Treatment, Prevention and Cost of Cardiovascular Disease: Current & Predicted Cost of Care and the Potential for Improved Individualized Care Using Clinical Decision Support Systems
Author, and Content Consultant to e-SERIES A: Cardiovascular Diseases: Justin Pearlman, MD, PhD, FACC
Author and Curator: Larry H Bernstein, MD, FACP
and
Curator: Aviva Lev-Ari, PhD, RN
This article has the following FIVE parts:
1. Forecasting the Impact of Heart Failure in the United States : A Policy Statement From the American Heart Association
2. A Case Study from the GENETIC CONNECTIONS — In The Family: Heart Disease Seeking Clues to Heart Disease in DNA of an Unlucky Family
3. Arterial Stiffness and Cardiovascular Events : The Framingham Heart Study
4. Arterial Elasticity in Quest for a Drug Stabilizer: Isolated Systolic Hypertension
caused by Arterial Stiffening Ineffectively Treated by Vasodilatation Antihypertensives
5. Clinical Decision Support Systems: Realtime Clinical Expert Support — Biomarkers of Cardiovascular Disease : Molecular Basis and Practical Considerations
PA Heidenreich, NM Albert, LA Allen, DA Bluemke, J Butler, et al. Circulation: Heart Failure 2013;6.
Print ISSN: 1941-3289, Online ISSN: 1941-3297.
Heart failure (HF) poses a major burden on productivity and cost of national healthcare expenditures
- among older Americans, more are hospitalized for HF than for any other medical condition.
As the population ages, the prevalence of HF is expected to increase.
The purpose of this report is to
- provide an in-depth look at how the changing demographics in the United States will impact the prevalence and cost of care for HF for different US populations.
Projections of HF Prevalence
Prevalence estimates for HF were determined from
Projections of the US Population With HF From 2010 to 2030 for Different Age Groups
|
Year
|
All ages
|
18-44 y
|
45-64 y
|
65-79 y
|
> 80
|
| 2012 |
5 813 262 |
396 578 |
1 907 141 |
2 192 233 |
1 317 310 |
| 2015 |
6 190 606 |
402 926 |
1 949 669 |
2 483 853 |
1 354 158 |
| 2020 |
6 859 623 |
417 600 |
1 974 585 |
3 004 002 |
1 463 436 |
| 2025 |
7 644 674 |
434 635 |
1 969 852 |
3 526 347 |
1 713 840 |
| 2030 |
8 489 428 |
450 275 |
2 000 896 |
3 857 729 |
2 180 528 |
Future Costs of HF
The future costs of HF were estimated by methods developed by the American Heart Association
- project the prevalence and costs of HF from 2012 to 2030
- factor out the costs attributable to comorbid conditions.
The model does this by assuming that
(1) HF prevalence percentages will remain constant by age, sex, and race/ethnicity;
(2) the costs of technological innovation will rise at the current rate.
HF prevalence and costs (direct and indirect) were projected using the following steps:
1. HF prevalence and average cost per person were estimated by age group (18–44, 45–64, 65–79, ≥80 years), gender (male, female), and race/ethnicity (white non-Hispanic, white Hispanic, black, other) [32]. The initial HF cost per person and rate of increase in cost was determined for each demographic group, as a percentage of total healthcare expeditures.
2. Inflation is separately addressed by correcting dollar values from Medical Expenditure Panel Survey (MEPS) to 2010 dollars.
3. Nursing home spending triggered an adjustment. The estimates project the incremental cost of care attributable to heart failure (HF).
4. Total HF population prevalence and costs were projected by multiplying the US Census–projected population of each demographic group by the percentage prevalence and average cost
5. The total work loss and home productivity loss costs were generated by multiplying per capita work days lost attributable to HF by (1) prevalence of HF, (2) the probability of employment given HF (for work loss costs only), (3) mean per capita daily earnings, and (4) US Census population projection counts.
Projections of Indirect Costs
Indirect costs of lost productivity from morbidity and premature mortality were estimated as detailed below.
Morbidity costs represent the value of lost earnings attributable to HF and include loss of work among
- currently employed individuals and those too sick to work, as well as
- home productivity loss, which is the value of household services performed by household members who do not receive pay for the services.
Total Costs Attributable to Heart Failure (HF)
Projections of Total Cost of Care ($ Billions) for HF for Different Age Groups of the US Population
| Year |
All |
18–44 |
45–64 |
65–79 |
≥ 80 |
| 2012 |
|
|
|
|
|
| Medical |
20.9 |
0.33 |
3.67 |
8.46 |
8.42 |
| Indirect: Morbidity |
5.42 |
0.52 |
1.92 |
2.05 |
0.93 |
| Indirect: Mortality |
4.35 |
0.66 |
2.53 |
0.98 |
0.18 |
| Total |
30.7 |
1.51 |
8.12 |
11.5 |
9.53 |
| 2020 |
|
|
|
|
|
| Medical |
31.1 |
0.43 |
4.58 |
14.2 |
11.8 |
| Indirect: Morbidity |
7.09 |
0.66 |
2.20 |
3.11 |
1.12 |
| Indirect: Mortality |
5.39 |
0.79 |
2.89 |
1.49 |
0.22 |
| Total |
43.6 |
1.88 |
9.67 |
18.8 |
13.2 |
| 2030 |
|
|
|
|
|
| Medical |
53.1 |
0.59 |
5.86 |
23.3 |
23.4 |
| Indirect: Morbidity |
9.80 |
0.91 |
2.54 |
4.48 |
1.87 |
| Indirect: Mortality |
6.84 |
0.98 |
3.32 |
2.16 |
0.37 |
| Total |
69.7 |
2.48 |
11.7 |
29.9 |
25.6 |
Excludes HF care costs that have been attributed to comorbid conditions.
Cost of Care
Total medical costs are projected to increase from $20.9 billion in 2012 to $53.1 billion in 2030, a 2.5-fold increase. Assuming continuation of current hospitalization practices, the majority (80%) of the costs stem from
- hospitalization. Also, the majority of increase is from directs costs. Indirect costs are expected to rise as well, but at a lower rate, from $9.8 billion to $16.6 billion, an increase of 69%.
Direct costs (cost of medical care) are expected to increase at a faster rate than indirect costs because of premature deaths and lost productivity.
The total cost of HF (direct and indirect costs) is expected to increase in 2030 from the current $30.7 billion to at least $69.8 billion. This will amount to $244 for every US adult in 2030.
Thus the burden of HF for the US healthcare system will grow substantially during the next 18 years if current trends continue.
It is estimated that
- by 2030, the prevalence of HF in the United States will increase by 25%, to 3.0%.
- >8 million people in the US (1 in every 33) will have HF by 2030.
- the projected total direct medical costs of HF between 2012 and 2030 (in 2010 dollars) will increase from $21 billion to $53 billion.
- Total costs, including indirect costs for HF, are estimated to increase from $31 billion in 2012 to $70 billion in 2030.
- If one assumes all costs of cardiac care for HF patients are attributable to HF
(no cost attribution to comorbid conditions), the 2030 projected cost estimates of treating patients with HF will be 3-fold higher ($160 billion in direct costs).
Projections can be lowered if action is taken to reduce the health and economic burden of HF. Strategies, plans, and implementation to prevent HF and improve the efficiency of care are needed.
Causes and Stages of HF
If the projections for accelerating HF costs are to be avoided, attention to the different causes of HF and their risk factors is warranted.
HF is a clinical syndrome that results from a variety of cardiac disorders
- idiopathic dilated cardiomyopathy
- cardiac valvular disease
- pericarditis or pericardial effusion
- ischemic heart disease
- primary or secondary hypertension
- renovascular disease
- advanced liver disease with decreased venous return
- pulmonary hypertension
- prolonged hypoalbuminemia with generalized interstitial edema
- diabetic nephropathy
- heart muscle infiltration disease such as primary or secondary amyloidosis
- myocarditis
- rhythm disorders
- congenital diseases
- accidental trauma (war, chest trauma)
- toxicities (methamphetamine, cocaine, heavy metals, chemotherapy)
HF generally causes symptoms:
- shortness of breath
- fatigue
- swelling (edema)
- inability to lay flat (orthopnea, paroxysmal nocturnal dyspnea)
- possibly cough, wheezing
In the Western world the predominant causes of HF are:
- coronary artery disease
- valvular disease
- hypertension
- viral, alcohol, methamphetamine or other drug toxicity cardiomyopathy
- stress (catechol toxicity, takotsubo “broken heart” cardiomyopathy)
- atrial fibrillation/rapid heart rates
- thyroid disease
In 2001, the American College of Cardiology and AHA practice guidelines for chronic HF promoted a classification system that encompasses 4 stages of HF.
- Stage A: Patients at high risk for developing HF in the future but no functional or structural heart disorder.
- Stage B: a structural heart disorder but no symptoms.
- Stage C: previous or current symptoms of heart failure, manageable with medical treatment.
- Stage D: advanced disease requiring hospital-based support, a heart transplant or palliative care.
Stages A and B are considered precursors to the clinical HF and are meant
- to alert healthcare providers to known risk factors for HF and
- the available therapies aimed at mitigating disease progression.
Stage A patients have risk factors for HF hypertension, atherosclerotic heart disease, and/or diabetes mellitus.
Patients with stage B are asymptomatic patients who have developed structural heart disease from a variety of potential insults to the heart muscle such as myocardial infarction or valvular heart disease.
Stages C and D represent the symptomatic phases of HF, with stage C manageable and stage D failing medical management, resulting in marked symptoms at rest or with minimal activity despite optimal medical therapy.
Therapeutic interventions include:
- dietary salt restriction and diuretics
- medications known to prolong survival (beta blockers, ACE inhibitors, aldosterone inhibitors)
- implantable devices such as pacemakers and defibrillators
- stoppage of tobacco, toxic drugs, excess alcohol
Classic demographic risk factors for the development of HF include
- older age, male gender, ethnicity, and low socioeconomic status.
- comorbid disease states contribute to the development of HF
- Ischemic heart disease
- Hypertension
Diabetes mellitus, insulin resistance, and obesity are also linked to HF development,
- with diabetes mellitus increasing the risk of HF by ≈2-fold in men and up to 5-fold in women.
Smoking remains the single largest preventable cause of disease and premature death in the United States.
Translation of Scientific Evidence into Clinical Practice
In multiple studies, failures to apply evidence-based management strategies are blamed for avoidable hospitalizations and/or deaths from HF
Improved implementation of guidelines can delay, mitigate or prevent the onset of HF, and improve survival. Performance improvement programs have facilitated the implementation of evidence-based therapies in both hospital and ambulatory care settings.
Care transition programs by hospitals have become more widespread
- in an effort to reduce avoidable readmissions.
The interventions used by these programs include
- initiating discharge planning early in the course of hospital care,
- actively involving patients and families or caregivers in the plan of care,
- providing new processes and systems that ensure patient understanding of the plan of care before discharge from the hospital, and
- improving quality of care by continually monitoring adherence to national evidence-based guidelines with appropriate adaptations for individual differences in needs and responses.
In multiple studies,adherence to the HF plan of care was associated with reduced all-cause mortality as well as HF hospitalization.
It is anticipated that care transition programs may increase appropriate admissions while decreasing inappropriate admissions
This would have a potentially benenficial impact on the 30-day all-cause readmission rate that has become
- a focus of public reporting in pay for performance.
More than a quarter of Medicare spending occurs in the last year of life, and
- the costs of care during the last 6 months for a patient with HF have been increasing (11% from 2000 to 2007).
Improving end-of-life care cost effectiveness for patients with stage D HF will require ongoing
- improved prediction of outcomes
- integration of multiple aspects of care
- educated examination of alternatives and priorities
- improved decision-making
- unbiased allocation of resources and coverage for this process rather than unbalanced coverage favoring catastrophic care
Palliative care, including formal hospice care, is increasingly advocated for patients with advanced HF.
Offering palliative care to patients with HF may lead to
- more conservative (and less expensive) treatment
- consistent with many patients’ goals for care
The use of hospice services is growing among the HF population,
- HF now the second most common reason for entering hospice
- but hospice declaration may impose automated restrictions on care that can impose an impediment to election of hospice
A recent study of patients in hospice care found that
- patients with HF were more likely than patients with cancer to use hospice services longer than 6 months or to be discharged from hospice care alive.
Highlights:
1. Increasing incidence and costs of care for heart failure projected from 2012 to 2030
2. Direct costs rising at greater rate than indirect costs
3. American Heart Association has defined 4 stages of HF, the last 2 of which are advanced
4. Stages C & D are clinically overt and contribute to rehospitalization
5. Stage D accounts for a significant use of end-of-life hospice care
6. There are evidence-based guidelines for the provision of coordinated care that are not widely applied at present
Basic questions raised:
1. If stages A & B are under the radar, then what measures can best trigger the use of evidence-based guidelines for care?
2. Why are evidence-based guidelines commonly not deployed?
- Flaws in the “evidence” due to bias, design errors, limted ability to extrapolate to the patients it should address
- Delays in education, convincing of caretakers, and deployment
- Inadequate resources
- Financial or other disincentives
The arguments for introducing coordinated care and for evidence-based guidelines is strong.
Arguments AGAINST slavish imposition of evidence based medicine include genetic individuality (what is best on average is not necessarily best for each genetically and behaviorly distinct individual). Strict adherence to evidence-based guidelines also stifles innovative explorations. None-the-less, deviations from evidence-based plans should be cautious, well-documented, and well-informed, not due to mal-aligned incentives, ignorance, carelessness or error.
The question of when and how to intervene most cost effectively is unanswered. If some patients are salt-sensitive as a contribution to the prevalence of hypertension and heart failure, should EVERYONE be salt restricted or should there be a more concerted effort to define who is salt sensitive? What if it proved more cost-effective to restrict salt intake for everyone, even though many might be fine with high sodium intake, and some might even benefit from or require high sodium intake? Is it reasonable to impose costs, hurdles, even possible harm on some as a cheaper way to achieve “greater good”?
These issues are highly relevant to the proposed emphasis on holistic solutions.
2. A Case Study from the GENETIC CONNECTIONS — In The Family: Heart Disease Seeking Clues to Heart Disease in DNA of an Unlucky Family
By GINA KOLATA 2013.05.13 New York Times
Scientists are studying the genetic makeup of the Del Sontro family for
- telltale mutations or aberrations in the DNA.
Robin Ashwood, one of Mr. Del Sontro’s sisters, found out she had extensive heart disease even though her electrocardiograms was normal. Six of her seven siblings also have heart disease, despite not having any of the traditional risk factors. Then, after a sister, just 47 years old, found out she had advanced heart disease, Mr. Del Sontro, then 43, went to a cardiologist. An X-ray of his arteries revealed the truth. Like his grand-father, his mother, his four brothers and two sisters, he had heart disease.
Now he and his extended family have joined an extraordinary federal research project that is using genetic sequencing to find factors that increase the risk of heart disease beyond the usual suspects — high cholesterol, high blood pressure, smoking and diabetes.“We don’t know yet how many pathways there are to heart disease,” said Dr. Leslie Biesecker, who directs the study Mr. Del Sontro joined. “That’s the power of genetics. To try and dissect that.”
“I had bought the dream: if you just do the right things and eat the right things, you will be O.K.,” said Mr. Del Sontro, whose cholesterol and blood pressure are reassuringly low.
3. Arterial Stiffness and Cardiovascular Events : The Framingham Heart Study
GF Mitchell, Shih-Jen Hwang, RS Vasan, MG Larson.
Circulation. 2010;121:505-511. http://circ.ahajournals.org/content/121/4/505
http://dx.doi.org/10.1161/CIRCULATIONAHA.109.886655
Various measures of arterial stiffness and wave reflection have been proposed as cardiovascular risk markers.
Prior studies have not assessed relations of a comprehensive panel of stiffness measures to prognosis.
First-onset major cardiovascular disease events in relation to arterial stiffness
- pulse wave velocity [PWV]
- wave reflection
- augmentation index
- carotid-brachial pressure amplification)
- central pulse pressure
were analyzed in 2232 participants (mean age, 63 years; 58% women) in the Framingham Heart Study by a proportional hazards model. During median follow-up of 7.8 (range, 0.2 to 8.9) years,
- 151 of 2232 participants (6.8%) experienced an event.
In multivariable models adjusted for
- age
- sex
- systolic blood pressure
- use of antihypertensive therapy
- total and high-density lipoprotein cholesterol concentrations
- smoking
- presence of diabetes mellitus
higher aortic PWV was associated with a 48% increase in cardiovascular disease risk (95% confidence interval, 1.16 to 1.91 per SD; P 0.002).
After PWV was added to a standard risk factor model, integrated discrimination improvement was 0.7% (95% confidence interval, 0.05% to 1.3%; P 0.05).
In contrast,
- augmentation index,
- central pulse pressure, and
- pulse pressure amplification
were not related to cardiovascular disease outcomes in multivariable models.
Higher aortic stiffness assessed by PWV
- is associated with increased risk for a first cardiovascular event.
Aortic PWV improves risk prediction when added to standard risk factors and may represent
- a valuable biomarker of cardiovascular disease risk
We shall here visit a recent article by Justin D. Pearlman and Aviva Lev-Ari, PhD, RN, on
Pros and Cons of Drug Stabilizers for Arterial Elasticity as an Alternative or Adjunct to Diuretics and Vasodilators in the Management of Hypertension, titled
4. Hypertension and Vascular Compliance: 2013 Thought Frontier – An Arterial Elasticity Focus
http://pharmaceuticalintelligence.com/2013/05/11/arterial-elasticity-in-quest-for-a-drug-stabilizer-isolated-systolic-hypertension-caused-by-arterial-stiffening-ineffectively-treated-by-vasodilatation-antihypertensives/
Speaking at the 2013 International Conference on Prehypertension and Cardiometabolic Syndrome, meeting cochair Dr Reuven Zimlichman (Tel Aviv University, Israel) argued that there is a growing number of patients for whom the conventional methods are inappropriate for
- the definitions of hypertension
- the risk-factor tables used to guide treatment
Most antihypertensives today work by producing vasodilation or decreasing blood volume which may be
- ineffective treatments for patients in whom average arterial diameter and circulating volume are not the causes of hypertension and as targets of therapy may promote decompensation
In the future, he predicts, “we will have to start looking for a totally different medication that will aim to
- improve or at least to stabilize arterial elasticity: medication that might affect factors that determine the stiffness of the arteries, like collagen, like fibroblasts.
Those are not the aim of any group of antihypertensive medications today.”
Zimlichman believes existing databases could be used to develop algorithms that focus on
- inelasticity as a mechanism of hypertensive disease
He also points out that
- ambulatory blood-pressure-monitoring devices can measure elasticity
http://www.theheart.org/article/1502067.do
A related article was published on the relationship between arterial stiffening and primary hypertension.
Arterial stiffening provides sufficient explanation for primary hypertension.
KH Pettersen, SM Bugenhagen, J Nauman, DA Beard, SW Omholt.
By use of empirically well-constrained computer models describing the coupled function of the baroreceptor reflex and mechanics of the circulatory system, we demonstrate quantitatively that
- arterial stiffening seems sufficient to explain age-related emergence of hypertension.
Specifically,
- the empirically observed chronic changes in pulse pressure with age
- the capacity of hypertensive individuals to regulate short-term changes in blood pressure becomes impaired
The results suggest that a major target for treating chronic hypertension in the elderly may include
- the reestablishment of a proper baroreflex response.
http://arxiv.org/abs/1305.0727v2?goback=%2Egde_4346921_member_240018699
5. Clinical Decision Support Systems: Realtime Clinical Expert Support: Biomarkers of Cardiovascular Disease — Molecular Basis and Practical Considerations
RS Vasan. Circulation. 2006;113:2335-2362
http://dx.doi.org/10.1161/CIRCULATIONAHA.104.482570
http://circ.ahajournals.org/content/113/19/2335
Substantial data indicate that CVD is a life course disease that begins with the evolution of risk factors that contribute to
- subclinical atherosclerosis.
Subclinical disease culminates in overt CVD. The onset of CVD itself portends an adverse prognosis with greater
- risks of recurrent adverse cardiovascular events, morbidity, and mortality.
Clinical assessment alone has limitations. Clinicians have used additional tools to aid clinical assessment and to enhance their ability to identify the “vulnerable” patient at risk for CVD, as suggested by a recent National Institutes of Health (NIH) panel.
Biomarkers are one such tool to better identify high-risk individuals, to diagnose disease conditions promptly for diagnosis, prognosis, and treatment guidance.
Biological marker (biomarker): A laboratory test value that is objectively measured and evaluated as an indicator of
- normal biological processes,
- pathogenic processes, or
- pharmacological responses to a therapeutic intervention.
Type 0 biomarker: A marker of the natural history of a disease
- Type 0 correlates longitudinally with known clinical indices/predicts outcomes.
Type I biomarker: A marker that captures the effects of a therapeutic intervention
- Type I assesses an aspect of treatment mechanism of action.
Type 2 biomarker (surrogate end point): A marker intended to predict outcomes on the basis of
- epidemiologic
- therapeutic
- pathophysiologic or
- other scientific evidence.
With biomarkers monitoring disease progression or response to therapy, the patient can serve as his or her own control (follow-up values may be compared to baseline values).
Costs may be less important for prognostic markers when they are largely restricted to people with disease (total cost=cost per person x number to be tested, plus down-stream costs). Some biomarkers (e.g., an exercise stress test) may be used for both diagnostic and prognostic purposes.
Generally there are cost differences in establishing a prognostic value versus diagnostic value of a biomarker:
- prognostic utility typically requires a large sample and a prospective design, whereas
- diagnostic value often can be determined with a smaller sample in a cross-sectional design
Regardless of the intended use, it is important to remember that biomarkers that do not change disease management
- cannot affect patient outcome and therefore
- are unlikely to be cost-effective (judged in terms of quality-adjusted life-years gained).
Typically, for a biomarker to change management, it is important to have evidence that risk reduction strategies should vary with biomarker levels, and/or biomarker-guided management achieves advantages over a management scheme that ignores the biomarker levels.
Typically it means that biomarker levels should be modifiable by therapy.
Gil David and Larry Bernstein have developed, in consultation with Prof. Ronald Coifman, in the Yale University Applied Mathematics Program, a software system that is the equivalent of an intelligent Electronic Health Records Dashboard that
- provides empirical medical reference and
- suggests quantitative diagnostics options.
The current design of the Electronic Medical Record (EMR) is a
linear presentation of portions of the record
- by services
- by diagnostic method, and
- by date
to cite examples.
This allows perusal through a graphical user interface (GUI) that
- partitions the information or necessary reports in a workstation entered by keying to icons.
- presents decision support
Examples of data partitions include:
- history
- medications
- laboratory reports
- imaging
- EKGs
The introduction of a DASHBOARD adds presentation of
- drug reactions
- allergies
- primary and secondary diagnoses, and
- critical information
about any patient the care giver needing access to the record.
A basic issue for such a tool is what information is presented and how it is displayed.
A determinant of the success of this endeavor is if it
- facilitates workflow
- facilitates decision-making process
- reduces medical error.
Continuing work is in progress in extending the capabilities with model datasets, and sufficient data based on the assumption that computer extraction of data from disparate sources will, in the long run, further improve this process.
For instance, there is synergistic value in finding coincidence of:
- ST shift on EKG
- elevated cardiac biomarker (troponin)
- in the absence of substantially reduced renal function.
Similarly, the conversion of hematology based data into useful clinical information requires the establishment of problem-solving constructs based on the measured data.
The most commonly ordered test used for managing patients worldwide is the hemogram that often incorporates
- morphologic review of a peripheral smear
- descriptive statistics
While the hemogram has undergone progressive modification of the measured features over time the subsequent expansion of the panel of tests has provided a window into the cellular changes in the
- production
- release
- or suppression
of the formed elements from the blood-forming organ into the circulation. In the hemogram one can view data reflecting the characteristics of a broad spectrum of medical conditions.
Progressive modification of the measured features of the hemogram has delineated characteristics expressed as measurements of
- size
- density, and
- concentration
resulting in many characteristic features of classification. In the diagnosis of hematological disorders
- proliferation of marrow precursors
- domination of a cell line
- suppression of hematopoiesis
Other dimensions are created by considering
- the maturity and size of the circulating cells.
The application of rules-based, automated problem solving should provide a valid approach to
- the classification and interpretation of the data used to determine a knowledge-based clinical opinion.
The exponential growth of knowledge since the mapping of the human genome enabled by parallel advances in applied mathematics that have not been a part of traditional clinical problem solving.
As the complexity of statistical models has increased
- the dependencies have become less clear to the individual.
Contemporary statistical modeling has a primary goal of finding an underlying structure in studied data sets.
The development of an evidence-based inference engine that can substantially interpret the data at hand and
- convert it in real time to a “knowledge-based opinion”
could improve clinical decision-making by incorporating into the model
- multiple complex clinical features as well as onset and duration .
An example of a difficult area for clinical problem solving is found in the diagnosis of Systemic Inflammatory Response Syndrome (SIRS) and associated sepsis. SIRS is a costly diagnosis in hospitalized patients. Failure to diagnose it in a timely manner increases the financial and safety hazard. The early diagnosis of SIRS/sepsis is made by the application of defined criteria by the clinician.
- temperature
- heartrate
- respiratory rate and
- WBC count
The application of those clinical criteria, however, defines the condition after it has developed, leaving unanswered the hope for
- a reliable method for earlier diagnosis of SIRS.
The early diagnosis of SIRS may possibly be enhanced by the measurement of proteomic biomarkers, including
- transthyretin
- C-reactive protein
- procalcitonin
- mean arterial pressure
Immature granulocyte (IG) measurement has been proposed as a
- readily available indicator of the presence of granulocyte precursors (left shift).
The use of such markers, obtained by automated systems in conjunction with innovative statistical modeling, provides
- a promising support to early accurate decision making.
Such a system aims to reduce medical error by utilizing
- the conjoined syndromic features of disparate data elements .
How we frame our expectations is important. It determines
- the data we collect to examine the process.
In the absence of data to support an assumed benefit, there is no proof of validity at whatever cost.
Potential arenas of benefit include:
- hospital operations
- nonhospital laboratory studies
- companies in the diagnostic business
- planners of health systems
The problem stated by LL WEED in “Idols of the Mind” (Dec 13, 2006):
“ a root cause of a major defect in the health care system is that, while we falsely admire and extol the intellectual powers of highly educated physicians, we do not search for the external aids their minds require.” Hospital information technology (HIT) use has been focused on information retrieval, leaving
- the unaided mind burdened with information processing.
We deal with problems in the interpretation of data presented to the physician, and how the situation could be improved through better
- design of the software that presents data .
The computer architecture that the physician uses to view the results is more often than not presented
- as the designer would prefer, and not as the end-user would like.
In order to optimize the interface for physician, the system could have a “front-to-back” design, with the call up for any patient
- A dashboard design that presents the crucial information that the physician would likely act on in an easily accessible manner
- Each item used has to be closely related to a corresponding criterion needed for a decision.
Feature Extraction.
Eugene Rypka contributed greatly to clarifying the extraction of features in a series of articles, which
- set the groundwork for the methods used today in clinical microbiology.
The method he describes is termed S-clustering, and
- will have a significant bearing on how we can view laboratory data.
He describes S-clustering as extracting features from endogenous data that
- amplify or maximize structural information to create distinctive classes.
The method classifies by taking the number of features with sufficient variety to generate maps.
The mapping is done by
- a truth table NxN of messages and choices
- each variable is scaled to assign values for each message choice.
For example, the message for an antibody titer would be converted from 0 + ++ +++ to 0 1 2 3.
Even though there may be a large number of measured values, the variety is reduced by this compression, even though it may represent less information.
The main issue is
- how a combination of variables falls into a table to convey meaningful information.
We are concerned with
- accurate assignment into uniquely variable groups by information in test relationships.
One determines the effectiveness of each variable by its contribution to information gain in the system. The reference or null set is the class having no information. Uncertainty in assigning to a classification can be countered by providing sufficient information.
One determines the effectiveness of each variable by its contribution to information gain in the system. The possibility for realizing a good model for approximating the effects of factors supported by data used
- for inference owes much to the discovery of Kullback-Liebler distance or “information”, and Akaike
- found a simple relationship between K-L information and Fisher’s maximized log-likelihood function.
In the last 60 years the application of entropy comparable to
- the entropy of physics, information, noise, and signal processing,
- developed by Shannon, Kullback, and others
- integrated with modern statistics,
- as a result of the seminal work of Akaike, Leo Goodman, Magidson and Vermunt, and work by Coifman
Akaike pioneered recognition that the choice of model influence results in a measurable manner. In particular, a larger number of variables promotes further explanations of variance, such that a model selection criterion is important that penalizes for the number of variables when success is measured by explanation of variance.
Gil David et al. introduced an AUTOMATED processing of the data available to the ordering physician and
- can anticipate an enormous impact in diagnosis and treatment of perhaps half of the top 20 most common
- causes of hospital admission that carry a high cost and morbidity.
For example:
- anemias (iron deficiency, vitamin B12 and folate deficiency, and hemolytic anemia or myelodysplastic syndrome);
- pneumonia; systemic inflammatory response syndrome (SIRS) with or without bacteremia;
- multiple organ failure and hemodynamic shock;
- electrolyte/acid base balance disorders;
- acute and chronic liver disease;
- acute and chronic renal disease;
- diabetes mellitus;
- protein-energy malnutrition;
- acute respiratory distress of the newborn;
- acute coronary syndrome;
- congestive heart failure;
- hypertension
- disordered bone mineral metabolism;
- hemostatic disorders;
- leukemia and lymphoma;
- malabsorption syndromes; and
- cancer(s)[breast, prostate, colorectal, pancreas, stomach, liver, esophagus, thyroid, and parathyroid].
- endocrine disorders
- prenatal and perinatal diseases
Rudolph RA, Bernstein LH, Babb J: Information-Induction for the diagnosis of
myocardial infarction. Clin Chem 1988;34:2031-2038.
Bernstein LH (Chairman). Prealbumin in Nutritional Care Consensus Group.
Measurement of visceral protein status in assessing protein and energy
malnutrition: standard of care. Nutrition 1995; 11:169-171.
Bernstein LH, Qamar A, McPherson C, Zarich S, Rudolph R. Diagnosis of myocardial infarction:
integration of serum markers and clinical descriptors using information theory.
Yale J Biol Med 1999; 72: 5-13.
Kaplan L.A.; Chapman J.F.; Bock J.L.; Santa Maria E.; Clejan S.; Huddleston D.J.; Reed R.G.;
Bernstein L.H.; Gillen-Goldstein J. Prediction of Respiratory Distress Syndrome using the
Abbott FLM-II amniotic fluid assay. The National Academy of Clinical Biochemistry (NACB)
Fetal Lung Maturity Assessment Project. Clin Chim Acta 2002; 326(8): 61-68.
Bernstein LH, Qamar A, McPherson C, Zarich S. Evaluating a new graphical ordinal logit method
(GOLDminer) in the diagnosis of myocardial infarction utilizing clinical features and laboratory
data. Yale J Biol Med 1999; 72:259-268.
Bernstein L, Bradley K, Zarich SA. GOLDmineR: Improving models for classifying patients with
chest pain. Yale J Biol Med 2002; 75, pp. 183-198.
Ronald Raphael Coifman and Mladen Victor Wickerhauser. Adapted Waveform Analysis as a Tool for Modeling, Feature Extraction, and Denoising. Optical Engineering, 33(7):2170–2174, July 1994.
R. Coifman and N. Saito. Constructions of local orthonormal bases for classification and regression.
C. R. Acad. Sci. Paris, 319 Série I:191-196, 1994.
Realtime Clinical Expert Support and validation System
We have developed a software system that is the equivalent of an intelligent Electronic Health Records Dashboard that provides empirical medical reference and suggests quantitative diagnostics options. The primary purpose is to gather medical information, generate metrics, analyze them in realtime and provide a differential diagnosis, meeting the highest standard of accuracy. The system builds its unique characterization and provides a list of other patients that share this unique profile, therefore
- utilizing the vast aggregated knowledge (diagnosis, analysis, treatment, etc.) of the medical community.
- The main mathematical breakthroughs are provided by accurate patient profiling and inference methodologies
- in which anomalous subprofiles are extracted and compared to potentially relevant cases.
As the model grows and its knowledge database is extended, the diagnostic and the prognostic become more accurate and precise.
We anticipate that the effect of implementing this diagnostic amplifier would result in
- higher physician productivity at a time of great human resource limitations,
- safer prescribing practices,
- rapid identification of unusual patients,
- better assignment of patients to observation, inpatient beds,
intensive care, or referral to clinic,
- shortened length of patients ICU and bed days.
The main benefit is a
- real time assessment as well as
- diagnostic options based on comparable cases,
- flags for risk and potential problems
as illustrated in the following case acquired on 04/21/10. The patient was diagnosed by our system with severe SIRS at a grade of 0.61 .

The patient was treated for SIRS and the blood tests were repeated during the following week. The full combined record of our system’s assessment of the patient, as derived from the further hematology tests, is illustrated below. The yellow line shows the diagnosis that corresponds to the first blood test (as also shown in the image above). The red line shows the next diagnosis that was performed a week later.

The MISSIVE(c) system, by Justin Pearlman, is an alternative approach that includes not only automated data retrieval and reformatting of data for decision support, but also an integrated set of tools to speed up analysis, structured for quality and error reduction, couplled to facilitated report generation, incorporation of just-in-time knowledge and group expertise, standards of care, evidence-based planning, and both physician and patient instruction.
See also in Pharmaceutical Intelligence:
The Cost Burden of Disease: U.S. and Michigan.CHRT Brief. January 2010. @www.chrt.org
The National Hospital Bill: The Most Expensive Conditions by Payer, 2006. HCUP Brief #59.
Rudolph RA, Bernstein LH, Babb J: Information-Induction for the diagnosis of myocardial infarction. Clin Chem 1988;34:2031-2038.
Bernstein LH, Qamar A, McPherson C, Zarich S, Rudolph R. Diagnosis of myocardial infarction:
integration of serum markers and clinical descriptors using information theory.
Yale J Biol Med 1999; 72: 5-13.
Kaplan L.A.; Chapman J.F.; Bock J.L.; Santa Maria E.; Clejan S.; Huddleston D.J.; Reed R.G.;
Bernstein L.H.; Gillen-Goldstein J. Prediction of Respiratory Distress Syndrome using the Abbott FLM-II amniotic fluid assay. The National Academy of Clinical Biochemistry (NACB) Fetal Lung Maturity Assessment Project. Clin Chim Acta 2002; 326(8): 61-68.
Bernstein LH, Qamar A, McPherson C, Zarich S. Evaluating a new graphical ordinal logit method (GOLDminer) in the diagnosis of myocardial infarction utilizing clinical features and laboratory
data. Yale J Biol Med 1999; 72:259-268.
Bernstein L, Bradley K, Zarich SA. GOLDmineR: Improving models for classifying patients with chest pain. Yale J Biol Med 2002; 75, pp. 183-198.
Ronald Raphael Coifman and Mladen Victor Wickerhauser. Adapted Waveform Analysis as a Tool for Modeling, Feature Extraction, and Denoising.
Optical Engineering 1994; 33(7):2170–2174.
R. Coifman and N. Saito. Constructions of local orthonormal bases for classification and regression. C. R. Acad. Sci. Paris, 319 Série I:191-196, 1994.
W Ruts, S De Deyne, E Ameel, W Vanpaemel,T Verbeemen, And G Storms. Dutch norm data for 13 semantic categoriesand 338 exemplars. Behavior Research Methods, Instruments,
& Computers 2004; 36 (3): 506–515.
De Deyne, S Verheyen, E Ameel, W Vanpaemel, MJ Dry, WVoorspoels, and G Storms. Exemplar by feature applicability matrices and other Dutch normative data for semantic
concepts. Behavior Research Methods 2008; 40 (4): 1030-1048
Landauer, T. K., Ross, B. H., & Didner, R. S. (1979). Processing visually presented single words: A reaction time analysis [Technical memorandum]. Murray Hill, NJ: Bell Laboratories.
Lewandowsky , S. (1991).
Weed L. Automation of the problem oriented medical record. NCHSR Research Digest Series DHEW. 1977;(HRA)77-3177.
Naegele TA. Letter to the Editor. Amer J Crit Care 1993;2(5):433.
Sheila Nirenberg/Cornell and Chethan Pandarinath/Stanford, “Retinal prosthetic strategy with the capacity to restore normal vision,” Proceedings of the National Academy of Sciences.
Other related articles published in this Open Access Online Scientific Journal include the following:
http://pharmaceuticalintelligence.com/2012/08/13/the-automated-second-opinion-generator/
http://pharmaceuticalintelligence.com/2012/09/21/the-electronic-health-record-how-far-we-
have-travelled-and-where-is-journeys-end/
http://pharmaceuticalintelligence.com/2013/02/18/the-potential-contribution-of-
informatics-to-healthcare-is-more-than-currently-estimated/
http://pharmaceuticalintelligence.com/2013/05/04/cardiovascular-diseases-decision-support-
systems-for-disease-management-decision-making/?goback=%2Egde_4346921_member_239739196
http://pharmaceuticalintelligence.com/2012/08/13/demonstration-of-a-diagnostic-clinical-
laboratory-neural-network-agent-applied-to-three-laboratory-data-conditioning-problems/
http://pharmaceuticalintelligence.com/2012/12/17/big-data-in-genomic-medicine/
http://pharmaceuticalintelligence.com/2013/02/13/cracking-the-code-of-human-life-
the-birth-of-bioinformatics-and-computational-genomics/
http://pharmaceuticalintelligence.com/2013/04/28/genetics-of-conduction-disease-
atrioventricular-av-conduction-disease-block-gene-mutations-transcription-excitability-
and-energy-homeostasis/
http://pharmaceuticalintelligence.com/2012/12/10/identification-of-biomarkers-that-
are-related–to-the-actin-cytoskeleton/
http://pharmaceuticalintelligence.com/2012/08/14/regression-a-richly-textured-method-
for-comparison-and-classification-of-predictor-variables/
http://pharmaceuticalintelligence.com/2012/08/02/diagnostic-evaluation-of-sirs-by-
immature-granulocytes/
http://pharmaceuticalintelligence.com/2012/08/01/automated-inferential-diagnosis-
of-sirs-sepsis-septic-shock/
http://pharmaceuticalintelligence.com/2012/08/12/1815/
http://pharmaceuticalintelligence.com/2012/08/15/1946/
http://pharmaceuticalintelligence.com/2013/05/13/vinod-khosla-20-doctor-included-speculations-
musings-of-a-technology-optimist-or-technology-will-replace-80-of-what-doctors-do/
http://pharmaceuticalintelligence.com/2013/05/05/bioengineering-of-vascular-and-tissue-models/
The Heart: Vasculature Protection – A Concept-based Pharmacological Therapy including THYMOSIN
Aviva Lev-Ari, PhD, RN 2/28/2013
http://pharmaceuticalintelligence.com/2013/02/28/the-heart-vasculature-protection-a-concept-
based-pharmacological-therapy-including-thymosin/
FDA Pending 510(k) for The Latest Cardiovascular Imaging Technology
Aviva Lev-Ari, PhD, RN 1/28/2013
http://pharmaceuticalintelligence.com/2013/01/28/fda-pending-510k-for-the-latest-
cardiovascular-imaging-technology/
PCI Outcomes, Increased Ischemic Risk associated with Elevated Plasma Fibrinogen not
Platelet Reactivity Aviva Lev-Ari, PhD, RN 1/10/2013
http://pharmaceuticalintelligence.com/2013/01/10/pci-outcomes-increased-ischemic-risk-
associated-with-elevated-plasma-fibrinogen-not-platelet-reactivity/
The ACUITY-PCI score: Will it Replace Four Established Risk Scores — TIMI, GRACE, SYNTAX,
and Clinical SYNTAX Aviva Lev-Ari, PhD, RN 1/3/2013
http://pharmaceuticalintelligence.com/2013/01/03/the-acuity-pci-score-will-it-replace-four-
established-risk-scores-timi-grace-syntax-and-clinical-syntax/
Coronary artery disease in symptomatic patients referred for coronary angiography: Predicted by
Serum Protein Profiles Aviva Lev-Ari, PhD, RN 12/29/2012
http://pharmaceuticalintelligence.com/2012/12/29/coronary-artery-disease-in-symptomatic-
patients-referred-for-coronary-angiography-predicted-by-serum-protein-profiles/
New Definition of MI Unveiled, Fractional Flow Reserve (FFR)CT for Tagging Ischemia
Aviva Lev-Ari, PhD, RN 8/27/2012
http://pharmaceuticalintelligence.com/2012/08/27/new-definition-of-mi-unveiled-
fractional-flow-reserve-ffrct-for-tagging-ischemia/
Herceptin Fab (antibody) – light and heavy chains (Photo credit: Wikipedia)

Personalized Medicine (Photo credit: Wikipedia)
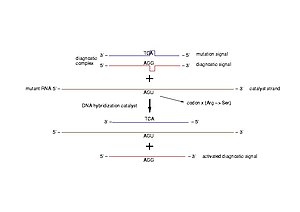
Diagnostic of pathogenic mutations. A diagnostic complex is a dsDNA molecule resembling a short part of the gene of interest, in which one of the strands is intact (diagnostic signal) and the other bears the mutation to be detected (mutation signal). In case of a pathogenic mutation, the transcribed mRNA pairs to the mutation signal and triggers the release of the diagnostic signal (Photo credit: Wikipedia)
Like this:
Like Loading...
Read Full Post »







 DNA decoder: Knome’s software can tease out medically relevant changes in DNA that could disrupt individual gene function or even a whole molecular pathway, as is highlighted here—certain mutations in the BRCA2 gene, which affects the function of many other genes, can be associated with an increased risk of breast cancer.
DNA decoder: Knome’s software can tease out medically relevant changes in DNA that could disrupt individual gene function or even a whole molecular pathway, as is highlighted here—certain mutations in the BRCA2 gene, which affects the function of many other genes, can be associated with an increased risk of breast cancer.