Ubiquitin Pathway Involved in Neurodegenerative Diseases
Larry H Bernstein, MD, FCAP
PINK1 and Parkin and Parkinson’s Disease
Studies of the familial Parkinson disease-related proteins PINK1 and Parkin have demonstrated that these factors promote the fragmentation and turnover of mitochondria following treatment of cultured cells with mitochondrial depolarizing agents. Whether PINK1 or Parkin influence mitochondrial quality control under normal physiological conditions in dopaminergic neurons, a principal cell type that degenerates in Parkinson disease, remains unclear. To address this matter, we developed a method to purify and characterize neural subtypes of interest from the adult Drosophila brain.
Using this method, we find that dopaminergic neurons from Drosophila parkin mutants accumulate enlarged, depolarized mitochondria, and that genetic perturbations that promote mitochondrial fragmentation and turnover rescue the mitochondrial depolarization and neurodegenerative phenotypes of parkin mutants. In contrast, cholinergic neurons from parkin mutants accumulate enlarged depolarized mitochondria to a lesser extent than dopaminergic neurons, suggesting that a higher rate of mitochondrial damage, or a deficiency in alternative mechanisms to repair or eliminate damaged mitochondria explains the selective vulnerability of dopaminergic neurons in Parkinson disease.
Our study validates key tenets of the model that PINK1 and Parkin promote the fragmentation and turnover of depolarized mitochondria in dopaminergic neurons. Moreover, our neural purification method provides a foundation to further explore the pathogenesis of Parkinson disease, and to address other neurobiological questions requiring the analysis of defined neural cell types.
Burmana JL, Yua S, Poole AC, Decala RB , Pallanck L. Analysis of neural subtypes reveals selective mitochondrial dysfunction in dopaminergic neurons from parkin mutants.
Autophagy in Parkinson’s Disease.
Parkinson’s disease is a common neurodegenerative disease in the elderly. To explore the specific role of autophagy and the ubiquitin-proteasome pathway in apoptosis, a specific proteasome inhibitor and macroautophagy inhibitor and stimulator were selected to investigate pheochromocytoma (PC12) cell lines transfected with human mutant (A30P) and wildtype (WT) -synuclein.
The apoptosis ratio was assessed by flow cytometry. LC3, heat shock protein 70 (hsp70) and caspase-3 expression in cell culture were determined by Western blot. The hallmarks of apoptosis and autophagy were assessed with transmission electron microscopy. Compared to the control group or the rapamycin (autophagy stimulator) group, the apoptosis ratio in A30P and WT cells was significantly higher after treatment with inhibitors of the proteasome and macroautophagy. The results of Western blots for caspase-3 expression were similar to those of flow cytometry; hsp70 protein was significantly higher in the proteasome inhibitor group than in control, but in the autophagy inhibitor and stimulator groups, hsp70 was similar to control. These findings show that inhibition of the proteasome and autophagy promotes apoptosis, and the macroautophagy stimulator rapamycin reduces the apoptosis ratio. And inhibiting or stimulating autophagy has less impact on hsp70 than the proteasome pathway.
In conclusion, either stimulation or inhibition of macroautophagy, has less impact on hsp70 than on the proteasome pathway. This study found that rapamycin decreased apoptotic cells in A30P cells independent of caspase-3 activity. Although several lines of evidence recently demonstrated crosstalk between autophagy and caspase-independent apoptosis, we could not confirm that autophagy activation protects cells from caspase-independent cell death. Undoubtedly, there are multiple connections between the apoptotic and autophagic processes.
Inhibition of autophagy may subvert the capacity of cells to remove damaged organelles or to remove misfolded proteins, which would favor apoptosis. However, proteasome inhibition activated macroautophagy and accelerated apoptosis. A likely explanation is inhibition of the proteasome favors oxidative reactions that trigger apoptosis, presumably through
- a direct effect on mitochondria, and
- the absence of NADPH2 and ATP
which may deinhibit the activation of caspase-2 or MOMP. Another possibility is that aggregated proteins induced by proteasome inhibition increase apoptosis.
Yang F, Yanga YP, Maoa CJ, Caoa BY, et al. Role of autophagy and proteasome degradation pathways in apoptosis of PC12 cells overexpressing human -synuclein. Neuroscience Letters 2009; 454:203–208. doi:10.1016/j.neulet.2009.03.027. www.elsevier.com/locate/neulet http://neurosciletters.com/ Role_of_autophagy_and_proteasome_degradation_pathways_in_apoptosis_of_PC12_cells_
overexpressing_human –synuclein/
Parkin-dependent Ubiquitination of Endogenous Bax
Autosomal recessive loss-of-function mutations within the PARK2 gene functionally inactivate the E3 ubiquitin ligase parkin, resulting in neurodegeneration of catecholaminergic neurons and a familial form of Parkinson disease. Current evidence suggests both a mitochondrial function for parkin and a neuroprotective role, which may in fact be interrelated. The antiapoptotic effects of Parkin have been widely reported, and may involve fundamental changes in the threshold for apoptotic cytochrome c release, but the substrate(s) involved in Parkin dependent protection had not been identified. Here, we demonstrate the Parkin-dependent ubiquitination of endogenous Bax comparing primary cultured neurons from WT and Parkin KO mice and using multiple Parkin-overexpressing cell culture systems. The direct ubiquitination of purified Bax was also observed in vitro following incubation with recombinant parkin. The authors found that Parkin prevented basal and apoptotic stress induced translocation of Bax to the mitochondria. Moreover, an engineered ubiquitination-resistant form of Bax retained its apoptotic function, but Bax KO cells complemented with lysine-mutant Bax did not manifest the antiapoptotic effects of Parkin that were observed in cells expressing WT Bax. These data suggest that Bax is the primary substrate responsible for the antiapoptotic effects of Parkin, and provide mechanistic insight into at least a subset of the mitochondrial effects of Parkin.
Johnson BN, Berger AK, Cortese GP, and LaVoie MJ. The ubiquitin E3 ligase Parkin regulates the proapoptotic function of Bax. PNAS 2012, pp 6. www.pnas.org/cgi/doi/10.1073/pnas.1113248109
http://PNAS.org/ The_ubiquitin_E3_ligase_Parkin_regulates_the_proapoptotic_function_of_Bax
Parkin Promotes Mitochondrial Loss in Autophagy
Parkin, an E3 ubiquitin ligase implicated in Parkinson’s disease, promotes degradation of dysfunctional mitochondria by autophagy. Using proteomic and cellular approaches, we show that upon translocation to mitochondria, Parkin activates the ubiquitin–proteasome system (UPS) for widespread degradation of outer membrane proteins. This is evidenced by an increase in K48-linked polyubiquitin on mitochondria, recruitment of the 26S proteasome and rapid degradation of multiple outer membrane proteins. The degradation of proteins by the UPS occurs independently of the autophagy pathway, and inhibition of the 26S proteasome completely abrogates Parkin-mediated mitophagy in HeLa, SH-SY5Y and mouse cells. Although the mitofusins Mfn1 and Mfn2 are rapid degradation targets of Parkin, degradation of additional targets is essential for mitophagy. These results indicate that remodeling of the mitochondrial outer membrane proteome is important for mitophagy, and reveal a causal link between the UPS and autophagy, the major pathways for degradation of intracellular substrates.
Chan NC, Salazar AM, Pham AH, Sweredoski MJ, et al. Broad activation of the ubiquitin–proteasome system by Parkin is critical for mitophagy. Human Molecular Genetics 2011; 20(9): 1726–1737. doi:10.1093/hmg/ddr048. http://HumMolecGenetics.com/ Broad_activation_of_the_ubiquitin–proteasome_system_by_Parkin_is_critical_for_mitophagy/
Interactome Networks and Protein Expression
Aloy P. Shaping the future of interactome networks. (A report of the third Interactome Networks Conference, Hinxton, UK, 29 August-1 September 2007). Genome Biology 2007; 8:316 (doi:10.1186/gb-2007-8-10-316)
Complex systems are often networked, and biology is no exception. Following on from the genome sequencing projects, experiments show that proteins in living organisms are highly connected, which helps to explain how such great complexity can be achieved by a comparatively small set of gene products. At a recent conference on interactome networks held outside Cambridge, UK, the most recent advances in research on cellular networks were discussed. This year’s conference focused on identifying the strengths and weaknesses of currently resolved interaction networks and the techniques used to determine them – reflecting the fact that the field of mapping interaction networks is maturing.
Peroutka RJ, Orcutt SJ, Strickler JE, and Butt TR. SUMO Fusion Technology for Enhanced Protein Expression and Purification in Prokaryotes and Eukaryotes. Chapter 2. in T.C. Evans, M.-Q. Xu (eds.), Heterologous Gene Expression in E. coli, Methods in Molecular Biology 705:15-29. DOI 10.1007/978-1-61737-967-3_2, © Springer Science+Business Media, LLC 2011
The preparation of sufficient amounts of high-quality protein samples is the major bottleneck for structural proteomics. The use of recombinant proteins has increased significantly during the past decades. The most commonly used host, Escherichia coli, presents many challenges including protein misfolding, protein degradation, and low solubility. A novel SUMO fusion technology appears to enhance protein expression and solubility (www.lifesensors.com). Efficient removal of the SUMO tag by SUMO protease in vitro facilitates the generation of target protein with a native N-terminus. In addition to its physiological relevance in eukaryotes, SUMO can be used as a powerful biotechnology tool forenhanced functional protein expression in prokaryotes and eukaryotes.
IL-6 regulation on mitochondrial remodeling/dysfunction
Muscle protein turnover regulation during cancer cachexia is being rapidly defined, and skeletal muscle mitochondria function appears coupled to processes regulating muscle wasting. Skeletal muscle oxidative capacity and the expression of proteins regulating mitochondrial biogenesis and dynamics are disrupted in severely cachectic ApcMin/+ mice. It has not been determined if these changes occur at the onset of cachexia and are necessary for the progression of muscle wasting. Exercise and anti-cytokine therapies have proven effective in preventing cachexia development in tumor bearing mice, while their effect on mitochondrial content, biogenesis and dynamics is not well understood.
The purposes of this study were to
1) determine IL-6 regulation on mitochondrial remodeling/dysfunction during the progression of cancer cachexia and
2) to determine if exercise training can attenuate mitochondrial dysfunction and the induction of proteolytic pathways during IL-6 induced cancer cachexia.
ApcMin/+ mice were examined during the progression of cachexia, after systemic interleukin (IL)-6r antibody treatment, or after IL-6 over-expression with or without exercise. Direct effects of IL-6 on mitochondrial remodeling were examined in cultured C2C12 myoblasts.
- Mitochondrial content was not reduced during the initial development of cachexia, while muscle PGC-1α and fusion (Mfn1, Mfn2) protein expression was repressed.
- With progressive weight loss mitochondrial content decreased, PGC-1α and fusion proteins were further suppressed, and fission protein (FIS1) was induced.
IL-6 receptor antibody administration after the onset of cachexia improved mitochondrial content,
- PGC-1α,
- Mfn1/Mfn2 and
- FIS1 protein expression.
IL-6 over-expression in pre-cachectic mice accelerated body weight loss and muscle wasting, without reducing mitochondrial content, while PGC-1α and Mfn1/Mfn2 protein expression was suppressed and FIS1 protein expression induced. Exercise normalized these IL-6 induced effects. C2C12 myotubes administered IL-6 had
- increased FIS1 protein expression,
- increased oxidative stress, and
- reduced PGC-1α gene expression
- without altered mitochondrial protein expression.
Altered expression of proteins regulating mitochondrial biogenesis and fusion are early events in the initiation of cachexia regulated by IL-6, which precede the loss of muscle mitochondrial content. Furthermore, IL-6 induced mitochondrial remodeling and proteolysis can be rescued with moderate exercise training even in the presence of high circulating IL-6 levels.
White JP, Puppa MJ, Sato S, Gao S. IL-6 regulation on skeletal muscle mitochondrial remodeling during cancer cachexia in the ApcMin/+ mouse. Skeletal Muscle 2012; 2:14-30.
http://www.skeletalmusclejournal.com/content/2/1/14
Starvation-induced Autophagy
Upon starvation cells undergo autophagy, a cellular degradation pathway important in the turnover of whole organelles and long lived proteins. Starvation-induced protein degradation has been regarded as an unspecific bulk degradation process. We studied global protein dynamics during amino acid starvation-induced autophagy by quantitative mass spectrometry and were able to record nearly 1500 protein profiles during 36 h of starvation. Cluster analysis of the recorded protein profiles revealed that cytosolic proteins were degraded rapidly, whereas proteins annotated to various complexes and organelles were degraded later at different time periods. Inhibition of protein degradation pathways identified the lysosomal/autophagosomal system as the main degradative route.
Thus, starvation induces degradation via autophagy, which appears to be selective and to degrade proteins in an ordered fashion and not completely arbitrarily as anticipated so far.
Kristensen AR, Schandorff S, Høyer-Hansen M, Nielsen MO, et al. Ordered Organelle Degradation during Starvation-induced Autophagy. Molecular & Cellular Proteomics 2008; 7:2419–2428.
http://MolecCellProteomics.com/Ordered_Organelle_Degradation_during_Starvation-induced_Autophagy/
Skeletal Muscle Macroautophagy
Skeletal muscles are the agent of motion and one of the most important tissues responsible for the control of metabolism. Coordinated movements are allowed by the highly organized structure of the cytosol of muscle fibers (or myofibers), the multinucleated and highly specialized cells of skeletal muscles involved in contraction. Contractile proteins are assembled into repetitive structures, the basal unit of which is the sarcomere, that are well packed into the myofiber cytosol. Myonuclei are located at the edge of the myofibers, whereas the various organelles such as mitochondria and sarcoplasmic reticulum are embedded among the myofibrils. Many different changes take place in the cytosol of myofibers during catabolic conditions:
- proteins are mobilized
- organelles networks are reorganized for energy needs
- the setting of myonuclei can be modified.
Further,
- strenuous physical activity,
- improper dietary regimens and
- aging
lead to mechanical and metabolic damages of
- myofiber organelles,
- especially mitochondria, and
- contractile proteins.
During aging the protein turnover is slowed down, therefore it is easier to accumulate aggregates of dysfunctional proteins. Therefore, a highly dynamic tissue such as skeletal muscle requires a rapid and efficient system for the removal of altered organelles, the elimination of protein aggregates, and the disposal of toxic products.
The two major proteolytic systems in muscle are the ubiquitin-proteasome and the autophagy-lysosome pathways. The proteasome system requires
- the transcription of the two ubiquitin ligases (atrogin-1 and MuRF1) and
- the ubiquitination of the substrates.
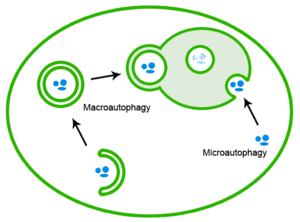
Therefore, the ubiquitin-proteasome system can provide the rapid elimination of single proteins or small aggregates. Conversely, the autophagic system is able to degrade entire organelles and large proteins aggregates. In the autophagy-lysosome system, double-membrane vesicles named autophagosomes are able to engulf a portion of the cytosol and fuse with lysosomes, where their content is completely degraded by lytic enzymes.
The autophagy flux can be biochemicaly monitored following LC3 lipidation and p62 degradation. LC3 is the mammalian homolog of the yeast Atg8 gene, which is lipidated when recruited for the double-membrane commitment and growth. p62 (SQSTM-1) is a polyubiquitin-binding protein involved in the proteasome system and that can either reside free in the cytosol and nucleus or occur within autophagosomes and lysosomes. The GFP-LC3 transgenic mouse model allows easy detection of autophagosomes by simply monitoring the presence of bright GFP-positive puncta inside the myofibrils and beneath the plasma membrane of the myofibers, thus investigate the activation of autophagy in skeletal muscles with different contents of slow and fast-twitching myofibers and in response to stimuli such as fasting. For example, in the fast-twiching extensor digitorum longus muscle few GFP-LC3 dots were observed before starvation, while many small GFP-LC3 puncta appeared between myofibrils and in the perinuclear regions after 24 h starvation. Conversely, in the slow-twitching soleus muscle, autophagic puncta were almost absent in standard condition and scarcely induced after 24 h starvation.
Autophagy in Muscle Homeostasis
The autophagic flux was found to be increased during certain catabolic conditions, such as fasting, atrophy , and denervation , thus contributing to protein breakdown. Food deprivation is one of the strongest stimuli known to induce autophagy in muscle. Indeed skeletal muscle, after the liver, is the most responsive tissue to autophagy activation during food deprivation. Since muscles are the biggest reserve of amino acids in the body, during fasting autophagy has the vital role to maintain the amino acid pool by digesting muscular protein and organelles. In mammalian cells, mTORC1, which consists of
- mTOR and
- Raptor,
is the nutrient sensor that negatively regulates autophagy.
During atrophy, protein breakdown is mediated by atrogenes, which are under the forkhead box O (FoxO) transcription factors control, and activation of autophagy seems to aggravate muscle loss during atrophy. In vivo and in vitro studies demonstrated that several genes coding for components of the autophagic machinery, such as
- LC3,
- GABARAP,
- Vps34,
- Atg12 and
- Bnip3,
are controlled by FoxO3 transcription factor. FoxO3 is able to regulate independently
- the ubiquitin-proteasome system and
- the autophagy-lysosome machinery in vivo and in vitro.
Denervation is also able to induce autophagy in skeletal muscle, although at a slower rate than fasting. This effect is mediated by RUNX1, a transcription factor upregulated during autophagy; the lack of RUNX1 results in
- excessive autophagic flux in denervated muscle and leads to atrophy.
The generation of Atg5 and Atg7 muscle-specific knockout mice have shown that
- with suppression of autophagy both models display muscle weakness and atrophy and
- a significant reduction of weight, which is
- correlated with the important loss of muscle tissue due to an atrophic condition.
An unbalanced autophagy flux is highly detrimental for muscle, as too much induces atrophy whereas too little leads to muscle weakness and degeneration. Muscle wasting associated with autophagy inhibition becomes evident and symptomatic only after a number of altered proteins and dysfunctional organelles are accumulated, a condition that becomes evident after months or even years. On the other hand, the excessive increase of autophagy flux is able to induce a rapid loss of muscle mass (within days or weeks). Alterations of autophagy are involved in the pathogenesis of several myopathies and dystrophies.
The maintenance of muscle homeostasis is finely regulated by the balance between catabolic and anabolic process. Macroautophagy (or autophagy) is a catabolic process that provides the degradation of protein aggregation and damaged organelles through the fusion between autophagosomes and lysosomes. Proper regulation of the autophagy flux is fundamental for
- the homeostasis of skeletal muscles during physiological situations and
- in response to stress.
Defective as well as excessive autophagy is harmful for muscle health and has a pathogenic role in several forms of muscle diseases.
Grumati P, Bonaldo P. Autophagy in Skeletal Muscle Homeostasis and in Muscular Dystrophies. Cells 2012, 1, 325-345; doi:10.3390/cells1030325. ISSN 2073-4409. www.mdpi.com/journal/cells
http://cell.com/ Autophagy in Skeletal Muscle Homeostasis and in Muscular Dystrophies/
Parkinson’s Disease Mutations
Mutations in parkin, a ubiquitin ligase, cause early-onset familial Parkinson’s disease (AR-JP). How Parkin suppresses Parkinsonism remains unknown. Parkin was recently shown to promote the clearance of impaired mitochondria by autophagy, termed mitophagy. Here, we show that Parkin promotes mitophagy by catalyzing mitochondrial ubiquitination, which in turn recruits ubiquitin-binding autophagic components, HDAC6 and p62, leading to mitochondrial clearance.
During the process, juxtanuclear mitochondrial aggregates resembling a protein aggregate-induced aggresome are formed. The formation of these “mito-aggresome” structures requires microtubule motor-dependent transport and is essential for efficient mitophagy. Importantly, we show that AR-JP–causing Parkin mutations are defective in supporting mitophagy due to distinct defects at
- recognition,
- transportation, or
- ubiquitination of impaired mitochondria,
thereby implicating mitophagy defects in the development of Parkinsonism. Our results show that impaired mitochondria and protein aggregates are processed by common ubiquitin-selective autophagy machinery connected to the aggresomal pathway, thus identifying a mechanistic basis for the prevalence of these toxic entities in Parkinson’s disease.
Lee JY,Nagano Y, Taylor JP,Lim KL, and Yao TP. Disease-causing mutations in Parkin impair mitochondrial ubiquitination, aggregation, and HDAC6-dependent mitophagy. J Cell Biol 2010; 189(4):671-679. www.jcb.org/cgi/doi/10.1083/jcb.201001039
http://JCellBiol.com/Disease-causing_mutations_in_Parkin_impair_mitochondrial_ubiquitination_ aggregation_and_HDAC6-dependent_mitophagy/
Drosophila Parkin Requires PINK1
Loss of the E3 ubiquitin ligase Parkin causes early onset Parkinson’s disease, a neurodegenerative disorder of unknown etiology. Parkin has been linked to multiple cellular processes including
- protein degradation,
- mitochondrial homeostasis, and
- autophagy;
however, its precise role in pathogenesis is unclear. Recent evidence suggests that Parkin is recruited to damaged mitochondria, possibly affecting
- mitochondrial fission and/or fusion,
- to mediate their autophagic turnover.
The precise mechanism of recruitment and the ubiquitination target are unclear. Here we show in Drosophila cells that PINK1 is required to recruit Parkin to dysfunctional mitochondria and promote their degradation. Furthermore, PINK1 and Parkin mediate the ubiquitination of the profusion factor Mfn on the outer surface of mitochondria. Loss of Drosophila PINK1 or parkin causes an increase in Mfn abundance in vivo and concomitant elongation of mitochondria. These findings provide a molecular mechanism by which the PINK1/Parkin pathway affects mitochondrial fission/fusion as suggested by previous genetic interaction studies. We hypothesize that Mfn ubiquitination may provide a mechanism by which terminally damaged mitochondria are labeled and sequestered for degradation by autophagy.
Ziviani E, Tao RN, and Whitworth AJ. Drosophila Parkin requires PINK1 for mitochondrial translocation and ubiquitinates Mitofusin. PNAS 2010. Pp6 www.pnas.org/cgi/doi/10.1073/pnas.0913485107
http://PNAS.org/ Drosophila_Parkin_requires_PINK1_for_ mitochondrial_translocation_and_ubiquitinates_Mitofusin
Dynamin-related protein 1 (Drp1) in Parkinson’s
Mutations in Parkin, an E3 ubiquitin ligase that regulates protein turnover, represent one of the major causes of familial Parkinson’s disease (PD), a neurodegenerative disorder characterized by the loss of dopaminergic neurons and impaired mitochondrial functions. The underlying mechanism by which pathogenic parkin mutations induce mitochondrial abnormality is not fully understood. Here we demonstrate that Parkin interacts with and subsequently ubiquitinates dynamin-related protein 1 (Drp1), for promoting its proteasome-dependent degradation. Pathogenic mutation or knockdown of Parkin inhibits the ubiquitination and degradation of Drp1, leading to an increased level of Drp1 for mitochondrial fragmentation. These results identify Drp1 as a novel substrate of Parkin and suggest a potential mechanism linking abnormal Parkin expression to mitochondrial dysfunction in the pathogenesis of PD.
Wang H, Song P, Du L, Tian W. Parkin ubiquitinates Drp1 for proteasome-dependent degradation: implication of dysregulated mitochondrial dynamics in Parkinson’s disease.
JBC Papers in Press. Published on February 3, 2011 as Manuscript M110.144238. http://www.jbc.org/cgi/doi/10.1074/jbc.M110.144238
http://JBC.org/ Parkin_ubiquitinates_Drp1_for_proteasome-dependent_degradation_implication_of_ dysregulated_mitochondrial_dynamics_in_Parkinson’s_disease
Pink1, Parkin, and DJ-1 Form a Complex
Mutations in the genes PTEN-induced putative kinase 1 (PINK1), PARKIN, and DJ-1 cause autosomal recessive forms of Parkinson disease (PD), and the Pink1/Parkin pathway regulates mitochondrial integrity and function. An important question is whether the proteins encoded by these genes function to regulate activities of other cellular compartments. A study in mice, reported by Xiong et al. in this issue of the JCI, demonstrates that Pink1, Parkin, and DJ-1 can form a complex in the cytoplasm, with Pink1 and DJ-1 promoting the E3 ubiquitin ligase activity of Parkin to degrade substrates via the proteasome (see the related article, doi:10.1172/ JCI37617).
This protein complex in the cytosol may or may not be related to the role of these proteins in regulating mitochondrial function or oxidative stress in vivo. Three models for the role of the PPD complex. In this issue of the JCI, Xiong et al. report that Pink1, Parkin, and DJ-1 bind to each other and form a PPD E3 ligase complex in which Pink1 and DJ-1 modulate Parkin-dependent ubiquitination and subsequent degradation of substrates via the proteasome. Previous work suggests that the Pink1/Parkin pathway regulates mitochondrial integrity and promotes mitochondrial fission in Drosophila.
(A) Parkin and DJ-1 may be recruited to the mitochondrial outer membrane during stress and interact with Pink1. These interactions may facilitate the ligase activity of Parkin, thereby facilitating the turnover of molecules that regulate mitochondrial dynamics and mitophagy. The PPD complex may have other roles in the cytosol that result in degradative ubiquitination and/or relay information from mitochondria to other cellular compartments.
(B) Alternatively, Pink1 may be released from mitochondria after cleavage to interact with DJ-1 and Parkin in the cytosol.
A and B differ in the site of action of the PPD complex and the cleavage status of Pink1.
The complex forms on the mitochondrial outer membrane potentially containing full-length Pink1 in A, and in the cytosol with cleaved Pink1 in B.
Lack of DJ-1 function results in phenotypes that are distinct from the mitochondrial phenotypes observed in null mutants of Pink1 or Parkin in Drosophila. Thus, although the PPD complex is illustrated here as regulating mitochondrial fission, the role of DJ-1 in vivo remains to be clarified.
(C) It is also possible that the action occurs in the cytosol and is independent of the function of Pink1/Parkin in regulating mitochondrial integrity and function.
The Xiong et al. study offers an entry point for explorations of the role of Pink1, Parkin, and DJ-1 in the cytoplasm. It remains to be shown whether Parkin, in complex with Pink1 and DJ-1, carries out protein degradation in vivo.
Li H, and Guo M. Protein degradation in Parkinson disease revisited: it’s complex. commentaries. J Clin Invest. doi:10.1172/JCI38619. http://www.jci.org
http://JCI.org/ Protein_degradation_in_Parkinson_disease_revisited_it’s_complex/
Xiong, H., et al. Parkin, PINK1, and DJ-1 form a ubiquitin E3 ligase complex promoting unfolded protein degradation. J. Clin. Invest. 2009; 119:650–660.
http://JCI.org/ Parkin_PINK1_DJ1_form_ubiquitin_E3_ligase_complex_promoting_unfolded_protein_degradation/
Mitochondrial Ubiquitin Ligase, MITOL, protects neuronal cells
Nitric oxide (NO) is implicated in neuronal cell survival. However, excessive NO production mediates neuronal cell death, in part via mitochondrial dysfunction. Here, we report that the mitochondrial ubiquitin ligase, MITOL, protects neuronal cells from mitochondrial damage caused by accumulation of S-nitrosylated microtubule associated protein 1B-light chain 1 (LC1). S-nitrosylation of LC1 induces a conformational change that serves both to activate LC1 and to promote its ubiquination by MITOL, indicating that microtubule stabilization by LC1 is regulated through its interaction with MITOL. Excessive NO production can inhibit MITOL, and MITOL inhibition resulted in accumulation of S-nitrosylated LC1 following stimulation of NO production by calcimycin and N-methyl-D-aspartate. LC1 accumulation under these conditions resulted in mitochondrial dysfunction and neuronal cell death. Thus, the balance between LC1 activation by S-nitrosylation and down-regulation by MITOL is critical for neuronal cell survival. Our findings may contribute significantly to an understanding of the mechanisms of neurological diseases caused by nitrosative stress-mediated mitochondrial dysfunction.
Yonashiro R, Kimijima Y, Shimura T, Kawaguchi K, et al. Mitochondrial ubiquitin ligase MITOL blocks S-nitrosylated MAP1B-light chain 1-mediated mitochondrial dysfunction and neuronal cell death. PNAS; 2012. pp 6. www.pnas.org/cgi/doi/10.1073/pnas.1114985109
Ubiquitin–Proteasome System in Neurodegeneration
A common histopathological hallmark of most neurodegenerative diseases is the presence of aberrant proteinaceous inclusions inside affected neurons. Because these protein aggregates are detected using antibodies against components of the ubiquitin–proteasome system (UPS), impairment of this machinery for regulated proteolysis has been suggested to be at the root of neurodegeneration. This hypothesis has been difficult to prove in vivo owing to the lack of appropriate tools. The recent report of transgenic mice with ubiquitous expression of a UPS-reporter protein should finally make it possible to test in vivo the role of the UPS in neurodegeneration.
Hernandez F, Dıaz-Hernandez M, Avila J and Lucas JJ. Testing the ubiquitin–proteasome hypothesis of neurodegeneration in vivo. TRENDS in Neurosciences 2004; 27(2): 66-68.
ALP in Parkinson’s
The ubiquitin-proteasome system (UPS) and autophagy-lysosome pathway (ALP) are the two most important mechanisms that normally repair or remove abnormal proteins. Alterations in the function of these systems to degrade misfolded and aggregated proteins are being increasingly recognized as playing a pivotal role in the pathogenesis of many neurodegenerative disorders such as Parkinson’s disease. Dysfunction of the UPS has been already strongly implicated in the pathogenesis of this disease and, more recently, growing interest has been shown in identifying the role of ALP in neurodegeneration. Mutations of a-synuclein and the increase of intracellular concentrations of non-mutant a-synuclein have been associated with Parkinson’s disease phenotype.
The demonstration that a-synuclein is degraded by both proteasome and autophagy indicates a possible linkage between the dysfunction of the UPS or ALP and the occurrence of this disorder.The fact that mutant a-synucleins inhibit ALP functioning by tightly binding to the receptor on the lysosomal membrane for autophagy pathway further supports the assumption that impairment of the ALP may be related to the development of Parkinson’s disease. In this review, we summarize the recent findings related to this topic and discuss the unique role of the ALP in this neurogenerative disorder and the putative therapeutic potential through ALP enhancement.
Pan Y, Kondo S, Le W, Jankovic J. The role of autophagy-lysosome pathway in neurodegeneration associated with Parkinson’s disease. Brain 2008; 131: 1969-1978. doi:10.1093/brain/awm318.
Ubiquitin-Proteasome System in Parkinson’s
There is growing evidence that dysfunction of the mitochondrial respiratory chain and failure of the cellular protein degradation machinery, specifically the ubiquitin-proteasome system, play an important role in the pathogenesis of Parkinson’s disease. We now show that the corresponding pathways of these two systems are linked at the transcriptomic level in Parkinsonian substantia nigra. We examined gene expression in medial and lateral substantia nigra (SN) as well as in frontal cortex using whole genome DNA oligonucleotide microarrays. In this study, we use a hypothesis-driven approach in analysing microarray data to describe the expression of mitochondrial and ubiquitin-proteasomal system (UPS) genes in Parkinson’s disease (PD).
Although a number of genes showed up-regulation, we found an overall decrease in expression affecting the majority of mitochondrial and UPS sequences. The down-regulated genes include genes that encode subunits of complex I and the Parkinson’s-disease-linked UCHL1. The observed changes in expression were very similar for both medial and lateral SN and also affected the PD cerebral cortex. As revealed by “gene shaving” clustering analysis, there was a very significant correlation between the transcriptomic profiles of both systems including in control brains.
Therefore, the mitochondria and the proteasome form a higher-order gene regulatory network that is severely perturbed in Parkinson’s disease. Our quantitative results also suggest that Parkinson’s disease is a disease of more than one cell class, i.e. that it goes beyond the catecholaminergic neuron and involves glia as well.
Duke DC, Moran LB, Kalaitzakis ME, Deprez M, et al. Transcriptome analysis reveals link between proteasomal and mitochondrial pathways in Parkinson’s disease. Neurogenetics 2006; 7:139-148.
The causes of various neurodegenerative diseases, particularly sporadic cases, remain unknown, but increasing evidence suggests that these diseases may share similar molecular and cellular mechanisms of pathogenesis. One prominent feature common to most neurodegenerative diseases is the accumulation of misfolded proteins in the form of insoluble protein aggregates or inclusion bodies. Although these aggregates have different protein compositions, they all contain ubiquitin and proteasome subunits, implying a failure of the ubiquitin-proteasome system (UPS) in the removal of misfolded proteins.
A direct link between UPS dysfunction and neurodegeneration has been provided by recent findings that genetic mutations in UPS components cause several rare, familial forms of neurodegenerative diseases. Furthermore, it is becoming increasingly clear that oxidative stress, which results from aging or exposure to environmental toxins, can directly damage UPS components, thereby contributing to the pathogenesis of sporadic forms of neurodegenerative diseases.
Aberrations in the UPS often result in defective proteasome-mediated protein degradation, leading to accumulation of toxic proteins and eventually to neuronal cell death. Interestingly, emerging evidence has begun to suggest that impairment in substrate-specific components of the UPS, such as E3 ubiquitin-protein ligases, may cause aberrant ubiquitination and neurodegeneration in a proteasome-independent manner. This provides an overview of the molecular components of the UPS and their impairment in familial and sporadic forms of neurodegenerative diseases, and summarizes present knowledge about the pathogenic mechanisms of UPS dysfunction in neurodegeneration.
Molecular mechanisms of protein ubiquitination and degradation by the UPS. Ubiquitination involves a highly specific enzyme cascade in which
- ubiquitin (Ub) is first activated by the ubiquitinactivating enzyme (E1),
- then transferred to an ubiquitin-conjugating enzyme (E2), and
- finally covalently attached to the substrate by an ubiquitin-protein ligase (E3).
Ubiquitination is a reversible posttranslational modification in which the removal of Ub is mediated by a deubiquitinating enzyme (DUB).
- Substrate proteins can be either monoubiquitinated or polyubiquitinated through successive conjugation of Ub moieties to an internal lysine residue in Ub.
- K48-linked poly-Ub chains are recognized by the 26S proteasome, resulting in degradation of the substrate and recycling of Ub.
Monoubiquitination or K63-linked polyubiquitination plays a number of regulatory roles in cells that are proteasome-independent.
Parkin
Loss-of-function mutations in parkin, a 465-amino-acid RING-type E3 ligase, were first identified as the cause for autosomal recessive juvenile Parkinsonism (AR-JP) and subsequently found to account for ~50% of all recessively transmitted early-onset PD cases. Interestingly, patients with parkin mutations do not exhibit Lewy body pathology.
Possible pathogenic mechanisms by which impaired UPS components cause neurodegeneration. Genetic mutations or oxidative stress from aging and/or exposure to environmental toxins have been shown to impair the ubiquitination machinery (particularly E3 ubiquitin-protein ligases) and deubiquitinating enzymes (DUBs), resulting in abnormal ubiquitination. Depending on the type of ubiquitination affected, the impairment could cause neurodegeneration through two different mechanisms.
- aberrant K48-linked polyubiquitination resulting from impaired E3s or DUBs alters protein degradation by the proteasome, leading to accumulation of toxic proteins and subsequent neurodegeneration. The proteasomes could be directly damaged by oxidative stress or might be inhibited by protein aggregation, which exacerbates the neurotoxicity.
- aberrant monoubiquitination or K63-linked polyubiquitination resulting from impaired E3s or DUBs alters crucial non-proteasomal functions, such as gene transcription and protein trafficking, thereby causing neurodegeneration without protein aggregation.
These two models are not mutually exclusive because a single E3 or DUB enzyme, such as parkin or UCH-L1, could regulate more than one type of ubiquitination. In addition, abnormal ubiquitination and neurodegeneration could also result from mutation or oxidative stress-induced structural changes in the protein substrates that alter their recognition and degradation by the UPS.
Lian Li and Chin LS. IMPAIRMENT OF THE UBIQUITIN-PROTEASOME SYSTEM: A COMMON PATHOGENIC MECHANISM IN NEURODEGENERATIVE DISORDERS. In The Ubiquitin Proteasome System…Chapter 23. (Eds: Eds: Mario Di Napoli and Cezary Wojcik) 553-577 © 2007 Nova Science Publishers, Inc. ISBN 978-1-60021-749-4.
filedesc Schematic diagram of the ubiquitylation system. Created by Roger B. Dodd (Photo credit: Wikipedia)
Current Noteworthy Work
Statins inhibit HMG-CoA reductase, a key enzyme in cholesterol synthesis, and are widely used to treat hypercholesterolemia.
These drugs can lead to a number of side effects in muscle, including muscle fiber breakdown; however, the mechanisms of muscle injury by statins are poorly understood. We report that lovastatin induced the expression of atrogin-1, a key gene involved in skeletal muscle atrophy, in humans with statin myopathy, in zebrafish embryos, and in vitro in murine skeletal muscle cells. In cultured mouse myotubes, atrogin-1 induction following lovastatin treatment was accompanied by distinct morphological changes, largely absent in atrogin-1 null cells. In zebrafish embryos, lovastatin promoted muscle fiber damage, an effect that was closely mimicked by knockdown of zebrafish HMG-CoA reductase. Moreover, atrogin-1 knockdown in zebrafish embryos prevented lovastatin-induced muscle injury. Finally, overexpression of PGC-1α, a transcriptional coactivator that induces mitochondrial biogenesis and protects against the development of muscle atrophy, dramatically prevented lovastatin-induced muscle damage and abrogated atrogin-1 induction both in fish and in cultured mouse myotubes. Collectively, our human, animal, and in vitro findings shed light on the molecular mechanism of statin-induced myopathy and suggest that atrogin-1 may be a critical mediator of the muscle damage induced by statins.
Inami Y, Waguri S, Sakamoto A, Kouno T, et al. Persistent activation of Nrf2 through p62 in hepatocellular carcinoma cells. J. Cell Biol. 2011; 193(2): 275–284. www.jcb.org/cgi/doi/10.1083/jcb.201102031
Macroautophagy (hereafter referred to as autophagy) is a cellular degradation system in which cytoplasmic components, including organelles, are sequestered by double membrane structures called autophagosomes and the sequestered materials are degraded by lysosomal hydrolases for supply of amino acids and for cellular homeostasis. Although autophagy has generally been considered nonselective, recent studies have shed light on another indispensable role for basal autophagy in cellular homeostasis, which is mediated by selective degradation of a specific substrate(s). p62 is a ubiquitously expressed cellular protein that is conserved in metazoa but not in plants and fungi, and recently it has been known as one of the selective substrates for autophagy.
This protein is localized at the autophagosome formation site and directly interacts with LC3, an autophagosome localizing protein . Subsequently, the p62 is incorporated into the autophagosome and then degraded. Therefore, impaired autophagy is accompanied by accumulation of p62 followed by the formation of p62 and ubiquitinated protein aggregates because of the nature of both self- oligomerization and ubiquitin binding of p62.
Epicrisis
This extensive review leaves little left unopened. We have seen the central role that the UPS system plays in normal organelle proteolysis in concert with autophagy. Impaired ubiquitination occurs from aging, and/or toxins, under oxidative stress involving E3s or DUBs.
This leads to altered gene transcripton, altered protein trafficking, and plays a role in neurodegenative disease, and muscle malfunction.
English: A cartoon representation of a lysine 48-linked diubiquitin molecule. The two ubiquitin chains are shown as green cartoons with each chain labelled. The components of the linkage are indicated and shown as orange sticks. Image was created using PyMOL from PDB id 1aar. (Photo credit: Wikipedia)
Different forms of protein ubiquitylation (Photo credit: Wikipedia)
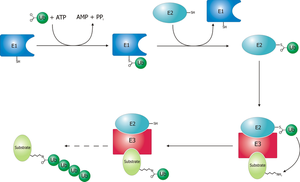
filedesc Schematic diagram of the ubiquitylation system. Created by Roger B. Dodd (Photo credit: Wikipedia)
English: Structure of the PARK2 protein. Based on PyMOL rendering of PDB 1iyf. (Photo credit: Wikipedia)
Related articles
Expanding the Genetic Alphabet and linking the genome to the metabolome. pharmaceuticalintelligence.files.wordpress.com
http://pharmaceuticalintelligence.com/Expanding the Genetic Alphabet and linking the genome to the metabolome
Mitochondria: Origin from oxygen free environment, role in aerobic glycolysis, metabolic adaptation. pharmaceuticalintelligence.files.wordpress.com
http://pharmaceuticalintelligence.com/Mitochondria: Origin from oxygen free environment, role in aerobic glycolysis, metabolic adaptation/
Nitric Oxide and Sepsis, Hemodynamic Collapse, and the Search for Therapeutic Options. pharmaceuticalintelligence.files.wordpress.com
http://pharmaceuticallintelligence.com/Nitric Oxide and Sepsis, Hemodynamic Collapse, and the Search for Therapeutic Options/
Is the Warburg Effect the cause or the effect of cancer: A 21st Century View? pharmaceuticalintelligence.files.wordpress.com
http://pharmaceuticalintelligence.com/Is the Warburg Effect the cause or the effect of cancer: A 21st Century View?/
Breast Cancer, drug resistance, and biopharmaceutical targets. pharmaceuticalintelligence.files.wordpress.com
http://pharmaceuticalintelligence.com/Breast Cancer, drug resistance, and biopharmaceutical targets/
Crucial role of Nitric Oxide in Cancer. pharmaceuticalintelligence.files.wordpress.com
http://pharmaceuticalintelligence.com/Crucial role of Nitric Oxide in Cancer/
Reveals from ENCODE project will invite high synergistic collaborations to discover specific targets. pharmaceuticalintelligence.files.wordpress.com
http://pharmaceuticalintelligence.com/Reveals from ENCODE project will invite high synergistic collaborations to discover specific targets/
The Ubiquitin Ligase Nedd4-1 Participates in Denervation-Induced Skeletal Muscle Atrophy in Mice (plosone.org)
Polyglutamine ‘toxicity’ (dundeemedstudentnotes.wordpress.com)
Inhibition of Xanthine Oxidase by Allopurinol Prevents Skeletal Muscle Atrophy: Role of p38 MAPKinase and E3 Ubiquitin Ligases (plosone.org)
Ubiquitin Dynamics in Complexes Reveal Molecular Recognition Mechanisms Beyond Induced Fit and Conformational Selection (ploscompbiol.org)
Molecule shows effectiveness against drug-resistant myeloma (zedie.wordpress.com)
Related articles
- Betulinic Acid Selectively Increases Protein Degradation and Enhances Prostate Cancer-Specific Apoptosis: Possible Role for Inhibition of Deubiquitinase Activity (plosone.org)
- The N-Terminal Region of Nurr1 (a.a 1 – 31) Is Essential for Its Efficient Degradation by the Ubiquitin Proteasome Pathway (plosone.org)
- Peptide found to induce autophagy resulting in defense against diseases (medicalxpress.com)
- BHD and apoptosis: a review (bhdresearch.scienceblog.com)
- Just One Position-Independent Lysine Residue Can Direct MelanA into Proteasomal Degradation following N-Terminal Fusion of Ubiquitin (plosone.org)
- OTUD7B Protects Enzyme That Controls Pathway Tied To Autoimmune Disease, Cancer (medicalnewstoday.com)
- Intracellular Caspase-Modulating Chimeric Antigen Receptor (readingthinkingandwriting.com)
- New
Related articles
- The N-Terminal Region of Nurr1 (a.a 1 – 31) Is Essential for Its Efficient Degradation by the Ubiquitin Proteasome Pathway (plosone.org)
- BHD and apoptosis: a review (bhdresearch.scienceblog.com)
- Immunotherapy for neurodegenerative diseases: Focus on α-synucleinopathies (scicombinator.com)
- Scientists uncover potential drug target to block cell death in Parkinson’s disease (sciencedaily.com)







![A Pot[age] to Die For A Pot[age] to Die For](https://i0.wp.com/farm5.static.flickr.com/4102/4874343314_6aa1f56fd4_m.jpg?resize=240%2C180)


