and
- PTX-3 (OR, 13.6 [95% CI, 6.4–28.9]; P < 0.001)
- were associated with higher SYNTAX scores,
- HT (OR, 0.44 [95% CI, 0.24–0.80]; P = 0.008) and
- DM (OR, 0.48 [95% CI, 0.25–0.91]; P = 0.02)
- were associated with lower SYNTAX scores.
In the multivariate analysis – age, DM, LDL-C, hs-CRP, and PTX-3 – were found to be
- independent predictors of “intermediate to high” SYNTAX score (Table 4).
Increased
- age (OR, 2.5 [95% CI, 1.3–4.8]; P = 0.007),
- LDL-C (OR, 2.8 [95% CI, 1.5–5.2]; P = 0.001),
- hs-CRP (OR, 3.3 [95% CI, 1.8–6.1]; P < 0.001), and
- PTX-3 (OR, 35.4 [95% CI, 10.1–123.6]; P < 0.001)
- were associated with increased SYNTAX scores,
whereas DM (OR, 0.08 [95% CI, 0.02–0.33]; P < 0.001) was associated with lower SYNTAX score (Table 4).
In the second univariate and multivariate analyses (where SYNTAX cutoff was 33),
- the parameters that showed significance in the univariate analysis were age, LDL-C, glucose, hs-CRP, and PTX-3.
- In the univariate analysis, increased
- age (OR, 1.5 [95% CI, 1.0–2.3]; P = 0.05),
- LDL-C (OR, 1.5 [95% CI, 0.97–2.2]; P = 0.07),
- hs-CRP (OR, 1.4 [95% CI, 0.97–2.1]; P = 0.072), and
- PTX-3 (OR, 18.5 [95% CI, 6.6–51.8]; P < 0.001)
- were found to be associated with increased SYNTAX scores.
When these parameters were evaluated with multivariate analysis, only PTX-3 (OR, 18.4 [95% CI, 6.2–54.2]; P < 0.001)
- was found to be an independent predictor for high SYNTAX score (Table 4).
Table 4 http://images.journals.lww.com/jinvestigativemed/Original.00042871-201302000-00007.TT4.jpeg
The ROC curve analysis further revealed that the PTX-3 level was a strong indicator of high SYNTAX score with
- an area under the curve (AUC) of 0.91 (95% CI, 0.86–0.96) (Fig. 2).
The optimal cutoff of PTX-3 for the high SYNTAX score was 0.75 ng/mL.
Sensitivity, specificity, positive predictive value, and negative predictive value to identify high SYNTAX score for the PTX-3 level
- were 90%, 84%, 97%, and 60%, respectively.
- the ROC curve analysis of PTX-3 for intermediate-high SYNTAX score revealed that the AUC value was 0.82 (95% CI, 0.75–0.89).
The optimal threshold of PTX-3 level that
- maximized the combined specificity and sensitivity to predict
- intermediate to high SYNTAX score was 0.73 ng/mL.
For the cutoff value of 0.73 ng/mL, sensitivity, specificity, positive predictive value, and negative predictive value
- to identify intermediate-high SYNTAX score were 56%, 98%, 97%, and 56%, respectively.
Figure 2 http://images.journals.lww.com/jinvestigativemed/Original.00042871-201302000-00007.FF2.jpeg
In the ROC analysis of hs-CRP for high SYNTAX scores, the AUC value was found to be 0.68 (95% CI, 0.59–0.77; P < 0.001).
The optimal threshold of hs-CRP that maximized the combined specificity and sensitivity to predict for high SYNTAX scores was 0.89 mg/dL.
Similarly, the ROC analysis of hs-CRP for the intermediate-high SYNTAX scores revealed an AUC of 0.74 (95% CI, 0.65–0.83; P = 0.001).
The cutoff value of hs-CRP to predict the intermediate-high SYNTAX scores with a maximized sensitivity and specificity was 0.66 mg/dL.
DISCUSSION
In this particular study, we investigated the relationship between the serum PTX-3 level and the severity of CAD
- assessed by SYNTAX and Gensini scores in patients with SAP.
The PTX-3, was significantly higher than control group in the patients with CAD, and the serum PTX-3 levels
- were associated with the SYNTAX and Gensini scores.
When compared with the hs-CRP, the PTX-3 was found to be more tightly associated with the complexity and severity of CAD in the patients with SAP.
Pentraxin 3, an acute-phase reactant that is functionally and structurally similar to CRP,1 is produced both by different kinds of cells such as
- macrophages, dendritic cells, neutrophils, fibroblasts, and vascular endothelial cells.2
- Pentraxin 3 is released following the inflammatory stimuli19; therefore, it may reflect the local inflammatory status in tissues.20
Serum PTX-3 levels were shown to be elevated in patients with
- vasculitis,6 acute myocardial infarction,7,8 and systemic inflammation or sepsis,9 psoriasis, unstable angina pectoris, and heart failure.10–13
Higher PTX3 levels were reported to be associated with worse cardiovascular outcomes
- after acute coronary syndromes,8,21
- in the elderly people without known cardiovascular disease22 and
- associated with overall mortality in patients with stable coronary disease,
- independent of systemic inflammation.14
There are 2 reports investigating the association of PTX-3 level and the atherosclerotic burden.15,16 In one of these reports,
- Knoflach et al.15 took B-mode ultrasonography as the atherosclerosis index.
They did not provide any information about coronary anatomy, and in the other report, Soeki et al.16 evaluated 40 patients who
- underwent coronary angiography and measured their Gensini scores.
However, in none of the studies were the SYNTAX score and Gensini score used together to assess the degree of coronary atherosclerotic burden.
To our knowledge, this is the first report that showed the association of PTX-3 levels with the complexity and severity of CAD assessed by
- SYNTAX and Gensini scores in patients with stable coronary disease.
Chronic low-grade inflammation has been thought to play a major role in the pathogenesis of atherosclerosis.23,24 Previous studies have reported that
- levels of inflammatory markers such as hs-CRP, interleukin 6, and so on were increased in atherosclerosis.25
In the present study, both the SYNTAX and the Gensini scores were found to be correlated with serum PTX-3 and hs-CRP levels,
- which in turn might reflect the degree of inflammation.
The SYNTAX score is an important tool in the classification of complex CAD26 and can give predictive information about short- and long-term outcomes
- in patients with stable CAD who undergo percutaneous coronary intervention.27–30
Although the SYNTAX score is currently used for assessing the angiographic complexity of CAD rather than the severity of coronary atherosclerotic burden,
- because more complex lesions tend to have more atherosclerotic burden,
- the SYNTAX scores may also reflect the severity of coronary atherosclerotic burden.
The Gensini score, a well-known and widely used scoring system to evaluate the severity of CAD,18 was measured and
- found to be well correlated with the SYNTAX score,
- which supports the idea that angiographically more complex lesions tend to have more atherosclerotic burden.
When compared with the hs-CRP,
- the PTX-3 seems to be more tightly associated with coronary disease burden (r = 0.36 vs r = 0.87).
We found out that the serum PTX-3 levels were higher than those in the control group, even in the low SYNTAX group.
On the other side, the serum hs-CRP levels were not different in the control and the low SYNTAX groups.
It was reported that the leukocytes mainly found in the coronary artery lumen are the neutrophils.31
It is also known that PTX-3 is stored in specific granules of neutrophils and released in response to inflammatory signals.32
The reason why serum PTX-3 levels seem more tightly associated with the coronary disease burden
- when compared with serum hs-CRP levels may be the association of the
- on-site presence of neutrophils and local inflammatory signal–triggered release of PTX-3.
On the other hand, some human studies revealed that PTX-3 was produced more in areas of atherosclerosis and may contribute to its pathogenesis.31
Some other studies suggested that PTX-3 may be part of a protective mechanism in
- vascular repair via inhibiting fibroblast growth factor 2 or some other growth factors responsible for smooth muscle proliferation.33,34
But still, the exact role of PTX-3 in the pathophysiology of atherosclerosis seems to be obscure for the time being. It is well established that atherosclerosis
has an inflammatory background in most of the cases. In addition to that, high blood CRP level is known as an indicator of future cardiovascular disease risk
even in healthy individuals.35 According to the results of univariate and multivariate analyses, for intermediate and high SYNTAX scores,
- age, DM, LDL-C, hs-CRP, and PTX-3 were found to be independent predictors, whereas for the presence of
- high SYNTAX score, only PTX-3 was found to be an independent predictor.
Because of the tighter association with atherosclerotic burden and the on-site vascular presence,
- PTX-3 may be a promising candidate marker for vascular inflammation and future cardiovascular events.
LIMITATIONS
The major limitation of the current study is the number of patients included. It would be better to include more patients to increase the statistical power.
Besides, the SYNTAX and Gensini scores give us an idea about the complexity and severity of coronary atherosclerosis; however,
with coronary angiography alone, it is not possible to understand the extent of coronary plaque. In addition to that, the coronary anatomy of the
control group was not known, which was another limitation. Our selected population was free of other confounders of systemic inflammation, and
we did not have data about inflammatory markers other than hs-CRP, such as interleukin 6, tumor necrosis factor α, and so on, which may be accepted
as a limitation. Another limitation of the current study is that because there was no long-term follow-up of the patients, it did not provide any prognostic
data in terms of future cardiovascular events.
CONCLUSIONS
Pentraxin 3, a novel inflammatory marker, is associated with the complexity and severity of the CAD assessed by the SYNTAX and the Gensini scores in patients with SAP and seems to be more tightly associated with coronary atherosclerotic burden than hs-CRP.
REFERENCES
1. Hansson GK. Inflammation, atherosclerosis, and coronary artery disease. N Engl J Med. 2005; 352: 1685–1695.
2. Brown DW, Giles WH, Croft JB. White blood cell count: an independent predictor of coronary heart disease mortality among a national cohort. J Clin Epidemiol. 2001; 54: 316–322.
3. Kannel WB, Anderson K, Wilson PW. White blood cell count and cardiovascular disease. Insights from the Framingham Study. JAMA. 1992; 267: 1253–1256.
4. Muscari A, Bozzoli C, Puddu GM, et al.. Association of serum C3 levels with the risk of myocardial infarction. Am J Med. 1995; 98: 357–364.
5. Ridker PM, Cushman M, Stampfer MJ, et al.. Inflammation, aspirin, and the risk of cardiovascular disease in apparently healthy men. N Engl J Med. 1997; 336: 973–979.
6. Fazzini F, Peri G, Doni A, et al.. PTX3 in small-vessel vasculitides: an independent indicator of disease activity produced at sites of inflammation. Arthritis Rheum. 2001; 44: 2841–2850.
7. Peri G, Introna M, Corradi D, et al.. PTX3, A prototypical long pentraxin, is an early indicator of acute myocardial infarction in humans. Circulation. 2000; 102: 636–641.
8. Latini R, Maggioni AP, Peri G, et al.. Prognostic significance of the long pentraxin PTX3 in acute myocardial infarction. Circulation. 2004; 110: 2349–2354.
9. Muller B, Peri G, Doni A, et al.. Circulating levels of the long pentraxin PTX3 correlate with severity of infection in critically ill patients. Crit Care Med. 2001; 29: 1404–1407.
10. Bevelacqua V, Libra M, Mazzarino MC, et al.. Long pentraxin 3: a marker of inflammation in untreated psoriatic patients. Int J Mol Med. 2006; 18: 415–423.
11. Inoue K, Sugiyama A, Reid PC, et al.. Establishment of a high sensitivity plasma assay for human pentraxin3 as a marker for unstable angina pectoris. Arterioscler Thromb Vasc Biol. 2007; 27: 161–167.
12. Suzuki S, Takeishi Y, Niizeki T, et al.. Pentraxin 3, a new marker for vascular inflammation, predicts adverse clinical outcomes in patients with heart failure. Am Heart J. 2008; 155: 75–81.
13. Matsubara J, Sugiyama S, Nozaki T, et al.. Pentraxin 3 is a new inflammatory marker correlated with left ventricular diastolic dysfunction and heart failure with normal ejection fraction. J Am Coll Cardiol. 2011; 57: 861–869.
14. Dubin R, Li Y, Ix JH, et al.. Associations of pentraxin-3 with cardiovascular events, incident heart failure, and mortality among persons with coronary heart disease: data from the Heart and Soul Study. Am Heart J. 2012; 163: 274–279.
16. Soeki T, Niki T, Kusunose K, et al.. Elevated concentrations of pentraxin 3 are associated with coronary plaque vulnerability. J Cardiol. 2011; 58: 151–157.
17. SYNTAX working group. SYNTAX score calculator. Available at http://www.syntaxscore.com. Accessed May 20, 2012.
18. Gensini GG. A more meaningful scoring system for determining the severity of coronary heart disease. Am J Cardiol. 1983; 51: 606.
20. Mantovani A, Garlanda C, Bottazzi B, et al.. The long pentraxin PTX3 in vascular pathology. Vascul Pharmacol. 2006; 45: 326–330.
21. Matsui S, Ishii J, Kitagawa F, et al.. Pentraxin 3 in unstable angina and non-ST-segment elevation myocardial infarction. Atherosclerosis. 2010; 210: 220–225.
22. Jenny NS, Arnold AM, Kuller LH, et al.. Associations of pentraxin 3 with cardiovascular disease and all-cause death: the Cardiovascular Health Study. Arterioscler Thromb Vasc Biol. 2009; 29: 594–599.
26. Serruys PW, Morice MC, Kappetein AP, et al.. Percutaneous coronary intervention versus coronary-artery bypass grafting for severe coronary artery disease. N Engl J Med. 2009; 360: 961–972.
27. van Gaal WJ, Ponnuthurai FA, Selvanayagam J, et al.. The SYNTAX score predicts peri-procedural myocardial necrosis during percutaneous coronary intervention. Int J Cardiol. 2009; 135: 60–65.
28. Lemesle G, Bonello L, de Labriolle A, et al.. Prognostic value of the SYNTAX score in patients undergoing coronary artery bypass grafting for three-vessel coronary artery disease. Catheter Cardiovasc Interv. 2009; 73: 612–617.
29. Capodanno D, Di Salvo ME, Cincotta G, et al.. Usefulness of the SYNTAX score for predicting clinical outcome after percutaneous coronary intervention of unprotected left main coronary artery disease. Circ Cardiovasc Interv. 2009; 2: 302–308.
30. Kim YH, Park DW, Kim WJ, et al.. Validation of SYNTAX (Synergy between PCI with Taxus and Cardiac Surgery) score for prediction of outcomes after unprotected left main coronary revascularization. JACC Cardiovasc Interv. 2010; 3: 612–623.
32. Jaillon S, Peri G, Delneste Y, et al.. The humoral pattern recognition receptor PTX3 is stored in neutrophil granules and localizes in extracellular traps. J Exp Med. 2007; 204: 793–804.
33. Inforzato A, Baldock C, Jowitt TA, et al.. The angiogenic inhibitor long pentraxin PTX3 forms an asymmetric octamer with two binding sites for FGF2. J Biol Chem. 2010; 285: 17681–17692.
34. Camozzi M, Zacchigna S, Rusnati M, et al.. Pentraxin 3 inhibits fibroblast growth factor 2–dependent activation of smooth muscle cells in vitro and neointima formation in vivo. Arterioscler Thromb Vasc Biol. 2005; 25: 1837–1842.
35. Koenig W, Sund M, Frohlich M, et al.. C-Reactive protein, a sensitive marker of inflammation, predicts future risk of coronary heart disease in initially healthy middle-aged men: results from the MONICA (Monitoring Trends and Determinants in Cardiovascular Disease) Augsburg Cohort Study, 1984 to 1992. Circulation. 1999; 99: 237–242.
Keywords: pentraxin 3; coronary artery disease; SYNTAX score; hs-CRP; inflammation
This is not the only recent finding that adds to the ability to evaluate these patients. An as yet unpublished paper, expected to be published soon reports on
QRS fragmentation as a Prognostic test in Acute Coronary Syndrome, and this reviewer expects the work to have a high impact. The authors state that
QRS complex fragmentation is a promising bed-side test for assessment of prognosis in those patients. Presence of fragmented QRS in surface ECG during ACS
- represents myocardial scar or fibrosis and reflect severity of coronary lesions and a correlation between fQRS and depression of Lv function is established.
There are still other indicators that need to be considered, such as the mean arterial blood pressure.
There has been review and revisions of the guidelines for treatment of UA/NSTEMI within the last year, with differences being resolved among the Europeans and US.
Guidelines Updated for Unstable Angina/Non-ST Elevation Myocardial Infarction
According to the current study by Jneid and colleagues, new evidence is available on the management of unstable angina. This report replaces the 2007 American College of Cardiology Foundation/American Heart Association (ACC/AHA) Guidelines for the Management of Patients With Unstable Angina/Non–ST-Elevation Myocardial Infarction (UA/NSTEMI) that were updated by the 2011 guidelines.
This guideline was reviewed by
- 2 official reviewers each nominated by the ACCF and the AHA, as well as
- 1 or 2 reviewers each from the American College of Emergency Physicians; the Society for Cardiovascular Angiography and Interventions; and the Society of Thoracic Surgeons; and
- 29 individual content reviewers, including members of the ACCF Interventional Scientific Council.
The recommendations in this focused update are considered current
- until they are superseded in another focused update or the full-text guideline is revised, and are official policy of both the ACCF and the AHA.
STUDY SYNOPSIS AND PERSPECTIVE
American cardiology societies have caught up with the European Society of Cardiology by
- issuing their second update to the UA/NSTEMI guidelines in 18 months,
- with the 2012 focused update replacing the 2011 guidelines [1].
The new recommendations include ticagrelor (Brilinta) as one of the options for antiplatelet therapy alongside prasugrel (Effient) and clopidogrel, bringing them in line with European.
The European guidance, however, gave precedence to the new antiplatelets over clopidogrel, whereas the American update “places ticagrelor on an equal footing with the other two antiplatelets available–
this is the main reason for the update,” lead author Dr Hani Jneid (Baylor College of Medicine, Houston, TX), told heartwire . “Doctors now have a choice for second-line therapy after aspirin, depending on
- the patient’s clinical scenario,
- physician preference, and cost,”
- now that clopidogrel is available generically.
The US decision to recommend
- first prasugrel–in its 2011 update to the UA/NSTEMI guidelines–and
- now ticagrelor as equivalent antiplatelet therapy choices to clopidogrel after aspirin
- puts it somewhat at odds with the Europeans,
- who reserve clopidogrel use for those who cannot take the newer agents.
The reason for the Americans differing stance is that because while they are faster acting and more potent–
- the cost-effectiveness of the new agents is not known.
- it isn’t clear how the efficacy observed in pivotal clinical trials of these agents is going to translate into real-world benefit,
- and issues such as bleeding with prasugrel and compliance with a twice-daily drug such as ticagrelor remain concerns.
Bulk of 2012 Update on How to Use Ticagrelor
The 2012 ACCF/AHA focused update for the management of UA/NSTEMI stresses that
- all patients at medium/high risk should receive dual antiplatelet therapy on admission,
- with aspirin being first-line, indefinite therapy.
The bulk of the update centers on how to use ticagrelor which–
- like prasugrel or clopidogrel–
- can be added to aspirin for up to 12 months (or longer, at the discretion of the treating clinician).
Jneid notes it’s important to remember that prasugrel can only be used in the cath lab
- in patients undergoing percutaneous coronary intervention (PCI),
- whereas ticagrelor, like clopidogrel, can be used in medically managed or PCI patients.
And he emphasizes that, in line with the FDA’s black-box warning on ticagrelor,
The 81-mg aspirin dose is also considered a reasonable option in preference to a higher maintenance dose of 325 mg in
- any acute coronary syndrome (ACS) patient following PCI, he adds, as
- this strategy is believed to result in equal efficacy and lower bleeding risk.
With regard to how long antiplatelet therapy should be stopped before planned cardiac surgery, the recommendation is
- five days for ticagrelor–the same as that advised for clopidogrel.
- and seven days prior to surgery for prasugrel.
Jneid also highlights other important recommendations from the 2011 focused update carried over to 2012:
It is “reasonable” to proceed with cardiac catheterization and revascularization within
- 12–24 hours of admission in initially stable, very high-risk patients with ACS.
An invasive strategy is “reasonable” in patients with
- mild and moderate chronic kidney disease.
In those with diabetes hospitalized with ACS, insulin use should target glucose levels <180 mg/dL,
- a less-intensive reduction than previously recommended.
Platelet function or genotype testing for clopidogrel resistance are both considered “reasonable”
- if clinicians think the results will alter management,
- but Jneid acknowledged that “there is not much evidence to support these assays” .
Committee Encourages Participation in Registries
Jneid observes that unstable angina and NSTEMI are “very common” conditions that carry a high risk of death and recurrent heart attacks,
- which is why “the AHA and ACCF constantly update their guidelines so that physicians can provide patients with
- the most appropriate, aggressive therapy with the goal of improving health and survival.”
To this end, he notes that the writing panel encourages
- clinicians and hospitals to participate in quality-of-care registries designed
- to track and measure outcomes, complications, and
- adherence to evidence-based medicines.
Conflicts of interest for the writing committee are listed in the paper.
References
Jneid H, Anderson JL, Wright SR, et al. 2012 ACCF/AHA focused update on the guideline for the management of patients with unstable angina/non-ST elevation myocardial infarction (Updating the 2007 guideline and replacing the 2011 focused update): A report of the ACCF/AHA.
Circulation 2012; Available at: http://circ.ahajournals.org/ http://dx.doi.org/10.1161/CIR0b013e3182566fleo
source http://www.medscape.org
Related articles
FDA Pending 510(k) for The Latest Cardiovascular Imaging Technology
New Definition of MI Unveiled, Fractional Flow Reserve (FFR)CT for Tagging Ischemia
Coronary CT Angiography versus Standard Evaluation in Acute Chest Pain
Dilated Cardiomyopathy: Decisions on implantable cardioverter-defibrillators (ICDs) using left ventricular ejection fraction (
LVEF) and Midwall Fibrosis: Decisions on Replacement using late gadolinium enhancement cardiovascular MR (LGE-CMR)
The potential contribution of informatics to healthcare is more than currently estimated
PCI Outcomes, Increased Ischemic Risk associated with Elevated Plasma Fibrinogen not Platelet Reactivity
Amplifying Information Using S-Clustering and Relationship to Kullback-Liebler Distance: An Application to Myocardial Infarction
Percutaneous Endocardial Ablation of Scar-Related Ventricular Tachycardia
Vascular Medicine and Biology: CLASSIFICATION OF FAST ACTING THERAPY FOR PATIENTS AT HIGH RISK FOR MACROVASCULAR EVENTS Macrovascular Disease – Therapeutic Potential of cEPCs
Ablation Devices Market to 2016 – Global Market Forecast and Trends Analysis by Technology, Devices & Applications
Endothelial Dysfunction, Diminished Availability of cEPCs, Increasing CVD Risk for Macrovascular Disease – Therapeutic Potential of cEPCs
Positioning a Therapeutic Concept for Endogenous Augmentation of cEPCs — Therapeutic Indications for Macrovascular Disease: Coronary, Cerebrovascular and Peripheral
Cardiovascular Outcomes: Function of circulating
Endothelial Progenitor Cells (cEPCs): Exploring Pharmaco-therapy targeted at Endogenous Augmentation of cEPCs
Competition in the Ecosystem of Medical Devices in Cardiac and Vascular Repair: Heart Valves, Stents, Catheterization Tools and Kits for Open Heart and
Minimally Invasive Surgery (MIS)
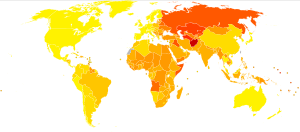
Age-standardised disability-adjusted life year (DALY) rates from Cardiovascular diseases by country (per 100,000 inhabitants). (Photo credit: Wikipedia)

English: Cardiovascular disease: PAD therapy with stenting Deutsch: PAVK Therapie: Kathetertherapie mit stenting (Photo credit: Wikipedia)
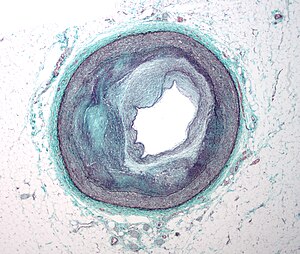
Micrograph of an artery that supplies the heart with significant atherosclerosis and marked luminal narrowing. Tissue has been stained using Masson’s trichrome. (Photo credit: Wikipedia)
Read Full Post »











