See on Scoop.it – Cardiovascular and vascular imaging
See on www.technologyreview.com
May 16, 2013 by 2012pharmaceutical
Posted in Uncategorized | Leave a Comment »
See on Scoop.it – Cardiovascular Disease: PHARMACO-THERAPY
If you check WebMD, Google or various online forums for answers before a doctor visit, or in substitution of one, you’re not alone.
See on mashable.com
Posted in Uncategorized | Leave a Comment »
Reporter: Aviva Lev-Ari, PhD, RN
See on Scoop.it – Cardiovascular Disease: PHARMACO-THERAPY
This year IBM dedicated its Five in Five series (an annual list of five technologies that are likely to advance dramatically) solely to sensors.
Digital sensors of the touch, sight,hearing, taste and smell kind along with their potential are all profiled by IBM Sensor technology is going through a renaissance as companies develop smart and innovative new ways to track data using them.
Sensor innovation is in-part driving the Digital Health Revolution as digital health companies find ingenius ways to integrate them in to apps, devices and other peripherals. The smartphone will play an increasing important role in all of this as they go from having six built-in sensors currently to having sixteen in the next five years.
If these predictions are correct then the next five years will be half-a-decade of sensor proliferation meaning the Digital Health Ecosystem will grow exponentially. In the meantime though there are already a plethora of digital health sensors in use or in the pipeline that are helping people improve and, in some instances, save lives.
See on bionic.ly
Posted in Digital HealthCare – biotech & internet joint ventures | Leave a Comment »
May 16, 2013 by 2012pharmaceutical
Posted in Uncategorized | Leave a Comment »
May 15, 2013 by 2012pharmaceutical
Author, and Content Consultant to e-SERIES A: Cardiovascular Diseases: Justin Pearlman, MD, PhD, FACC
Author and Curator: Larry H Bernstein, MD, FACP
and
Curator: Aviva Lev-Ari, PhD, RN
This article has the following FIVE parts:
1. Forecasting the Impact of Heart Failure in the United States : A Policy Statement From the American Heart Association
2. A Case Study from the GENETIC CONNECTIONS — In The Family: Heart Disease Seeking Clues to Heart Disease in DNA of an Unlucky Family
3. Arterial Stiffness and Cardiovascular Events : The Framingham Heart Study
4. Arterial Elasticity in Quest for a Drug Stabilizer: Isolated Systolic Hypertension
caused by Arterial Stiffening Ineffectively Treated by Vasodilatation Antihypertensives
5. Clinical Decision Support Systems: Realtime Clinical Expert Support — Biomarkers of Cardiovascular Disease : Molecular Basis and Practical Considerations
PA Heidenreich, NM Albert, LA Allen, DA Bluemke, J Butler, et al. Circulation: Heart Failure 2013;6.
Print ISSN: 1941-3289, Online ISSN: 1941-3297.
Heart failure (HF) poses a major burden on productivity and cost of national healthcare expenditures
As the population ages, the prevalence of HF is expected to increase.
The purpose of this report is to
Prevalence estimates for HF were determined from
|
Year |
All ages |
18-44 y |
45-64 y |
65-79 y |
> 80 |
| 2012 | 5 813 262 | 396 578 | 1 907 141 | 2 192 233 | 1 317 310 |
| 2015 | 6 190 606 | 402 926 | 1 949 669 | 2 483 853 | 1 354 158 |
| 2020 | 6 859 623 | 417 600 | 1 974 585 | 3 004 002 | 1 463 436 |
| 2025 | 7 644 674 | 434 635 | 1 969 852 | 3 526 347 | 1 713 840 |
| 2030 | 8 489 428 | 450 275 | 2 000 896 | 3 857 729 | 2 180 528 |
The future costs of HF were estimated by methods developed by the American Heart Association
The model does this by assuming that
(1) HF prevalence percentages will remain constant by age, sex, and race/ethnicity;
(2) the costs of technological innovation will rise at the current rate.
HF prevalence and costs (direct and indirect) were projected using the following steps:
1. HF prevalence and average cost per person were estimated by age group (18–44, 45–64, 65–79, ≥80 years), gender (male, female), and race/ethnicity (white non-Hispanic, white Hispanic, black, other) [32]. The initial HF cost per person and rate of increase in cost was determined for each demographic group, as a percentage of total healthcare expeditures.
2. Inflation is separately addressed by correcting dollar values from Medical Expenditure Panel Survey (MEPS) to 2010 dollars.
3. Nursing home spending triggered an adjustment. The estimates project the incremental cost of care attributable to heart failure (HF).
4. Total HF population prevalence and costs were projected by multiplying the US Census–projected population of each demographic group by the percentage prevalence and average cost
5. The total work loss and home productivity loss costs were generated by multiplying per capita work days lost attributable to HF by (1) prevalence of HF, (2) the probability of employment given HF (for work loss costs only), (3) mean per capita daily earnings, and (4) US Census population projection counts.
Indirect costs of lost productivity from morbidity and premature mortality were estimated as detailed below.
Morbidity costs represent the value of lost earnings attributable to HF and include loss of work among
Projections of Total Cost of Care ($ Billions) for HF for Different Age Groups of the US Population
| Year | All | 18–44 | 45–64 | 65–79 | ≥ 80 |
| 2012 | |||||
| Medical | 20.9 | 0.33 | 3.67 | 8.46 | 8.42 |
| Indirect: Morbidity | 5.42 | 0.52 | 1.92 | 2.05 | 0.93 |
| Indirect: Mortality | 4.35 | 0.66 | 2.53 | 0.98 | 0.18 |
| Total | 30.7 | 1.51 | 8.12 | 11.5 | 9.53 |
| 2020 | |||||
| Medical | 31.1 | 0.43 | 4.58 | 14.2 | 11.8 |
| Indirect: Morbidity | 7.09 | 0.66 | 2.20 | 3.11 | 1.12 |
| Indirect: Mortality | 5.39 | 0.79 | 2.89 | 1.49 | 0.22 |
| Total | 43.6 | 1.88 | 9.67 | 18.8 | 13.2 |
| 2030 | |||||
| Medical | 53.1 | 0.59 | 5.86 | 23.3 | 23.4 |
| Indirect: Morbidity | 9.80 | 0.91 | 2.54 | 4.48 | 1.87 |
| Indirect: Mortality | 6.84 | 0.98 | 3.32 | 2.16 | 0.37 |
| Total | 69.7 | 2.48 | 11.7 | 29.9 | 25.6 |
Excludes HF care costs that have been attributed to comorbid conditions.
Total medical costs are projected to increase from $20.9 billion in 2012 to $53.1 billion in 2030, a 2.5-fold increase. Assuming continuation of current hospitalization practices, the majority (80%) of the costs stem from
Direct costs (cost of medical care) are expected to increase at a faster rate than indirect costs because of premature deaths and lost productivity.
The total cost of HF (direct and indirect costs) is expected to increase in 2030 from the current $30.7 billion to at least $69.8 billion. This will amount to $244 for every US adult in 2030.
Thus the burden of HF for the US healthcare system will grow substantially during the next 18 years if current trends continue.
It is estimated that
Projections can be lowered if action is taken to reduce the health and economic burden of HF. Strategies, plans, and implementation to prevent HF and improve the efficiency of care are needed.
If the projections for accelerating HF costs are to be avoided, attention to the different causes of HF and their risk factors is warranted.
HF is a clinical syndrome that results from a variety of cardiac disorders
HF generally causes symptoms:
In the Western world the predominant causes of HF are:
In 2001, the American College of Cardiology and AHA practice guidelines for chronic HF promoted a classification system that encompasses 4 stages of HF.
Stages A and B are considered precursors to the clinical HF and are meant
Stage A patients have risk factors for HF hypertension, atherosclerotic heart disease, and/or diabetes mellitus.
Patients with stage B are asymptomatic patients who have developed structural heart disease from a variety of potential insults to the heart muscle such as myocardial infarction or valvular heart disease.
Stages C and D represent the symptomatic phases of HF, with stage C manageable and stage D failing medical management, resulting in marked symptoms at rest or with minimal activity despite optimal medical therapy.
Therapeutic interventions include:
Classic demographic risk factors for the development of HF include
Diabetes mellitus, insulin resistance, and obesity are also linked to HF development,
Smoking remains the single largest preventable cause of disease and premature death in the United States.
In multiple studies, failures to apply evidence-based management strategies are blamed for avoidable hospitalizations and/or deaths from HF
Improved implementation of guidelines can delay, mitigate or prevent the onset of HF, and improve survival. Performance improvement programs have facilitated the implementation of evidence-based therapies in both hospital and ambulatory care settings.
Care transition programs by hospitals have become more widespread
The interventions used by these programs include
In multiple studies,adherence to the HF plan of care was associated with reduced all-cause mortality as well as HF hospitalization.
It is anticipated that care transition programs may increase appropriate admissions while decreasing inappropriate admissions
This would have a potentially benenficial impact on the 30-day all-cause readmission rate that has become
More than a quarter of Medicare spending occurs in the last year of life, and
Improving end-of-life care cost effectiveness for patients with stage D HF will require ongoing
Palliative care, including formal hospice care, is increasingly advocated for patients with advanced HF.
Offering palliative care to patients with HF may lead to
The use of hospice services is growing among the HF population,
A recent study of patients in hospice care found that
1. Increasing incidence and costs of care for heart failure projected from 2012 to 2030
2. Direct costs rising at greater rate than indirect costs
3. American Heart Association has defined 4 stages of HF, the last 2 of which are advanced
4. Stages C & D are clinically overt and contribute to rehospitalization
5. Stage D accounts for a significant use of end-of-life hospice care
6. There are evidence-based guidelines for the provision of coordinated care that are not widely applied at present
1. If stages A & B are under the radar, then what measures can best trigger the use of evidence-based guidelines for care?
2. Why are evidence-based guidelines commonly not deployed?
The arguments for introducing coordinated care and for evidence-based guidelines is strong.
Arguments AGAINST slavish imposition of evidence based medicine include genetic individuality (what is best on average is not necessarily best for each genetically and behaviorly distinct individual). Strict adherence to evidence-based guidelines also stifles innovative explorations. None-the-less, deviations from evidence-based plans should be cautious, well-documented, and well-informed, not due to mal-aligned incentives, ignorance, carelessness or error.
The question of when and how to intervene most cost effectively is unanswered. If some patients are salt-sensitive as a contribution to the prevalence of hypertension and heart failure, should EVERYONE be salt restricted or should there be a more concerted effort to define who is salt sensitive? What if it proved more cost-effective to restrict salt intake for everyone, even though many might be fine with high sodium intake, and some might even benefit from or require high sodium intake? Is it reasonable to impose costs, hurdles, even possible harm on some as a cheaper way to achieve “greater good”?
These issues are highly relevant to the proposed emphasis on holistic solutions.
By GINA KOLATA 2013.05.13 New York Times
Scientists are studying the genetic makeup of the Del Sontro family for
Robin Ashwood, one of Mr. Del Sontro’s sisters, found out she had extensive heart disease even though her electrocardiograms was normal. Six of her seven siblings also have heart disease, despite not having any of the traditional risk factors. Then, after a sister, just 47 years old, found out she had advanced heart disease, Mr. Del Sontro, then 43, went to a cardiologist. An X-ray of his arteries revealed the truth. Like his grand-father, his mother, his four brothers and two sisters, he had heart disease.
Now he and his extended family have joined an extraordinary federal research project that is using genetic sequencing to find factors that increase the risk of heart disease beyond the usual suspects — high cholesterol, high blood pressure, smoking and diabetes.“We don’t know yet how many pathways there are to heart disease,” said Dr. Leslie Biesecker, who directs the study Mr. Del Sontro joined. “That’s the power of genetics. To try and dissect that.”
“I had bought the dream: if you just do the right things and eat the right things, you will be O.K.,” said Mr. Del Sontro, whose cholesterol and blood pressure are reassuringly low.
GF Mitchell, Shih-Jen Hwang, RS Vasan, MG Larson.
Circulation. 2010;121:505-511. http://circ.ahajournals.org/content/121/4/505
http://dx.doi.org/10.1161/CIRCULATIONAHA.109.886655
Various measures of arterial stiffness and wave reflection have been proposed as cardiovascular risk markers.
Prior studies have not assessed relations of a comprehensive panel of stiffness measures to prognosis.
First-onset major cardiovascular disease events in relation to arterial stiffness
were analyzed in 2232 participants (mean age, 63 years; 58% women) in the Framingham Heart Study by a proportional hazards model. During median follow-up of 7.8 (range, 0.2 to 8.9) years,
In multivariable models adjusted for
higher aortic PWV was associated with a 48% increase in cardiovascular disease risk (95% confidence interval, 1.16 to 1.91 per SD; P 0.002).
After PWV was added to a standard risk factor model, integrated discrimination improvement was 0.7% (95% confidence interval, 0.05% to 1.3%; P 0.05).
In contrast,
were not related to cardiovascular disease outcomes in multivariable models.
Higher aortic stiffness assessed by PWV
Aortic PWV improves risk prediction when added to standard risk factors and may represent
We shall here visit a recent article by Justin D. Pearlman and Aviva Lev-Ari, PhD, RN, on
Pros and Cons of Drug Stabilizers for Arterial Elasticity as an Alternative or Adjunct to Diuretics and Vasodilators in the Management of Hypertension, titled
Speaking at the 2013 International Conference on Prehypertension and Cardiometabolic Syndrome, meeting cochair Dr Reuven Zimlichman (Tel Aviv University, Israel) argued that there is a growing number of patients for whom the conventional methods are inappropriate for
Most antihypertensives today work by producing vasodilation or decreasing blood volume which may be
In the future, he predicts, “we will have to start looking for a totally different medication that will aim to
Those are not the aim of any group of antihypertensive medications today.”
Zimlichman believes existing databases could be used to develop algorithms that focus on
He also points out that
http://www.theheart.org/article/1502067.do
A related article was published on the relationship between arterial stiffening and primary hypertension.
KH Pettersen, SM Bugenhagen, J Nauman, DA Beard, SW Omholt.
By use of empirically well-constrained computer models describing the coupled function of the baroreceptor reflex and mechanics of the circulatory system, we demonstrate quantitatively that
Specifically,
The results suggest that a major target for treating chronic hypertension in the elderly may include
http://arxiv.org/abs/1305.0727v2?goback=%2Egde_4346921_member_240018699
RS Vasan. Circulation. 2006;113:2335-2362
http://dx.doi.org/10.1161/CIRCULATIONAHA.104.482570
http://circ.ahajournals.org/content/113/19/2335
Substantial data indicate that CVD is a life course disease that begins with the evolution of risk factors that contribute to
Subclinical disease culminates in overt CVD. The onset of CVD itself portends an adverse prognosis with greater
Clinical assessment alone has limitations. Clinicians have used additional tools to aid clinical assessment and to enhance their ability to identify the “vulnerable” patient at risk for CVD, as suggested by a recent National Institutes of Health (NIH) panel.
Biomarkers are one such tool to better identify high-risk individuals, to diagnose disease conditions promptly for diagnosis, prognosis, and treatment guidance.
Biological marker (biomarker): A laboratory test value that is objectively measured and evaluated as an indicator of
Type 0 biomarker: A marker of the natural history of a disease
Type I biomarker: A marker that captures the effects of a therapeutic intervention
Type 2 biomarker (surrogate end point): A marker intended to predict outcomes on the basis of
With biomarkers monitoring disease progression or response to therapy, the patient can serve as his or her own control (follow-up values may be compared to baseline values).
Costs may be less important for prognostic markers when they are largely restricted to people with disease (total cost=cost per person x number to be tested, plus down-stream costs). Some biomarkers (e.g., an exercise stress test) may be used for both diagnostic and prognostic purposes.
Generally there are cost differences in establishing a prognostic value versus diagnostic value of a biomarker:
Regardless of the intended use, it is important to remember that biomarkers that do not change disease management
Typically, for a biomarker to change management, it is important to have evidence that risk reduction strategies should vary with biomarker levels, and/or biomarker-guided management achieves advantages over a management scheme that ignores the biomarker levels.
Typically it means that biomarker levels should be modifiable by therapy.
Gil David and Larry Bernstein have developed, in consultation with Prof. Ronald Coifman, in the Yale University Applied Mathematics Program, a software system that is the equivalent of an intelligent Electronic Health Records Dashboard that
The current design of the Electronic Medical Record (EMR) is a
linear presentation of portions of the record
to cite examples.
This allows perusal through a graphical user interface (GUI) that
Examples of data partitions include:
The introduction of a DASHBOARD adds presentation of
about any patient the care giver needing access to the record.
A basic issue for such a tool is what information is presented and how it is displayed.
A determinant of the success of this endeavor is if it
Continuing work is in progress in extending the capabilities with model datasets, and sufficient data based on the assumption that computer extraction of data from disparate sources will, in the long run, further improve this process.
For instance, there is synergistic value in finding coincidence of:
Similarly, the conversion of hematology based data into useful clinical information requires the establishment of problem-solving constructs based on the measured data.
The most commonly ordered test used for managing patients worldwide is the hemogram that often incorporates
While the hemogram has undergone progressive modification of the measured features over time the subsequent expansion of the panel of tests has provided a window into the cellular changes in the
of the formed elements from the blood-forming organ into the circulation. In the hemogram one can view data reflecting the characteristics of a broad spectrum of medical conditions.
Progressive modification of the measured features of the hemogram has delineated characteristics expressed as measurements of
resulting in many characteristic features of classification. In the diagnosis of hematological disorders
Other dimensions are created by considering
The application of rules-based, automated problem solving should provide a valid approach to
The exponential growth of knowledge since the mapping of the human genome enabled by parallel advances in applied mathematics that have not been a part of traditional clinical problem solving.
As the complexity of statistical models has increased
Contemporary statistical modeling has a primary goal of finding an underlying structure in studied data sets.
The development of an evidence-based inference engine that can substantially interpret the data at hand and
could improve clinical decision-making by incorporating into the model
An example of a difficult area for clinical problem solving is found in the diagnosis of Systemic Inflammatory Response Syndrome (SIRS) and associated sepsis. SIRS is a costly diagnosis in hospitalized patients. Failure to diagnose it in a timely manner increases the financial and safety hazard. The early diagnosis of SIRS/sepsis is made by the application of defined criteria by the clinician.
The application of those clinical criteria, however, defines the condition after it has developed, leaving unanswered the hope for
The early diagnosis of SIRS may possibly be enhanced by the measurement of proteomic biomarkers, including
Immature granulocyte (IG) measurement has been proposed as a
The use of such markers, obtained by automated systems in conjunction with innovative statistical modeling, provides
Such a system aims to reduce medical error by utilizing
How we frame our expectations is important. It determines
In the absence of data to support an assumed benefit, there is no proof of validity at whatever cost.
Potential arenas of benefit include:
The problem stated by LL WEED in “Idols of the Mind” (Dec 13, 2006):
“ a root cause of a major defect in the health care system is that, while we falsely admire and extol the intellectual powers of highly educated physicians, we do not search for the external aids their minds require.” Hospital information technology (HIT) use has been focused on information retrieval, leaving
We deal with problems in the interpretation of data presented to the physician, and how the situation could be improved through better
The computer architecture that the physician uses to view the results is more often than not presented
In order to optimize the interface for physician, the system could have a “front-to-back” design, with the call up for any patient
Eugene Rypka contributed greatly to clarifying the extraction of features in a series of articles, which
The method he describes is termed S-clustering, and
He describes S-clustering as extracting features from endogenous data that
The method classifies by taking the number of features with sufficient variety to generate maps.
The mapping is done by
For example, the message for an antibody titer would be converted from 0 + ++ +++ to 0 1 2 3.
Even though there may be a large number of measured values, the variety is reduced by this compression, even though it may represent less information.
The main issue is
We are concerned with
One determines the effectiveness of each variable by its contribution to information gain in the system. The reference or null set is the class having no information. Uncertainty in assigning to a classification can be countered by providing sufficient information.
One determines the effectiveness of each variable by its contribution to information gain in the system. The possibility for realizing a good model for approximating the effects of factors supported by data used
In the last 60 years the application of entropy comparable to
Akaike pioneered recognition that the choice of model influence results in a measurable manner. In particular, a larger number of variables promotes further explanations of variance, such that a model selection criterion is important that penalizes for the number of variables when success is measured by explanation of variance.
Gil David et al. introduced an AUTOMATED processing of the data available to the ordering physician and
For example:
Rudolph RA, Bernstein LH, Babb J: Information-Induction for the diagnosis of
myocardial infarction. Clin Chem 1988;34:2031-2038.
Bernstein LH (Chairman). Prealbumin in Nutritional Care Consensus Group.
Measurement of visceral protein status in assessing protein and energy
malnutrition: standard of care. Nutrition 1995; 11:169-171.
Bernstein LH, Qamar A, McPherson C, Zarich S, Rudolph R. Diagnosis of myocardial infarction:
integration of serum markers and clinical descriptors using information theory.
Yale J Biol Med 1999; 72: 5-13.
Kaplan L.A.; Chapman J.F.; Bock J.L.; Santa Maria E.; Clejan S.; Huddleston D.J.; Reed R.G.;
Bernstein L.H.; Gillen-Goldstein J. Prediction of Respiratory Distress Syndrome using the
Abbott FLM-II amniotic fluid assay. The National Academy of Clinical Biochemistry (NACB)
Fetal Lung Maturity Assessment Project. Clin Chim Acta 2002; 326(8): 61-68.
Bernstein LH, Qamar A, McPherson C, Zarich S. Evaluating a new graphical ordinal logit method
(GOLDminer) in the diagnosis of myocardial infarction utilizing clinical features and laboratory
data. Yale J Biol Med 1999; 72:259-268.
Bernstein L, Bradley K, Zarich SA. GOLDmineR: Improving models for classifying patients with
chest pain. Yale J Biol Med 2002; 75, pp. 183-198.
Ronald Raphael Coifman and Mladen Victor Wickerhauser. Adapted Waveform Analysis as a Tool for Modeling, Feature Extraction, and Denoising. Optical Engineering, 33(7):2170–2174, July 1994.
R. Coifman and N. Saito. Constructions of local orthonormal bases for classification and regression.
C. R. Acad. Sci. Paris, 319 Série I:191-196, 1994.
We have developed a software system that is the equivalent of an intelligent Electronic Health Records Dashboard that provides empirical medical reference and suggests quantitative diagnostics options. The primary purpose is to gather medical information, generate metrics, analyze them in realtime and provide a differential diagnosis, meeting the highest standard of accuracy. The system builds its unique characterization and provides a list of other patients that share this unique profile, therefore
As the model grows and its knowledge database is extended, the diagnostic and the prognostic become more accurate and precise.
We anticipate that the effect of implementing this diagnostic amplifier would result in
The main benefit is a
as illustrated in the following case acquired on 04/21/10. The patient was diagnosed by our system with severe SIRS at a grade of 0.61 .
The patient was treated for SIRS and the blood tests were repeated during the following week. The full combined record of our system’s assessment of the patient, as derived from the further hematology tests, is illustrated below. The yellow line shows the diagnosis that corresponds to the first blood test (as also shown in the image above). The red line shows the next diagnosis that was performed a week later.
The MISSIVE(c) system, by Justin Pearlman, is an alternative approach that includes not only automated data retrieval and reformatting of data for decision support, but also an integrated set of tools to speed up analysis, structured for quality and error reduction, couplled to facilitated report generation, incorporation of just-in-time knowledge and group expertise, standards of care, evidence-based planning, and both physician and patient instruction.
See also in Pharmaceutical Intelligence:
The National Hospital Bill: The Most Expensive Conditions by Payer, 2006. HCUP Brief #59.
Rudolph RA, Bernstein LH, Babb J: Information-Induction for the diagnosis of myocardial infarction. Clin Chem 1988;34:2031-2038.
Bernstein LH, Qamar A, McPherson C, Zarich S, Rudolph R. Diagnosis of myocardial infarction:
integration of serum markers and clinical descriptors using information theory.
Yale J Biol Med 1999; 72: 5-13.
Kaplan L.A.; Chapman J.F.; Bock J.L.; Santa Maria E.; Clejan S.; Huddleston D.J.; Reed R.G.;
Bernstein L.H.; Gillen-Goldstein J. Prediction of Respiratory Distress Syndrome using the Abbott FLM-II amniotic fluid assay. The National Academy of Clinical Biochemistry (NACB) Fetal Lung Maturity Assessment Project. Clin Chim Acta 2002; 326(8): 61-68.
Bernstein LH, Qamar A, McPherson C, Zarich S. Evaluating a new graphical ordinal logit method (GOLDminer) in the diagnosis of myocardial infarction utilizing clinical features and laboratory
data. Yale J Biol Med 1999; 72:259-268.
Bernstein L, Bradley K, Zarich SA. GOLDmineR: Improving models for classifying patients with chest pain. Yale J Biol Med 2002; 75, pp. 183-198.
Ronald Raphael Coifman and Mladen Victor Wickerhauser. Adapted Waveform Analysis as a Tool for Modeling, Feature Extraction, and Denoising.
Optical Engineering 1994; 33(7):2170–2174.
R. Coifman and N. Saito. Constructions of local orthonormal bases for classification and regression. C. R. Acad. Sci. Paris, 319 Série I:191-196, 1994.
W Ruts, S De Deyne, E Ameel, W Vanpaemel,T Verbeemen, And G Storms. Dutch norm data for 13 semantic categoriesand 338 exemplars. Behavior Research Methods, Instruments,
& Computers 2004; 36 (3): 506–515.
De Deyne, S Verheyen, E Ameel, W Vanpaemel, MJ Dry, WVoorspoels, and G Storms. Exemplar by feature applicability matrices and other Dutch normative data for semantic
concepts. Behavior Research Methods 2008; 40 (4): 1030-1048
Landauer, T. K., Ross, B. H., & Didner, R. S. (1979). Processing visually presented single words: A reaction time analysis [Technical memorandum]. Murray Hill, NJ: Bell Laboratories.
Lewandowsky , S. (1991).
Weed L. Automation of the problem oriented medical record. NCHSR Research Digest Series DHEW. 1977;(HRA)77-3177.
Naegele TA. Letter to the Editor. Amer J Crit Care 1993;2(5):433.
Sheila Nirenberg/Cornell and Chethan Pandarinath/Stanford, “Retinal prosthetic strategy with the capacity to restore normal vision,” Proceedings of the National Academy of Sciences.
http://pharmaceuticalintelligence.com/2012/08/13/the-automated-second-opinion-generator/
http://pharmaceuticalintelligence.com/2013/05/04/cardiovascular-diseases-decision-support-
systems-for-disease-management-decision-making/?goback=%2Egde_4346921_member_239739196
http://pharmaceuticalintelligence.com/2012/12/17/big-data-in-genomic-medicine/
http://pharmaceuticalintelligence.com/2012/12/10/identification-of-biomarkers-that-
are-related–to-the-actin-cytoskeleton/
http://pharmaceuticalintelligence.com/2012/08/12/1815/
http://pharmaceuticalintelligence.com/2012/08/15/1946/
http://pharmaceuticalintelligence.com/2013/05/05/bioengineering-of-vascular-and-tissue-models/
The Heart: Vasculature Protection – A Concept-based Pharmacological Therapy including THYMOSIN
Aviva Lev-Ari, PhD, RN 2/28/2013
http://pharmaceuticalintelligence.com/2013/02/28/the-heart-vasculature-protection-a-concept-
based-pharmacological-therapy-including-thymosin/
FDA Pending 510(k) for The Latest Cardiovascular Imaging Technology
Aviva Lev-Ari, PhD, RN 1/28/2013
http://pharmaceuticalintelligence.com/2013/01/28/fda-pending-510k-for-the-latest-
cardiovascular-imaging-technology/
PCI Outcomes, Increased Ischemic Risk associated with Elevated Plasma Fibrinogen not
Platelet Reactivity Aviva Lev-Ari, PhD, RN 1/10/2013
http://pharmaceuticalintelligence.com/2013/01/10/pci-outcomes-increased-ischemic-risk-
associated-with-elevated-plasma-fibrinogen-not-platelet-reactivity/
The ACUITY-PCI score: Will it Replace Four Established Risk Scores — TIMI, GRACE, SYNTAX,
and Clinical SYNTAX Aviva Lev-Ari, PhD, RN 1/3/2013
http://pharmaceuticalintelligence.com/2013/01/03/the-acuity-pci-score-will-it-replace-four-
established-risk-scores-timi-grace-syntax-and-clinical-syntax/
Coronary artery disease in symptomatic patients referred for coronary angiography: Predicted by
Serum Protein Profiles Aviva Lev-Ari, PhD, RN 12/29/2012
http://pharmaceuticalintelligence.com/2012/12/29/coronary-artery-disease-in-symptomatic-
patients-referred-for-coronary-angiography-predicted-by-serum-protein-profiles/
New Definition of MI Unveiled, Fractional Flow Reserve (FFR)CT for Tagging Ischemia
Aviva Lev-Ari, PhD, RN 8/27/2012
http://pharmaceuticalintelligence.com/2012/08/27/new-definition-of-mi-unveiled-
fractional-flow-reserve-ffrct-for-tagging-ischemia/
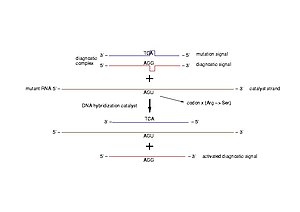
Diagnostic of pathogenic mutations. A diagnostic complex is a dsDNA molecule resembling a short part of the gene of interest, in which one of the strands is intact (diagnostic signal) and the other bears the mutation to be detected (mutation signal). In case of a pathogenic mutation, the transcribed mRNA pairs to the mutation signal and triggers the release of the diagnostic signal (Photo credit: Wikipedia)
Posted in Cardiovascular Pharmacogenomics, Cell Biology, Signaling & Cell Circuits, Computational Biology/Systems and Bioinformatics, FDA Regulatory Affairs, Frontiers in Cardiology and Cardiovascular Disorders, Genomic Testing: Methodology for Diagnosis, Health Economics and Outcomes Research, HealthCare IT, HTN, International Global Work in Pharmaceutical, Medical and Population Genetics, Medical Devices R&D Investment, Medical Imaging Technology, Image Processing/Computing, MRI, CT, Nuclear Medicine, Ultra Sound, Molecular Genetics & Pharmaceutical, Origins of Cardiovascular Disease, Personalized and Precision Medicine & Genomic Research, Pharmaceutical Analytics, Pharmaceutical Industry Competitive Intelligence, Pharmaceutical R&D Investment, Pharmacogenomics, Pharmacotherapy of Cardiovascular Disease, Population Health Management, Genetics & Pharmaceutical, Regulated Clinical Trials: Design, Methods, Components and IRB related issues, Statistical Methods for Research Evaluation, Technology Transfer: Biotech and Pharmaceutical, Uncategorized | Tagged American Heart Association, Cardiovascular disease, Clinical decision support system, Heart disease, Heart Failure | 11 Comments »
Curator: Larry H Bernstein, MD, FCAP
This article is a renewal of a previous discussion on the role of genomics in discovery of therapeutic targets which focused on:
“The Birth of BioInformatics & Computational Genomics” lays the manifold multivariate systems analytical tools that has moved the science forward to a ground that ensures clinical application. Their is a web-like connectivity between inter-connected scientific discoveries, as significant findings have led to novel hypotheses and has driven our understanding of biological and medical processes at an exponential pace owing to insights into the chemical structure of DNA,
In addition, the emergence of methods for
Three-Dimensional Folding and Functional Organization Principles of The Drosophila Genome Sexton T, Yaffe E, Kenigeberg E, Bantignies F,…Cavalli G. Institute de Genetique Humaine, Montpelliere GenomiX, and Weissman Institute, France and Israel. Cell 2012; 148(3): 458-472. http://dx.doi.org/10.1016/j.cell.2012.01.010 http://www.ncbi.nlm.nih.gov/pubmed/22265598 Chromosomes are the physical realization of genetic information and thus form the basis for its
Here we present a high-resolution chromosomal contact map derived from a modified genome-wide chromosome conformation capture approach applied to Drosophila embryonic nuclei. The entire genome is linearly partitioned into well-demarcated physical domains that overlap extensively with
Chromosomal contacts are hierarchically organized between domains. Global modeling of contact density and clustering of domains show that
Together, these observations allow for quantitative prediction of the Drosophila chromosomal contact map, laying the foundation for detailed studies of
“Mr. President; The Genome is Fractal !” Eric Lander (Science Adviser to the President and Director of Broad Institute) et al. delivered the message on Science Magazine cover (Oct. 9, 2009) and generated interest in this by the International HoloGenomics Society at a Sept meeting. First, it may seem to be trivial to rectify the statement in “About cover” of Science Magazine by AAAS. The statement
While the paper itself does not make this statement, the new Editorship of the AAAS Magazine might be even more advanced if the previous Editorship did not reject (without review)
Second, it may not be sufficiently clear for the reader that the reasonable requirement for the
A “knot-free” structure could be spooled by an ordinary “knitting globule” (such that the DNA polymerase does not bump into a “knot” when duplicating the strand; just like someone knitting can go through the entire thread without encountering an annoying knot):
Note, however, that the “strand” can be accessed only at its beginning – it is impossible to e.g.
This is where certain fractals provide a major advantage – that could be the “Eureka” moment. For instance, the mentioned Hilbert-curve is not only “knot free” – but provides an easy access to
If the Hilbert curve starts from the lower right corner and ends at the lower left corner, for instance
Likewise, even the path from the beginning of the Hilbert-curve is about equally easy to access – easier than to reach from the origin a point that is about 2/3 down the path. The Hilbert-curve provides an easy access between two points within the “spooled thread”; from a point that is about 1/5 of the overall length to about 3/5 is also in a “close neighborhood”. This marvellous fractal structure is illustrated by the 3D rendering of the Hilbert-curve. Once you observe such fractal structure,
It will dawn on you that the genome is orders of magnitudes more finessed than we ever thought so. Those embarking at a somewhat complex review of some historical aspects of the power of fractals may wish to consult the ouvre of Mandelbrot (also, to celebrate his 85th birthday). For the more sophisticated readers, even the fairly simple Hilbert-curve (a representative of the Peano-class) becomes even more stunningly brilliant than just some “see through density”. Those who are familiar with the classic “Traveling Salesman Problem” know that “the shortest path along which every given n locations can be visited once, and only once” requires fairly sophisticated algorithms (and tremendous amount of computation if n>10 (or much more). Some readers will be amazed, therefore, that for n=9 the underlying Hilbert-curve helps to provide an empirical solution. refer to pellionisz@junkdna.com Briefly, the significance of the above realization, that the (recursive) Fractal Hilbert Curve is intimately connected to the (recursive) solution of TravelingSalesman Problem, a core-concept of Artificial Neural Networks can be summarized as below. Accomplished physicist John Hopfield (already a member of the National Academy of Science) aroused great excitement in 1982 with his (recursive) design of artificial neural networks and learning algorithms which were able to find solutions to combinatorial problems such as the Traveling SalesmanProblem. (Book review Clark Jeffries, 1991; see J Anderson, Rosenfeld, and A Pellionisz (eds.), Neurocomputing 2: Directions for research, MIT Press, Cambridge, MA, 1990): “Perceptions were modeled chiefly with neural connections in a “forward” direction: A -> B -* C — D. The analysis of networks with strong backward coupling proved intractable. All our interesting results arise as consequences of the strong back-coupling” (Hopfield, 1982). The Principle of Recursive Genome Function surpassed obsolete axioms that blocked, for half a Century, entry of recursive algorithms to interpretation of the structure-and function of (Holo)Genome. This breakthrough,
is particularly important for those who focused on Biological (actually occurring) Neural Networks (rather than abstract algorithms that may not, or because of their core-axioms, simply could not represent neural networks under the governance of DNA information). If biophysicist Andras Pellionisz is correct, genetic science may be on the verge of yielding its third — and by far biggest — surprise. With a doctorate in physics, Pellionisz is the holder of Ph.D.’s in computer sciences and experimental biology from the prestigious Budapest Technical University and the Hungarian National Academy of Sciences. A biophysicist by training, the 59-year-old is a former research associate professor of physiology and biophysics at New York University, author of numerous papers in respected scientific journals and textbooks, a past winner of the prestigious Humboldt Prize for scientific research, a former consultant to NASA and holder of a patent on the world’s first artificial cerebellum, a technology that has already been integrated into research on advanced avionics systems. Because of his background, the Hungarian-born brain researcher might also become one of the first people to successfully launch a new company by
The genes we know about today, Pellionisz says, can be thought of as something similar to machines that make bricks (proteins, in the case of genes), with certain junk-DNA sections providing a blueprint for the different ways those proteins are assembled. The notion that at least certain parts of junk DNA might have a purpose for example, many researchers
In a provisional patent application filed July 31, Pellionisz claims to have
plays in growth — and in life itself. His patent application covers all attempts to
the fractal properties of introns for diagnostic and therapeutic purposes.
| The FractoGene Decade from Inception in 2002 Proofs of Concept and Impending Clinical Applications by 2012Junk DNA Revisited (SF Gate, 2002)The Future of Life, 50th Anniversary of DNA (Monterey, 2003)Mandelbrot and Pellionisz (Stanford, 2004)Morphogenesis, Physiology and Biophysics (Simons, Pellionisz 2005)PostGenetics; Genetics beyond Genes (Budapest, 2006)ENCODE-conclusion (Collins, 2007)The Principle of Recursive Genome Function (paper, YouTube, 2008)You Tube Cold Spring Harbor presentation of FractoGene (Cold Spring Harbor, 2009)Mr. President, the Genome is Fractal! (2009)HolGenTech, Inc. Founded (2010)Pellionisz on the Board of Advisers in the USA and India (2011)ENCODE – final admission (2012) Recursive Genome Function is Clogged by Fractal Defects in Hilbert-Curve (2012) Geometric Unification of Neuroscience and Genomics (2012) US Patent Office issues FractoGene 8,280,641 to Pellionisz (2012) http://www.junkdna.com/the_fractogene_decade.pdf |
To fully understand Pellionisz’ idea, one must first know what a fractal is. Fractals are a way that nature organizes matter. Fractal patterns can be found in anything that has a nonsmooth surface (unlike a billiard ball), such as
Some, but not all, fractals are self-similar and stop repeating their patterns at some stage;
Because they are geometric, meaning they have a shape, fractals can be described in mathematical terms. It’s similar to the way a circle can be described by using a number to represent its radius (the distance from its center to its outer edge). When that number is known, it’s possible to draw the circle it represents without ever having seen it before. Although the math is much more complicated, the same is true of fractals. If one has the formula for a given fractal, it’s possible to use that formula to construct, or reconstruct, an image of whatever structure it represents, no matter how complicated. The mysteriously repetitive but not identical strands of genetic material are in reality
It’s this pattern of fractal instructions, he says, that tells genes what they must do in order to form living tissue, everything from the wings of a fly to the entire body of a full-grown human. In a move sure to alienate some scientists, Pellionisz chose the unorthodox route of making his initial disclosures online on his own Web site. He picked that strategy, he says, because it is the fastest way he can document his claims and find scientific collaborators and investors. Most mainstream scientists usually blanch at such approaches, preferring more traditionally credible methods, such as publishing articles in peer-reviewed journals. Pellionisz’ idea is that a fractal set of building instructions in the DNA plays a role in organizing life itself. Decode the language, and in theory it could be reverse engineered. Just as knowing the radius of a circle lets one create that circle. The fractal-based formula
The idea is encourage new collaborations across the boundaries that separate the intertwined
Hal Plotkin, Special to SF Gate. Thursday, November 21, 2002. http://www.junkdna.com/ http://www.junkdna.com/the_fractogene_decade.pdf http://www.sciencentral.com/articles/view.php3?article_id=218392305 http://www.news-medical.net/health/Junk-DNA-What-is-Junk-DNA.aspx http://www.kurzweilai.net/junk-dna-plays-active-role-in-cancer-progression-researchers-find http://marginalrevolution.com/marginalrevolution/2013/05/the-battle-over-junk-dna http://profiles.nlm.nih.gov/SC/B/B/F/T/_/scbbft.pdf
The human genome: a multifractal analysis. Moreno PA, Vélez PE, Martínez E, et al. BMC Genomics 2011, 12:506. http://www.biomedcentral.com/1471-2164/12/506 Several studies have shown that genomes can be studied via a multifractal formalism. These researchers used a multifractal approach to study the genetic information content of the Caenorhabditis elegans genome. They investigated the possibility that the human genome shows a similar behavior to that observed in the nematode. They report
This behavior correlates strongly on the presence of Alu elements and to a lesser extent on CpG islands and (G+C) content.
They propose a descriptive non-linear model for the structure of the human genome. This model reveals a multifractal regionalization where many regions coexist that are
Given the role of Alu sequences in
these quantifications are especially useful.
Bun C, Ziccardi W, Doering J and Putonti C. Evolutionary Bioinformatics 2012:8 293-300. http://dx.doi.org/10.4137/EBO.S9248 Repetitive elements within genomic DNA are both functionally and evolution-wise informative. Discovering these sequences ab initio is computationally challenging, compounded by the fact that sequence identity between repetitive elements can vary significantly. These investigators present a new application, the Monomer Identification and Isolation Program (MiIP),
To compare MiIP’s performance with other repeat detection tools, analysis was conducted for synthetic sequences as well as several a21-II clones and HC21 BAC sequences. The main benefit of MiIP is
The DNA double helix can under certain conditions accommodate
Researchers in the UK presented a complete set of four variant nucleotides that makes it
Natural DNA only forms a triplex if the targeted strand is rich in purines – guanine (G) and adenine (A) – which in addition to the bonds of the Watson-Crick base pairing
Any Cs or Ts in the target strand of the duplex will only bind very weakly, as
Moreover, the recognition of G requires the C in the probe strand to be protonated,
To overcome all these problems, the groups of Tom Brown and Keith Fox at the University of Southampton have developed modified building blocks, and have now completed
They tested the binding of a 19-mer of these designer nucleotides to a double helix target sequence in comparison with the corresponding triplex-forming oligonucleotide made from natural DNA bases. Using fluorescence-monitored thermal melting and DNase I footprinting, the researchers showed that
Tests with mutated versions of the target sequence showed that
DA Rusling et al, Nucleic Acids Res. 2005, 33, 3025 http://nucleicacidsres.com/Rusling_DA KM Vasquez et al, Science 2000, 290, 530 http://Science.org/2000/290.530/Vazquez_KM/ Frank-Kamenetskii MD, Mirkin SM. Annual Rev Biochem 1995; 64:69-95. http://www.annualreviews.org/aronline/1995/Frank-Kamenetski_MD/64.69/ Since the pioneering work of Felsenfeld, Davies, & Rich, double-stranded polynucleotides containing purines in one strand and pydmidines in the other strand [such as poly(A)/poly(U), poly(dA)/poly(dT), or poly(dAG)/ poly(dCT)] have been known to be able to undergo a stoichiometric transition forming a triple-stranded structure containing one polypurine and two poly-pyrimidine strands. Early on, it was assumed that the third strand was located in the major groove and associated with the duplex via non-Watson-Crick interactions now
Triple helices consisting of one pyrimidine and two purine strands were also proposed. However, notwithstanding the fact that single-base triads in tRNA structures were well- documented, triple-helical DNA escaped wide attention before the mid-1980s. The interest in DNA triplexes arose due to two partially independent developments.
These complexes were shown to be triplex structures rather than D-loops, where
A characteristic feature of all these triplexes is that the two
These findings led explosive growth in triplex studies. One can easily imagine numerous “geometrical” ways to form a triplex, and those that have been studied experimentally. The canonical intermolecular triplex consists of either
Triplex formation strongly depends on the oligonucleotide(s) concentration. A single DNA
To comply with the sequence and polarity requirements for triplex formation, such a DNA strand must have a peculiar sequence: It contains a mirror repeat
Such DNA sequences fold into triplex configuration much more readily than do the corresponding intermolecular triplexes, because all triplex forming segments are brought together within the same molecule. It has become clear that both
can be met by DNA target sequences built of clusters of purines and pyrimidines. The third strand consists of adjacent homopurine and homopyrimidine blocks forming Hoogsteen hydrogen bonds with purines on alternate strands of the target duplex, and
These structures, called alternate-strand triplexes, have been experimentally observed as both intra- and inter-molecular triplexes. These results increase the number of potential targets for triplex formation in natural DNAs somewhat by adding sequences composed of purine and pyrimidine clusters, although arbitrary sequences are still not targetable because
References: Lyamichev VI, Mirkin SM, Frank-Kamenetskii MD. J. Biomol. Stract. Dyn. 1986; 3:667-69. http://JbiomolStractDyn.com/1986/Lyamichev_VI/3.667/ Filippov SA, Frank-Kamenetskii MD. Nature 1987; 330:495-97. http://Nature.com/1987/Fillipov_SA/330.495/ Demidov V, Frank-Kamenetskii MD, Egholm M, Buchardt O, Nielsen PE. Nucleic Acids Res. 1993; 21:2103-7. http://NucleicAcidsResearch.com/1993/Demidov_V/21.2103/ Mirkin SM, Frank-Kamenetskii MD. Anna. Rev. Biophys. Biomol. Struct. 1994; 23:541-76. http://AnnRevBiophysBiomolecStructure.com/1994/Mirkin_SM/23.541/ Hoogsteen K. Acta Crystallogr. 1963; 16:907-16 http://ActaCrystallogr.com/1963/Hoogsteen_K/16.907/ Malkov VA, Voloshin ON, Veselkov AG, Rostapshov VM, Jansen I, et al. Nucleic Acids Res. 1993; 21:105-11. http://NucleicAcidsResearch.com/1993/Malkov_VA/21.105 Malkov VA, Voloshin ON, Soyfer VN, Frank-Kamenetskii MD. Nucleic Acids Res. 1993; 21:585-91 http://NucleicAcidsRes.com/1993/Malkov_VA/21.585/ Chemy DY, Belotserkovskii BP, Frank-Kamenetskii MD, Egholm M, Buchardt O, et al. Proc. Natl. Acad. Sci. USA 1993; 90:1667-70 http://PNAS.org/1993/Chemy_DY/90.1667/ Triplex forming oligonucleotides Triplex forming oligonucleotides: sequence-specific tools for genetic targeting. Knauert MP, Glazer PM. Human Molec Genetics 2001; 10(20):2243-2251. http://HumanMolecGenetics.com/2001/Knauert_ MP/10.2243/ Triplex forming oligonucleotides (TFOs) bind in the major groove of duplex DNA with a
Because of these characteristics, TFOs have been proposed as
These investigators review work demonstrating the ability of TFOs and related molecules
TFOs can mediate targeted gene knock out in mice, providing a foundation for potential
Novagon DNA John Allen Berger, founder of Novagon DNA and The Triplex Genetic Code Over the past 12+ years, Novagon DNA has amassed a vast array of empirical findings which
We propose that our new Novagon DNA 6 nucleotide Triplex Genetic Code has more validity than
Our goal is to conduct a “world class” validation study to replicate and extend our findings.
An Integrated Statistical Approach to Compare Transcriptomics Data Across Experiments: A Case Study on the Identification of Candidate Target Genes of the Transcription Factor PPARα Ullah MO, Müller M and Hooiveld GJEJ. Bioinformatics and Biology Insights 2012;6: 145–154. http://dx.doi.org/10.4137/BBI.S9529 http://www.ncbi.nlm.nih.gov/pubmed/22783064 Corresponding author email: guido.hooiveld@wur.nl http://edepot.wur.nl/213859 An effective strategy to elucidate the signal transduction cascades activated by a transcription factor
Many statistical tests have been proposed for analyzing gene expression data, but
Since the analysis of microarrays involves the testing of multiple hypotheses within one study,
However, this may be an inappropriate metric for
Here we propose the simultaneous testing and integration of
These three contrasts test for the effect of a treatment in
We compare differential expression of genes across experiments while
Vincent Van Buren & Hailin Chen. Scientific Reports 2012; 2, Article number: 1011 http://dx.doi.org/10.1038/srep01011 The complexity of biomolecular interactions and influences is a major obstacle
Visualizing knowledge of biomolecular interactions increases
The rapidly changing landscape of high-content experimental results also presents a challenge
Distributing the responsibility for maintenance of a knowledgebase to a community of
Cognoscente serves these needs
Imputing functional associations
Comprehension of the complexity of this corpus of knowledge will be facilitated by effective
Cognoscente (http://vanburenlab.medicine.tamhsc.edu/cognoscente.html) was designed and implemented to serve these roles as a knowledgebase and as
Cognoscente currently contains over 413,000 documented interactions, with coverage across multiple species. Perl, HTML, GraphViz1, and a MySQL database were used in the development of Cognoscente. Cognoscente was motivated by the need to update the knowledgebase of
Satisfying these needs provides a strong foundation for developing new hypotheses about
Several existing tools provide functions that are similar to Cognoscente.
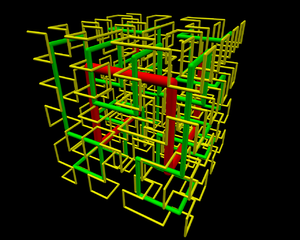
0tj, 1st and 2nd iteration of Hilbert curve in 3D. If you’re looking for the source file, contact me. (Photo credit: Wikipedia)
Posted in Biological Networks, Gene Regulation and Evolution, Chemical Biology and its relations to Metabolic Disease, Chemical Genetics, Computational Biology/Systems and Bioinformatics, Genome Biology, International Global Work in Pharmaceutical, Molecular Genetics & Pharmaceutical, Personalized and Precision Medicine & Genomic Research, Pharmaceutical Analytics, Scientist: Career considerations, Statistical Methods for Research Evaluation, Technology Transfer: Biotech and Pharmaceutical | Tagged artificial neural network, DNA, Eric Lander, FRANCE, Hilbert, Hilbert curve, John Hopfield, National Academy of Science | 2 Comments »
May 15, 2013 by 2012pharmaceutical
Reporter: Aviva Lev-Ari, PhD, RN
|
About 13th Annual Biotech in Europe Investor Forum The 13th Annual Biotech in Europe Investor Forum will be held on 30th September – 1st October 2013 in the Hilton Zurich Airport Hotel, Switzerland. The forum is recognised as the leading international stage for those interested in investing in the biotech and life science industry and is highly transactional, drawing together an exciting cross-section of early-stage/pre-IPO, late-stage and public companies with leading investors, analysts, money managers and pharmas. Supported and designed by leading figures within Europe’s bio industry, this event will once again be covered by our regular media partners. We expect around 350 delegates and 80 presenting companies. The two-day conference programme will give you access to some of the leading players in the industry and will provide you with truly excellent networking opportunities as it boasts an online partnering system with 500 meeting slots available for all attendees to book before the event. The program will comprise of plenary panels including:
Speakers & Chairs confirmed for the event include:
NEW FOR 2013: A series of lectures and panels covering;
Confirmed speakers include:
Presenting Opportunities Presenting at the forum offers excellent opportunities to showcase your company to some of the leading global investors and corporates. It will offer you the opportunity to communicate your projected capital raising plans or simply help you in finding the right partner for your business. Presenting companies from Europe and the US will benefit from specially designed panels and keynote addresses from leading industry figures as well as access to some of the leading analysts and investors from Europe and beyond. This year, we aim to expand the audience and provide once again, opportunities for executive-level networking, deal-making and strategic partnering. The forum is recognised as the leading international stage for those interested in investing in the biotech and life science industry and is highly transactional, drawing together an exciting cross-section of early-stage/pre-IPO, late-stage and public companies with leading investors, analysts, money managers and pharmas. Supported and designed by leading figures within Europe’s bio industry, this event will once again be covered by our regular media partners. Sponsorship Sachs Associates has developed an extensive knowledge of the key individuals operating within the European and global biotech industry. This together with a growing reputation for excellence puts Sachs Associates at the forefront of the industry and provides a powerful tool by which to increase the position of your company in this market. Raise your company’s profile directly with your potential clients. All of our sponsorship packages are tailor made to each client, allowing your organisation to gain the most out of attending our industry driven events. To learn more about presenting, exhibition or sponsorship opportunities, please contact |
SOURCE:
http://www.sachsforum.com/zurich13/zurich13-speakers.html
Posted in Pharmaceutical Industry Competitive Intelligence, Pharmaceutical R&D Investment | Leave a Comment »
May 15, 2013 by sjwilliamspa
Writer and Reporter: Stephen J. Williams, Ph.D.
In the November 23, 2012 issue of Science, Jocelyn Kaiser reports (Genetic Influences On Disease Remain Hidden in News and Analysis)[1] on the difficulties that many genomic studies are encountering correlating genetic variants to high risk of type 2 diabetes and heart disease. At the recent American Society of Human Genetics annual 2012 meeting, results of several DNA sequencing studies reported difficulties in finding genetic variants and links to high risk type 2 diabetes and heart disease. These studies were a part of an international effort to determine the multiple genetic events contributing to complex, common diseases like diabetes. Unlike Mendelian inherited diseases (like ataxia telangiectasia) which are characterized by defects mainly in one gene, finding genetic links to more complex diseases may pose a problem as outlined in the article:
Although many genome-wide-associations studies have found SNPs that have causality to increasing risk diseases such as cancer, diabetes, and heart disease, most individual SNPs for common diseases raise risk by about only 20-40% and would be useless for predicting an individual’s chance they will develop disease and be a candidate for a personalized therapy approach. Therefore, for common diseases, investigators are relying on direct exome sequencing and whole-genome sequencing to detect these medium-rare risk variants, rather than relying on genome-wide association studies (which are usually fine for detecting the higher frequency variants associated with common diseases).
Three of the many projects (one for heart risk and two for diabetes risk) are highlighted in the article:
1. National Heart, Lung and Blood Institute Exome Sequencing Project (ESP)[2]: heart, lung, blood
2. T2D-GENES Consortium: diabetes
Sequenced 5,300 exomes of type 2 diabetes patients and controls from five ancestry groups
SNP in PAX4 gene associated with disease in East Asians
No low-frequency variant with large effect though
3. GoT2D: diabetes
A nice article by Dr. Sowmiya Moorthie entitled Involvement of rare variants in common disease can be found at the PGH Foundation site http://www.phgfoundation.org/news/5164/ further discusses this conundrum, and is summarized below:
“Although GWAs have identified many SNPs associated with common disease, they have as yet had little success in identifying the causative genetic variants. Those that have been identified have only a weak effect on disease risk, and therefore only explain a small proportion of the heritable, genetic component of susceptibility to that disease. This has led to the common disease-common variant hypothesis, which predicts that common disease-causing genetic variants exist in all human populations, but each individual variant will necessarily only have a small effect on disease susceptibility (i.e. a low associated relative risk).
An alternative hypothesis is the common disease, many rare variants hypothesis, which postulates that disease is caused by multiple strong-effect variants, each of which is only found in a few individuals. Dickson et al. in a paper in PLoS Biology postulate that these rare variants can be indirectly associated with common variants; they call these synthetic associations and demonstrate how further investigation could help explain findings from GWA studies [Dickson et al. (2010) PLoS Biol. 8(1):e1000294][3]. In simulation experiments, 30% of synthetic associations were caused by the presence of rare causative variants and furthermore, the strength of the association with common variants also increased if the number of rare causative variants increased. “
Figure from Dr. Moorthie’s article showing the problem of “finding one in many”.
(please click to enlarge)
Indeed, other examples of such issues concerning gene variant association studies occur with other common diseases such as neurologic diseases and obesity, where it has been difficult to clearly and definitively associate any variant with prediction of risk.
For example, Nuytemans et. al.[4] used exome sequencing to find variants in the vascular protein sorting 3J (VPS35) and eukaryotic transcription initiation factor 4 gamma1 (EIF4G1) genes, tow genes causally linked to Parkinson’s Disease (PD). Although they identified novel VPS35 variants none of these variants could be correlated to higher risk of PD. One EIF4G1 variant seemed to be a strong Parkinson’s Disease risk factor however there was “no evidence for an overall contribution of genetic variability in VPS35 or EIF4G1 to PD development”.
These negative results may have relevance as companies such as 23andme (www.23andme.com) claim to be able to test for Parkinson’s predisposition. To see a description of the LLRK2 mutational analysis which they use to determine risk for the disease please see the following link: https://www.23andme.com/health/Parkinsons-Disease/. This company and other like it have been subjects of posts on this site (Personalized Medicine: Clinical Aspiration of Microarrays)
However there seems to be more luck with strategies focused on analyzing intronic sequence rather than exome sequence. Jocelyn Kaiser’s Science article notes this in a brief interview with Harry Dietz of Johns Hopkins University where he suspects that “much of the missing heritability lies in gene-gene interactions”. Oliver Harismendy and Kelly Frazer and colleagues’ recent publication in Genome Biology http://genomebiology.com/content/11/11/R118 support this notion[5]. The authors used targeted resequencing of two endocannabinoid metabolic enzyme genes (fatty-acid-amide hydrolase (FAAH) and monoglyceride lipase (MGLL) in 147 normal weight and 142 extremely obese patients.
These patients were enrolled in the CRESCENDO trial and patients analyzed were of European descent. However, instead of just exome sequencing, the group resequenced exome AND intronic sequence, especially focusing on promoter regions. They identified 1,448 single nucleotide variants but using a statistical filter (called RareCover which is referred to as a collapsing method) they found 4 variants in the promoters and intronic areas of the FAAH and MGLL genes which correlated to body mass index. It should be noted that anandamide, a substrate for FAAH, is elevated in obese patients. The authors did note some issues though mentioning that “some other loci, more weakly or inconsistently associated in the original GWASs, were not replicated in our samples, which is not too surprising given the sample size of our cohort is inadequate to replicate modest associations”.
https://www.youtube.com/watch?v=-Qr5ahk1HEI
REFERENCES
http://www.phgfoundation.org/news/5164/ PHG Foundation
1. Kaiser J: Human genetics. Genetic influences on disease remain hidden. Science 2012, 338(6110):1016-1017.
2. Tennessen JA, Bigham AW, O’Connor TD, Fu W, Kenny EE, Gravel S, McGee S, Do R, Liu X, Jun G et al: Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science 2012, 337(6090):64-69.
3. Dickson SP, Wang K, Krantz I, Hakonarson H, Goldstein DB: Rare variants create synthetic genome-wide associations. PLoS biology 2010, 8(1):e1000294.
4. Nuytemans K, Bademci G, Inchausti V, Dressen A, Kinnamon DD, Mehta A, Wang L, Zuchner S, Beecham GW, Martin ER et al: Whole exome sequencing of rare variants in EIF4G1 and VPS35 in Parkinson disease. Neurology 2013, 80(11):982-989.
5. Harismendy O, Bansal V, Bhatia G, Nakano M, Scott M, Wang X, Dib C, Turlotte E, Sipe JC, Murray SS et al: Population sequencing of two endocannabinoid metabolic genes identifies rare and common regulatory variants associated with extreme obesity and metabolite level. Genome biology 2010, 11(11):R118.
Other posts on this site related to Genomics include:
Cancer Biology and Genomics for Disease Diagnosis
Genomics-based cure for diabetes on-the-way
Personalized Medicine: Clinical Aspiration of Microarrays
Late Onset of Alzheimer’s Disease and One-carbon Metabolism
Genetics of Disease: More Complex is How to Creating New Drugs
Mitochondrial Metabolism and Cardiac Function
Pancreatic Cancer: Genetics, Genomics and Immunotherapy
Quantum Biology And Computational Medicine
Personalized Cardiovascular Genetic Medicine at Partners HealthCare and Harvard Medical School
Consumer Market for Personal DNA Sequencing: Part 4
Personalized Medicine: An Institute Profile – Coriell Institute for Medical Research: Part 3
Whole-Genome Sequencing Data will be Stored in Coriell’s Spin off For-Profit Entity
Posted in Alzheimer's Disease, Biological Networks, Gene Regulation and Evolution, Cardiovascular Pharmacogenomics, Cerebrovascular and Neurodegenerative Diseases, Chemical Biology and its relations to Metabolic Disease, Chemical Genetics, Etiology, Frontiers in Cardiology and Cardiovascular Disorders, Genome Biology, Genomic Testing: Methodology for Diagnosis, Health Economics and Outcomes Research, Molecular Genetics & Pharmaceutical, Personalized and Precision Medicine & Genomic Research, Pharmacogenomics, Reproductive Andrology, Embryology, Genomic Endocrinology, Preimplantation Genetic Diagnosis and Reproductive Genomics | Tagged Alzheimers Disease, American Society of Human Genetics, Aviva Lev-Ari, Biology, Conditions and Diseases, Congenital disorder of glycosylation, DNA, DNA Sequencing, Exome sequencing, Full genome sequencing, gene, genetic variants, genetics, Genome-wide association study, health, Heart disease, metabolic syndrome, Parkinsons disease, Personalized medicine, Single Nucleotide polymorphisms, Single-nucleotide polymorphism, type 2 diabetes | 5 Comments »
May 15, 2013 by tildabarliya
Author: Tilda Barliya PhD
The field of DNA and RNA nanotechnologies are considered one of the most dynamic research areas in the field of drug delivery in molecular medicine. Both DNA and RNA have a wide aspect of medical application including: drug deliveries, for genetic immunization, for metabolite and nucleic acid detection, gene regulation, siRNA delivery for cancer treatment (I), and even analytical and therapeutic applications.
Seeman (6,7) pioneered the concept 30 years ago of using DNA as a material for creating nanostructures; this has led to an explosion of knowledge in the now well-established field of DNA nanotechnology. The unique properties in terms of free energy, folding, noncanonical base-pairing, base-stacking, in vivo transcription and processing that distinguish RNA from DNA provides sufficient rationale to regard RNA nanotechnology as its own technological discipline. Herein, we will discuss the advantages of DNA nanotechnology and it’s use in medicine.
So What is the rational of using DNA nanotechnology(3)?
Structural DNA nanotechnology rests on three pillars: [1] Hybridization; [2] Stably branched DNA; and [3] Convenient synthesis of designed sequences.
Hybridization
Hybridization. The self-association (self=assembly) of complementary nucleic acid molecules or parts of molecules, is implicit in all aspects of structural DNA nanotechnology. Individual motifs are formed by the hybridization of strands designed to produce particular topological species. A key aspect of hybridization is the use of sticky ended cohesion to combine pieces of linear duplex DNA; this has been a fundamental component of genetic engineering for over 35 years (7). Not only is hybridization critical to the formation of structure, but it is deeply involved in almost all the sequence-dependent nanomechanical devices that have been constructed, and it is central to many attempts to build structural motifs in a sequential fashion (7,8 ).
Stably Branched DNA
branched DNA molecules are central to DNA nanotechnology. It is the combination of in vitro hybridization and synthetic branched DNA that leads to the ability to use DNA as a construction material. Such branched DNA is thought to be intermediates in genetic recombination (such as Holliday junctions).
Convenient Synthesis of Designed Sequences
Biologically derived branched DNA molecules, such as Holliday junctions, are inherently unstable, because they exhibit sequence symmetry; i.e., the four strands actually consist of two pairs of strands with the same sequence. This symmetry enables an isomerization known as branch migration that allows the branch point to relocate. DNA nanotechnology entailed sequence design that attempted to minimize sequence symmetry in every way possible.
One of the most remarkable innovations in structural DNA-nanotechnology in recent years is DNA origami, which was invented in 2006 by Paul Rothemund (1) (see Fig above). DNA origami utilizes the genome from a virus together with a large number of shorter DNA strands to enable the creation of numerous DNA-based structures (Figure 1). The shorter DNA strands forces the long viral DNA to fold into a pattern that is defined by the interaction between the long and the short DNA strands (1,2).
Rothemund believes that an application of patterned DNA origami would be the creation of a ‘nanobreadboard’, to which diverse components could be added. The attachment of proteins23, for example, might allow novel biological experiments aimed at modelling complex protein assemblies and examining the effects of spatial organization, whereas molecular electronic or plasmonic circuits might be created by attaching nanowires, carbon nanotubes or gold nanoparticles (1).
DNA nanotechnology and Biological Application
The physical and chemical properties of nanomaterials such as polymers, semiconductors, and metals present diverse advantages for various in vivo applications (3,9 ). For example:
Summary:
DNA nanotechnology is an evolving field that affects medicine, computation, material sciences, and physics. DNA nanostructures offer unprecedented control over shape, size, mechanical flexibility and anisotropic surface modification. Clearly, proper control over these aspects can increase circulation times by orders of magnitude, as can be seen for longcirculating particles such as erythrocytes and various pathogenic particles evolved to overcome this issue. The use of DNA in DNA/protein-based matrices makes these structures inherently amenable to structural tunability. More research in this direction will certainly be developed, making DNA a promising biomaterial in tissue engineering. future development of novel ways in which DNA would be utilized to have a much more comprehensive role in biological computation and data storage is envisaged.
REFERENCES
1. Paul W. K. Rothemund. Folding DNA to create nanoscale shapes and patterns. NATURE 2006 (March 16)|Vol 440: 297-302. http://www.nature.com/nature/journal/v440/n7082/full/nature04586.html
http://www.dna.caltech.edu/Papers/DNAorigami-nature.pdf
2. Andre V. Pinheiro, Dongran Han, William M. Shih and Hao Yan. Challenges and opportunities for structural DNA nanotechnology. Nature Nanotechnology 2011 Dec | VOL 6: 763-772. http://www.nature.com/nnano/journal/v6/n12/pdf/nnano.2011.187.pdf
2b. Thi Huyen La, Thi Thu Thuy Nguyen, Van Phuc Pham, Thi Minh Huyen Nguyen and Quang Huan Le. Using DNA nanotechnology to produce a drug delivery system. Adv. Nat. Sci.: Nanosci. Nanotechnol. 4 (2013) 015002 (7pp). http://iopscience.iop.org/2043-6262/4/1/015002. http://iopscience.iop.org/2043-6262/4/1/015002/pdf/2043-6262_4_1_015002.pdf
3. Muniza Zahid, Byeonghoon Kim, Rafaqat Hussain, Rashid Amin and Sung H Park. DNA nanotechnology: a future perspective. Nanoscale Research Letters 2013, 8:119. http://www.nanoscalereslett.com/content/8/1/119
4.By: Cientifica Ltd 2007. The Nanotech Revolution in Drug Delivery. http://www.cientifica.com/WhitePapers/054_Drug%20Delivery%20White%20Paper.pdf
5. Gemma Campbell. Nanotechnology and its implications for the health of the E.U citizen: Diagnostics, drug discovery and drug delivery. Institute of Nanotechnology and Nanoforum. http://www.nano.org.uk/nanomednet/images/stories/Reports/diagnostics,%20drug%20discovery%20and%20drug%20delivery.pdf
6.Peixuan Guo., Haque F., Brent Hallahan, Randall Reif and Hui Li. Uniqueness, Advantages, Challenges, Solutions, and Perspectives in Therapeutics Applying RNA Nanotechnology. Nucleic Acid Ther. 2012 August; 22(4): 226–245. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3426230/
7. SEEMAN N.C. Nanomaterials based on DNA. Annu. Rev. Biochem. 2010;79:65–87. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3454582/
8. Yin P, Choi HMT, Calvert CR, Pierce NA. Programming biomolecular self-assembly pathways. Nature.2008;451:318–323. http://www.ncbi.nlm.nih.gov/pubmed/18202654
9. Yan Lee P, Wong KY: Nanomedicine: a new frontier in cancer therapeutics. Curr Drug Deliv 2011, 8(3):245-253. http://www.eurekaselect.com/73728/article
10. Qian, L.L., Winfree, E., and Bruck, J. Neural Network Computation with DNA Strand Displacement Cascades. Nature 2011 475, 368-372. http://www.nature.com/nature/journal/v475/n7356/full/nature10262.html
11. Acharya S, Dilnawaz F, Sahoo SK: Targeted epidermal growth factor receptor nanoparticle bioconjugates for breast cancer therapy. Biomaterials 2009, 30(29):5737-5750. http://www.sciencedirect.com/science/article/pii/S0142961209006929
12. Bohunicky B, Mousa SA: Biosensors: the new wave in cancer diagnosis. Nanotechnology, Science and Applications 2011, 4:1-10. http://www.dovepress.com/biosensors-the-new-wave-in-cancer-diagnosis-peer-reviewed-article-NSA-recommendation1
13. Sanvicens N, Mannelli I, Salvador J, Valera E, Marco M: Biosensors for pharmaceuticals based on novel technology. Trends Anal Chem 2011, 30:541-553. http://www.sciencedirect.com/science/article/pii/S016599361100015X
14. Amin R, Kulkarni A, Kim T, Park SH: DNA thin film coated optical fiber biosensor. Curr Appl Phys 2011, 12(3):841-845. http://www.sciencedirect.com/science/article/pii/S1567173911005888
15. Choi, Y.; Baker, J. R. Targeting Cancer Cells with DNA Assembled Dendrimers: A Mix and Match Strategy for Cancer. Cell Cycle 2005, 4, 669–671. http://www.ncbi.nlm.nih.gov/pubmed/15846063 http://www.landesbioscience.com/journals/cc/article/1684/
I. By: Ziv Raviv PhD. The Development of siRNA-Based Therapies for Cancer. http://pharmaceuticalintelligence.com/2013/05/09/the-development-of-sirna-based-therapies-for-cancer/
II. By: Tilda Barliya PhD. Nanotechnology, personalized medicine and DNA sequencing. http://pharmaceuticalintelligence.com/2013/01/09/nanotechnology-personalized-medicine-and-dna-sequencing/
III. By: Larry Bernstein MD FACP. DNA Sequencing Technology. http://pharmaceuticalintelligence.com/2013/03/03/dna-sequencing-technology/
IV. By: Venkat S Karra PhD. Measuring glucose without needle pricks: nano-sized biosensors made the test easy. http://pharmaceuticalintelligence.com/2012/09/04/measuring-glucose-without-needle-pricks-nano-sized-biosensors-made-the-test-easy/
Posted in Bio Instrumentation in Experimental Life Sciences Research, Biological Networks, Gene Regulation and Evolution, Biomarkers & Medical Diagnostics, BioSimilars, CANCER BIOLOGY & Innovations in Cancer Therapy, Cell Biology, Signaling & Cell Circuits, Disease Biology, Small Molecules in Development of Therapeutic Drugs, Drug Delivery Platform Technology, Genomic Testing: Methodology for Diagnosis, Health Economics and Outcomes Research, Medical Devices R&D Investment, Nanotechnology for Drug Delivery, Personalized and Precision Medicine & Genomic Research, Technology Transfer: Biotech and Pharmaceutical | Tagged Biomimic systems, Biosensors, Cancer - General, computational algorithms, DNA, DNA nanotechnology, DNA origami, DNA sequencers, drug delivery system, Energy transfer, hybridization, Nucleic acid, Nucleic acid sequence, RNA, Small interfering RNA, stable branched DNA | 3 Comments »
May 15, 2013 by 2012pharmaceutical
This article has two parts:
Executive Insights, 2013;14(12). http://www.LEK.com
Companion diagnostics and their companion therapies is defined here as a method enabling
The result of this statement is that the diagnostics permitted to specific patient types gives access to
The concept is new because:
(1) The diagnoses may be closely related by classical criteria, but at the same time they are
not alike with respect to efficacy of treatment with a standard therapy.
(2) The companion diagnostics is restricted to dealing with a targeted drug-specific question
without regard to other clinical issues.
(3) The efficacy issue it clarifies is reliant on a deep molecular/metabolic insight that is not available, except through
emergent genomic/proteomic analysis that has become available and which has rapidly declining cost to obtain.
The limitation example given is HER2 testing for use of Herceptin in therapy for non-candidates (HER2 negative patients).
The problem is that the current format is a “one test/one drug” match, but decision support may require a combination of
While HER2 negative patients are more likely to be pre-menopausal with a more aggressive tumor than postmenopausal,
So the Herceptin would be given in combination, but with what other drug in a non-candidate?
The point that L.E.K. makes is that providing highly validated biomarkers linked to approved therapies, it is necessary to pursue more holistic decision support tests that interrogate multiple biomarkers (panels of companion diagnostic markers) and discovery of signatures for treatments that are also used with a broad range of information, such as,
A comprehensive solution of this nature appears to be a distance from realization. However, is this the direction that will lead to tomorrows treatment decision support approaches?
As a starting point, L.E.K. characterized the landscape of available tests in the U.S. that inform treatment decisions compiled from ~50 leading diagnostics companies operating in the U.S. between 2004-2011. L.E.K. identified more than 200 decision support tests that were classified by test purpose, and more specifically, whether tests inform treatment decisions for a single drug/class (e.g., companion diagnostics) vs. more holistic treatment decisions across multiple drugs/classes (i.e., multiagent response tests).
Companion Diagnostics
Single drug/class
Predict response/safety or guide dosing of a single drug or class
HercepTest Dako
Determines HER2 protein overexpression for Herceptin treatment selection
Multiple drugs/classes
Vysis ALK Break
Apart FISH
Abbott Labs Predicts the NSCLC patient response to Xalkori
Other Decision Support
Provide prognostic and predictive information on the benefit of treatment
Oncotype Dx Genomic Health, Inc.
Predicts both recurrence of breast cancer and potential patient benefit to chemotherapy regimens
PML-RARα Clarient, Inc.
Predicts response to all-trans retinoic acid (ATRA) and other chemotherapy agents
TRUGENE Siemens
Measures resistence to multiple HIV-1 anti-retroviral agents
Multi-agent Response
Inform targeted therapy class selection by interrogating a panel of biomarkers
Target Now Caris Life Sciences
Examines tumor’s molecular profile to tailor treatment options
ResponseDX: Lung Response Genetics, Inc.
Examines multiple biomarkers to guide therapeutic treatment decisions for NSCLC patients
Source: L.E.K. Analysis
Includes IVD and LDT tests from
For descriptive purposes only, may not map to exact regulatory labeling
Most tests are companion diagnostics and other decision support tests that provide guidance on
However, holistic decision support tests (e.g., multi-agent response) are growing the fastest at 56% CAGR.
The emergence of multi-agent response tests suggests diagnostics companies are already seeing the need to aggregate individual tests (e.g., companion diagnostics) into panels of appropriate markers addressing a given clinical decision need. L.E.K. believes this trend is likely to continue as
to become deployed in the clinic.
L.E.K. also completed an assessment of publicly available personalized medicine partnership activity from 2009-2011 for ~150 leading organizations operating in the U.S. to look at broader decision support trends and emergence of more holistic solutions beyond diagnostic tests.
Survey of partnerships deals was conducted for
Across the sample we identified 189 publicly announced partnerships of which ~65% focused on more traditional areas (biomarker discovery, companion diagnostics and targeted therapies). However, a significant portion (~30%) included elements geared towards creating more holistic decision support models.
Partnerships categorized as holistic decision support by L.E.K. were focused on
Interestingly, holistic decision support partnerships often included stakeholders outside of biopharma and diagnostics such as
This finding suggests that these new stakeholders will be increasingly important in influencing care decisions going forward.
Holistic Treatment Decision Support
| Holistic Decision Support Focus |
Technology Provider Partners |
Holistic Decision Support Activities |
|
| Molecular Profiling |
Life Technologies |
TGEN/US |
Sequencing of triple-negative breast cancer patients to identify potential treatment strategies |
|
Foundation Medicine |
Novartis |
Deployment of cancer genomics analysis platform to support Novartis clinical research efforts | |
| Predictive genomics |
Clarient, Inc. |
Acorn |
Biomarker profiling of patients within Acorn’s network of providers to support clinical research efforts |
|
GenomeQuest |
Beth Israel Deaconess |
Whole genome analysis and to guide patient management | |
| Outcomes Data Mining |
AstraZeneca |
WellPoint |
Evaluate comparative effectiveness of selected marketed therapies |
|
23andMe |
NIH |
Leverage information linking drug response and CYP2C9/CYP2C19 variation | |
|
Pfizer |
Medco |
Leverage patient genotype, phenotype and outcome for treatment decisions and target therapeutics | |
| Healthcare IT Infrastructure |
IBM |
WellPoint |
Deploy IBM’s Watson-based solution to evidence-based healthcare decision-making support |
|
Oracle |
Moffitt Cancer Center |
Deploy Oracle’s informatics platform to store and manage patient medical information | |
| Data Integration |
Siemens Diagnostics |
Susquehanna Health |
Integration of imaging and laboratory diagnostics |
|
Cernostics |
Geisinger |
Integration of advanced tissue diagnostics, digital pathology, annotated biorepository and EMR to create solutions |
|
| next-generation treatment decision support solutions |
CardioDx |
GE Healthcare |
Integration of genomics with imaging data in CVD |
L.E.K. believes the likely debate won’t center on which models and companies will prevail. It appears that the industry is now moving along the continuum to a truly holistic capability.
The mainstay of personalized medicine today will become integrated and enhanced by other data.
The companies that succeed will be able to capture vast amounts of information
Holistic models will be powered by increasingly larger datasets and sophisticated decision-making algorithms.
This will require the participation of an increasingly broad range of participants to provide the
There are a number of questions posed by this study, but only some are of interest to this discussion:
Group A. Pharmaceuticals and Devices
Group B. Diagnostics and Decision Support
– Testing technologies
– Information (e.g., associations, outcomes, trial databases, records)
– IT infrastructure for data integration and management, simulation and reporting
Group C. Providers and Payers
Gil David and Larry Bernstein have developed, in consultation with Prof. Ronald Coifman,
in the Yale University Applied Mathematics Program,
A software system that is the equivalent of an intelligent Electronic Health Records Dashboard that
The current design of the Electronic Medical Record (EMR) is a linear presentation of portions of the record
This allows perusal through a graphical user interface (GUI) that partitions the information or necessary reports
This requires that the medical practitioner finds the
The introduction of a DASHBOARD has allowed a presentation of
about any patient the care giver needing access to the record.
The advantage of this innovation is obvious. The startup problem is what information is presented and
We are proposing an innovation that supercedes the main design elements of a DASHBOARD and utilizes
So the important determinant of the success of this endeavor is that
Continuing work is in progress in extending the capabilities with model datasets, and sufficient data because
For instance, the finding of both ST depression on EKG coincident with an elevated cardiac biomarker (troponin), particularly in the absence of substantially reduced renal function. The conversion of hematology based data into useful clinical information requires the establishment of problem-solving constructs based on the measured data.
The most commonly ordered test used for managing patients worldwide is the hemogram that often incorporates
While the hemogram has undergone progressive modification of the measured features over time the subsequent expansion of the panel of tests has provided a window into the cellular changes in the
of the formed elements from the blood-forming organ into the circulation. In the hemogram one can view
Progressive modification of the measured features of the hemogram has delineated characteristics expressed as measurements of
resulting in many characteristic features of classification. In the diagnosis of hematological disorders
provide a two dimensional model. Other dimensions are created by considering
The application of rules-based, automated problem solving should provide a valid approach to
The exponential growth of knowledge since the mapping of the human genome enabled by parallel advances in applied mathematics that have not been a part of traditional clinical problem solving.
As the complexity of statistical models has increased
Contemporary statistical modeling has a primary goal of finding an underlying structure in studied data sets.
The development of an evidence-based inference engine that can substantially interpret the data at hand and
could improve clinical decision-making by incorporating
An example of a difficult area for clinical problem solving is found in the diagnosis of SIRS and associated sepsis. SIRS (and associated sepsis) is a costly diagnosis in hospitalized patients. Failure to diagnose sepsis in a timely manner creates a potential financial and safety hazard. The early diagnosis of SIRS/sepsis is made by the application of defined criteria by the clinician.
The application of those clinical criteria, however, defines the condition after it has developed and
The early diagnosis of SIRS may possibly be enhanced by the measurement of proteomic biomarkers, including
Immature granulocyte (IG) measurement has been proposed as a
The use of such markers, obtained by automated systems
Such a system utilizes the conjoined syndromic features of
How we frame our expectations is so important that it determines
In the absence of data to support an assumed benefit, there is no proof of validity at whatever cost.
This has meaning for
The problem stated by LL WEED in “Idols of the Mind” (Dec 13, 2006): “ a root cause of a major defect in the health care system is that, while we falsely admire and extol the intellectual powers of highly educated physicians, we do not search for the external aids their minds require”. HIT use has been
We deal with problems in the interpretation of data presented to the physician, and how through better
The computer architecture that the physician uses to view the results is more often than not presented
In order to optimize the interface for physician, the system would have a “front-to-back” design, with
the call up for any patient ideally consisting of a dashboard design that presents the crucial information
The key point is that each item used has to be closely related to a corresponding criterion needed for a decision.
This further breakdown in the modern era is determined by genetically characteristic gene sequences
that are transcribed into what we measure. Eugene Rypka contributed greatly to clarifying the extraction
of features in a series of articles, which
The method he describes is termed S-clustering, and
He describes S-clustering as extracting features from endogenous data that
The method classifies by taking the number of features
The mapping is done by
The number of messages and the number of choices forms an N-by N table. He points out that the message
Even though there may be a large number of measured values, the variety is reduced
Yet the real issue is how a combination of variables falls into a table with meaningful information. We are concerned with accurate assignment into uniquely variable groups by information in test relationships. One determines the effectiveness of each variable by
The reference or null set is the class having no information. Uncertainty in assigning to a classification is
The possibility for realizing a good model for approximating the effects of factors supported by data used
In the last 60 years the application of entropy comparable to
Gil David et al. introduced an AUTOMATED processing of the data available to the ordering physician and
For example: anemias (iron deficiency, vitamin B12 and folate deficiency, and hemolytic anemia or myelodysplastic syndrome); pneumonia; systemic inflammatory response syndrome (SIRS) with or without bacteremia; multiple organ failure and hemodynamic shock; electrolyte/acid base balance disorders; acute and chronic liver disease; acute and chronic renal disease; diabetes mellitus; protein-energy malnutrition; acute respiratory distress of the newborn; acute coronary syndrome; congestive heart failure; disordered bone mineral metabolism; hemostatic disorders; leukemia and lymphoma; malabsorption syndromes; and cancer(s)[breast, prostate, colorectal, pancreas, stomach, liver, esophagus, thyroid, and parathyroid].
Rudolph RA, Bernstein LH, Babb J: Information-Induction for the diagnosis of myocardial infarction. Clin Chem 1988;34:2031-2038.
Bernstein LH (Chairman). Prealbumin in Nutritional Care Consensus Group.
Measurement of visceral protein status in assessing protein and energy malnutrition: standard of care. Nutrition 1995; 11:169-171.
Bernstein LH, Qamar A, McPherson C, Zarich S, Rudolph R. Diagnosis of myocardial infarction: integration of serum markers and clinical descriptors using information theory. Yale J Biol Med 1999; 72: 5-13.
Kaplan L.A.; Chapman J.F.; Bock J.L.; Santa Maria E.; Clejan S.; Huddleston D.J.; Reed R.G.; Bernstein L.H.; Gillen-Goldstein J. Prediction of Respiratory Distress Syndrome using the Abbott FLM-II amniotic fluid assay. The National Academy of Clinical Biochemistry (NACB) Fetal Lung Maturity Assessment Project. Clin Chim Acta 2002; 326(8): 61-68.
Bernstein LH, Qamar A, McPherson C, Zarich S. Evaluating a new graphical ordinal logit method (GOLDminer) in the diagnosis of myocardial infarction utilizing clinical features and laboratory data. Yale J Biol Med 1999; 72:259-268.
Bernstein L, Bradley K, Zarich SA. GOLDmineR: Improving models for classifying patients with chest pain. Yale J Biol Med 2002; 75, pp. 183-198.
Ronald Raphael Coifman and Mladen Victor Wickerhauser. Adapted Waveform Analysis as a Tool for Modeling, Feature Extraction, and Denoising. Optical Engineering, 33(7):2170–2174, July 1994.
R. Coifman and N. Saito. Constructions of local orthonormal bases for classification and regression. C. R. Acad. Sci. Paris, 319 Série I:191-196, 1994.
We have developed a software system that is the equivalent of an intelligent Electronic Health Records Dashboard that provides empirical medical reference and suggests quantitative diagnostics options.
The primary purpose is to
The system builds its unique characterization and provides a list of other patients that share this unique profile, therefore utilizing the vast aggregated knowledge (diagnosis, analysis, treatment, etc.) of the medical community. The
As the model grows and its knowledge database is extended, the diagnostic and the prognostic become more accurate and precise. We anticipate that the effect of implementing this diagnostic amplifier would result in
The main benefit is a real time assessment as well as diagnostic options based on
as illustrated in the following case acquired on 04/21/10. The patient was diagnosed by our system with severe SIRS at a grade of 0.61 .
The patient was treated for SIRS and the blood tests were repeated during the following week. The full combined record of our system’s assessment of the patient, as derived from the further hematology tests, is illustrated below. The yellow line shows the diagnosis that corresponds to the first blood test (as also shown in the image above). The red line shows the next diagnosis that was performed a week later.
Chemistry of Herceptin [Trastuzumab] is explained with images in
http://www.chm.bris.ac.uk/motm/herceptin/index_files/Page450.htm
REFERENCES
The Cost Burden of Disease: U.S. and Michigan CHRT Brief. January 2010.
@www.chrt.org
The National Hospital Bill: The Most Expensive Conditions by Payer, 2006. HCUP Brief #59.
Rudolph RA, Bernstein LH, Babb J: Information-Induction for the diagnosis of myocardial infarction. Clin Chem 1988;34:2031-2038.
Bernstein LH, Qamar A, McPherson C, Zarich S, Rudolph R. Diagnosis of myocardial infarction: integration of serum markers and clinical descriptors using information theory. Yale J Biol Med 1999; 72: 5-13.
Kaplan L.A.; Chapman J.F.; Bock J.L.; Santa Maria E.; Clejan S.; Huddleston D.J.; Reed R.G.; Bernstein L.H.; Gillen-Goldstein J. Prediction of Respiratory Distress Syndrome using the Abbott FLM-II amniotic fluid assay. The National Academy of Clinical Biochemistry (NACB) Fetal Lung Maturity Assessment Project. Clin Chim Acta 2002; 326(8): 61-68.
Bernstein LH, Qamar A, McPherson C, Zarich S. Evaluating a new graphical ordinal logit method (GOLDminer) in the diagnosis of myocardial infarction utilizing clinical features and laboratory data. Yale J Biol Med 1999; 72:259-268.
Bernstein L, Bradley K, Zarich SA. GOLDmineR: Improving models for classifying patients with chest pain. Yale J Biol Med 2002; 75, pp. 183-198.
Ronald Raphael Coifman and Mladen Victor Wickerhauser. Adapted Waveform Analysis as a Tool for Modeling, Feature Extraction, and Denoising. Optical Engineering 1994; 33(7):2170–2174.
W Ruts, S De Deyne, E Ameel, W Vanpaemel,T Verbeemen, And G Storms. Dutch norm data for 13 semantic categories and 338 exemplars. Behavior Research Methods, Instruments, & Computers 2004; 36 (3): 506–515.
De Deyne, S Verheyen, E Ameel, W Vanpaemel, MJ Dry, WVoorspoels, and G Storms. Exemplar by feature applicability matrices and other Dutch normative data for semantic concepts. Behavior Research Methods 2008; 40 (4): 1030-1048
Landauer, T. K., Ross, B. H., & Didner, R. S. (1979). Processing visually presented single words: A reaction time analysis [Technical memorandum]. Murray Hill, NJ: Bell Laboratories. Lewandowsky, S. (1991).
Weed L. Automation of the problem oriented medical record. NCHSR Research Digest Series DHEW. 1977;(HRA)77-3177.
Naegele TA. Letter to the Editor. Amer J Crit Care 1993:2(5):433.
Retinal prosthetic strategy with the capacity to restore normal vision, Sheila Nirenberg and Chethan Pandarinath
http://www.pnas.org/content/109/37/15012
Larry H Bernstein, MD, FCAP
https://pharmaceuticalintelligence.com/2012/08/13/the-automated-second-opinion-generator/
Larry H Bernstein, MD, FCAP
Larry H Bernstein, MD, FCAP
Justin Pearlman, MD, PhD, FACC and Aviva Lev-Ari, PhD, RN
Larry H Bernstein, MD, FCAP
Larry H Bernstein, MD, FCAP
Aviva Lev-Ari, PhD, RN
Larry H Bernstein, MD, FCAP
Larry H Bernstein, MD, FCAP
Larry H Bernstein, MD, FCAP
Larry H Bernstein, MD, FCAP
https://pharmaceuticalintelligence.com/2012/12/17/big-data-in-genomic-medicine/
Larry H Bernstein, MD, FCAP
Isaac E. Mayzlin, Ph.D., David Mayzlin and Larry H. Bernstein, MD, FCAP
https://pharmaceuticalintelligence.com/2012/08/12/1815/
Reporter: Aviva Lev-Ari, PhD, RN
https://pharmaceuticalintelligence.com/2012/08/15/1946/
Aviva Lev-Ari, PhD, RN
Larry H Bernstein, MD, FACP and Aviva Lev-Ari, PhD, RN
https://pharmaceuticalintelligence.com/2013/05/05/bioengineering-of-vascular-and-tissue-models/
Aviva Lev-Ari, PhD, RN 2/28/2013
Aviva Lev-Ari, PhD, RN 1/28/2013
Aviva Lev-Ari, PhD, RN 1/10/2013
Aviva Lev-Ari, PhD, RN 1/3/2013
Aviva Lev-Ari, PhD, RN 12/29/2012
Aviva Lev-Ari, PhD, RN 8/27/2012
Additional Related articles
Posted in Cardiovascular Pharmacogenomics, Cell Biology, Signaling & Cell Circuits, Computational Biology/Systems and Bioinformatics, FDA Regulatory Affairs, Frontiers in Cardiology and Cardiovascular Disorders, Genomic Testing: Methodology for Diagnosis, Health Economics and Outcomes Research, HealthCare IT, International Global Work in Pharmaceutical, Medical and Population Genetics, Medical Devices R&D Investment, Medical Imaging Technology, Image Processing/Computing, MRI, CT, Nuclear Medicine, Ultra Sound, Molecular Genetics & Pharmaceutical, Origins of Cardiovascular Disease, Personalized and Precision Medicine & Genomic Research, Pharmaceutical Analytics, Pharmaceutical Industry Competitive Intelligence, Pharmaceutical R&D Investment, Pharmacogenomics, Population Health Management, Genetics & Pharmaceutical, Regulated Clinical Trials: Design, Methods, Components and IRB related issues, Statistical Methods for Research Evaluation, Technology Transfer: Biotech and Pharmaceutical | Tagged biomarker, breast cancer, Clinical decision support system, Compound annual growth rate, Diagnosis, HER2, Herceptin, Non-small-cell lung carcinoma, NSCLC, Personalized medicine, Trastuzumab | 6 Comments »





