
Original description – :Cartoon representation of ubiquitin protein, highlighting the secondary structure. α-helices are coloured in blue and the β-sheet in green. The normal attachment point for a further ubiquitin molecule in polyubiquitin chain formation, lysine 48, is shown in pink. :Image was created using PyMOL (Photo credit: Wikipedia)
Ubiquinin-Proteosome pathway, autophagy, the mitochondrion, proteolysis and cell apoptosis
Larry H Bernstein, MD, FACP, Curator, Reporter, AEW
The work reviewed follows a seminal contribution by two Israeli and an American molecular biologists who shared the Nobel Prize in Chemistry in 2004.
The Royal Swedish Academy of Sciences awarded the Nobel Prize in Chemistry for 2004 “for the discovery of ubiquitin-mediated protein degradation” jointly to Aaron Ciechanover Technion – Israel Institute of Technology, Haifa, Israel, Avram Hershko Technion – Israel Institute of Technology, Haifa, Israel and Irwin Rose – University of California, Irvine, USA.
Aaron Ciechanover, born 1947 (57 years) in Haifa, Israel (Israeli citizen) received a Doctor’s degree in medicine in 1975 at Hebrew University of Jerusalem, and in biology in 1982 at the Technion (Israel Institute of Technology), Haifa. He is a Distinguished Professor at the Center for Cancer and Vascular Biology, and the Rappaport Faculty of Medicine and Research Institute at the Technion, Haifa,
Israel.
Avram Hershko, born 1937 (67 years) in Karcag, Hungary (Israeli citizen) earned the Doctor’s degree in medicine in 1969 at the Hadassah and the Hebrew University Medical School, Jerusalem. He is a Distinguished Professor at the Rappaport Family Institute for Research in Medical Sciences at the Technion (Israel Institute of Technology), Haifa, Israel.
Irwin Rose, born 1926 (78 years) in New York, USA (American citizen) achieved a Doctor’s degree in 1952 at the University of Chicago, USA. Specialist at the Department of Physiology and Biophysics, College of Medicine, University of California, Irvine, USA.
Proteins labelled for destruction
Proteins build up all living things: plants, animals and therefore us humans. In the past few decades biochemistry has come a long way towards explaining how the cell produces all its various proteins. But as to the breaking down of proteins, not so many researchers were interested. Aaron Ciechanover, Avram Hershko and Irwin Rose went against the stream and at the beginning of the 1980s discovered one of the cell’s most important cyclical processes, regulated protein degradation. For this, they are being rewarded
with the 2004 Nobel Prize in Chemistry.
The label consists of a molecule called ubiquitin. This fastens to the protein to be destroyed, accompanies it to the proteasome where it is recognised as the key in a lock, and signals that a protein is on the way for disassembly. Shortly before the protein is squeezed into the proteasome, its ubiquitin label is disconnected for re-use.
Aaron Ciechanover, Avram Hershko and Irwin Rose have brought us to realise that the cell functions as a highly-efficient checking station where proteins are built up and broken down at a furious rate. The degradation is not indiscriminate but takes place through a process that is controlled in detail so that the proteins to be broken down at any given moment are given a molecular label, a ‘kiss of death’, to be dramatic. The labelled proteins are then fed into the cells’ “waste disposers”, the so called proteasomes, where they are chopped into small pieces and destroyed.
Animation (Plug in requirement: Flash Player 6)
Thanks to the work of the three Laureates it is now possible to understand at molecular level how the cell controls a number of central processes by breaking down certain proteins and not others. Examples of processes governed by ubiquitin-mediated protein degradation are cell division, DNA repair, quality control of newly-produced proteins, and important parts of the immune defence. When the degradation does not work correctly, we fall ill. Cervical cancer and cystic fibrosis are two examples. Knowledge of
ubiquitin-mediated protein degradation offers an opportunity to develop drugs against these diseases and others.
Aaron Ciechanover and Ronen Ben-Saadon. N-terminal ubiquitination: more protein substrates join in. TRENDS in Cell Biology 2004; 14 (3):103-106.
The ubiquitin–proteasome system (UPS) is involved in selective targeting of innumerable cellular proteins through a complex pathway that plays important roles in a broad array of processes. An important step in the proteolytic cascade is specific recognition of the substrate by one of many ubiquitin ligases, E3s, which is followed by generation of the polyubiquitin degradation signal. For most substrates, it is believed that the first ubiquitin moiety is conjugated, through its C-terminal Gly76 residue, to an 1-NH2 group of an internal Lys residue. Recent findings indicate that, for several proteins, the first ubiquitin moiety is fused linearly to the a-NH2 group of the N-terminal residue.
The ubiquitin–proteasome system (UPS). Ubiquitin is first activated to a high-energy intermediate by E1. It is then transferred to a member of the E2 family of enzymes. From E2 it can be transferred directly to the substrate (S, red) that is bound specifically to a member of the ubiquitin ligase family of proteins, E3
- (a). This occurs when the E3 belongs to the RING finger family of ligases. In the case of a HECT-domain-containing ligase
- (b), the activated ubiquitin is transferred first to the E3 before it is conjugated to the E3-bound substrate . Additional ubiquitin moieties are added successively to the previously conjugated moiety to generate a polyubiquitin chain.
- The polyubiquitinated substrate binds to the 26S proteasome complex (comprising 19S and 20S sub-complexes): the substrate is degraded to short peptides, and free and reusable ubiquitin is released through the activity of de-ubiquitinating enzymes (DUBs).
Ubiquitination on an internal lysine and on the N-terminal residue of the target substrate.
- (a) The first ubiquitin moiety is conjugated, through its C-terminal Gly76 residue, to the 1-NH2 group of an internal lysine residue of the target substrate (Kn).
- (b) The first ubiquitin moiety is conjugated to a free a-NH2 group of the N-terminal residue, X.
- In both cases, successive addition of activated ubiquitin moieties to internal Lys48 on the previously conjugated ubiquitin moiety leads to the synthesis of a polyubiquitin chain that serves as the degradation signal for the 26S proteasome
A UPS Autophagy Review
Summary: This discussion is another in a series discussing mitochondrial metabolism, energetics and regulatory function, and dysfunction, and the process leading to apoptosis and a larger effect on disease, with a specific targeting of neurodegeneration. Why neurological and muscle damage are more sensitive than other organs is not explained easily, but recall in the article on mitochondrial oxidation-reduction reactions and repair that there are organ specific differences in the rates of organelle mutation errors and in the rates of repair. In addition, consider the effect of iron-binding in the function of the cell, and Ca2+ binding in the creation of the mechanic work or signal transmission carried out by the neuromuscular system. We target the previously mentioned role of ubiquitin-proteosome, and interaction with autophagy, mitophagy, and disease.
Keywords: autophagy, ubiquitin-proteosome, UPS, protein degradation, defective organelle removal, selective degradation, E3, neurodegenerative disease, mitochondria, mitophagy, proteolysis, ribosomes, apoptosis, Ca++, rapamycin, TORC1, atg1p kinase, ubiqitization, trafficking pathways, unfolded protein response (UPS), p52/sequestrome, IC3, nitrogen starvation, acetaldehyde dehydrogenase (Ald6p), Ut1hp, toxisomes, Pex3/14 proteins, Bax, E3 Ligase, TRAP1, TNF-a, NFkB.
Ubiquitin-Proteosome Pathway
Three recent papers, describing three apparently independent biological processes, highlight the role of the ubiquitin-proteasome system as a major, however selective, proteolytic and regulatory pathway. Using specific inhibitors to the proteasome, Rock et al. (1994) demonstrate a role for this protease in the degradation of the major bulk of cellular proteins. They also showed that antigen processing requires the ubiquitin-activating enzyme, El. This indicates that antigen processing is both ubiquitin dependent and proteasome dependent. Furthermore, inhibitors to the proteasome prevent tumor necrosis factor a (TNFa)-induced activation of mature NFKB and its entry into the nucleus. The two studies clearly demonstrate that the ubiquitin-proteasome system is involved not only in complete destruction of its protein substrates, but also in limited proteolysis and posttranslational processing in which biologically active peptides or fragments are generated. In addition, the unstable c-Jut but not the stable v-Jun, is multiubiquitinated and degraded. The escape of the oncogenic v-Jun from ubiquitin-dependent degradation suggests a novel route to malignant transformation. Presented here is a review of the components, mechanisms of action, and cellular physiology of the ubiquitin-proteasome pathway.
Experimental evidence implicates the ubiquitin system in the degradation of
- mitotic cyclins,
- oncoproteins,
- the tumor suppressor protein p53,
- several cell surface receptors,
- transcriptional regulators, and
- mutated and damaged proteins.
Some of the proteolytic processes occur throughout the cell cycle, whereas others are tightly programmed and occur following cell cycle-dependent posttranslational modifications of the components involved. Signaling and degradation of other proteins (cell surface receptors, for example) may occur only following structural changes or modification(s) in the target molecule that results from ligand binding. Cell cycle-and modification-dependent degradation, as well the ability of the system to destroy completely or only partially its protein substrates, reflects the complexity involved in regulated intracellular protein degradation.
Enzymes of the System
The reaction occurs in two distinct steps:
- signaling of the protein by covalent attachment of multiple ubiquitin molecules and
- degradation of the targeted protein with the release of free and reutilizable ubiquitin.
Conjugation of ubiquitin to proteins destined for degradation proceeds, in general, in a three-step mechanism.
- Initially, the C-terminal Gly of ubiquitin is activated by ATP to a high energy thiol ester intermediate in a reaction catalyzed by the ubiquitin-activating enzyme, El.
- Following activation, E2 (ubiquitin carrier protein or ubiquitin-conjugating enzyme [USC]) transfers ubiquitin from El to the substrate that is bound to a ubiquitin-protein ligase, E3.
- Here an isopeptide bond is formed between the activated C-terminal Gly of ubiquitin and an c-NH2 group of a Lys residue of the substrate.
As E3 enzymes specifically synthesized by processive transfer of ubiquitin moieties to Lys-48 of the previous (and already conjugated) ubiquitin molecule. In many cases, E2 transfers activated ubiquitin directly to the protein substrate. Thus, E2 enzymes also play an important role in substrate recognition, although, in most cases, this modification is of the monoubiquitin type.
The Ubiquitin-Mediated Proteolytic Pathway
(1) Activation of ubiquitin by El and E2.
(2) Binding of the protein substrate to E3.
(3) EP dependent but EM independent monoubiquitination.
(4) EP-dependent but EM independent polyubiquitination?
(5) Ed-dependent polyubiquitination.
(6) Degradation of ubiquitin-protein conjugate by the 26s protease.
(7) “Correction” function of C-terminal hydrolase(s).
(6) Release of ubiquitin from terminal proteolytic products by &terminal hydrolase(s).
It is essential for the system that ubiquitin recycles. This function is carried out by ubiquitin C-terminal hydrolases (isopeptidases). In protein degradation, hydrolase(s) is required to release ubiquitin from isopeptide linkage with Lys residues of the protein substrate at the final stage of the proteolytic process. A ubiquitin C-terminal hydrolytic activity is also required to disassemble polyubiquitin chains linked to the protein substrate, following or during the degradative process. A “proofreading” function has been proposed for hydrolases to release free protein from “incorrectly” ubiquitinated proteins. Another possibility is that ubiquitin C-terminal hydrolases are required for trimming polyubitin chains.
Hydrolases are probably required for the processing of biosynthetic precursors of ubiquitin, since most ubiquitin genes are arranged either in linear polyubiquitin arrays or are fused to ribosomal proteins. Yet another hydrolase may be required for the removal of extra amino acid residues that are encoded by certain genes at the C-termini of some polyubiquitin molecules. Ubiquitin C-terminal hydrolases may have other functions as well. High energy El-ubiquitin and E2-ubiquitin thiol esters may react with intracellular nucleophiles (such as glutathione or polyamines). Such reactions may lead to rapid depletion of free ubiquitin unless such side products are rapidly cleaved.
Recognition of Substrates
Short-lived proteins contain a region enriched with Pro, Glu, Ser, and Thr (PEST region). However, it has not been shown that this region indeed serves as a consensus proteolysis targeting signal. An interesting problem involves the evolution of the N-end rule pathway and its physiological roles. Proteins that are derived from processing of polyproteins (Sindbis virus RNA polymerase, for example) may contain destabilizing N-termini and thus are proteolyzed via the N-end rule pathway.
Using a “synthetic lethal” screen, Ota and Varshavsky attempted to isolate a mutant that requires the N-end rule pathway for viability. They characterized an extragenic suppressor of the mutation and found that it encodes a protein with a strong correlation to protein phosphotyrosine phosphatase. The target protein or the connection between dephosphorylation of phosphotyrosine and the N-end rule pathway is still obscure. In an additional study, these researchers have shown that a missense mutation in SLNI, a member of a two-component signal transduction system in yeast, is lethal in the absence, but not in the presence, of the N-end rule pathway. Further studies are required to isolate the target protein and identify the signal transduction pathway.
Two recent studies have shed light on the role of the ubiquitin system and the proteasome in the process. Michalek et al. (1993) have shown that a mutant cell that harbors a thermolabile El cannot present peptides derived from ovalbumin following inactivation of the enzyme. In contrast, presentation of a minigene-expressed antigene peptide or presentation of exogenous similar peptide was not perturbed at the nonpermissive temperature. The important conclusion of the researchers is that the processing of the protein to peptides requires the complete ubiquitin pathway. In a complementary study, Rock et al. (1994) have shown that inhibitors that block the chymotryptic activity of the proteasome also block antigen presentation, most probably by inhibiting proteolysis of the antigen (ovalbumin). Thus, it appears that processing of MHC restricted class I antigens requires both ubiquitination and subsequent degradation by the proteasome. It is likely that the proteasome catalyzes processing of these antigens as part of the 26s protease complex.
Ciechanover A. The Ubiquitin-Proteasome Proteolytic Pathway. Cell 1994; 79:13-21.
Regulation of autophagy
The protein content of the cell is determined by the balance between protein synthesis and protein degradation. At constant intracellular protein concentration, i.e. at steady state, rates of protein synthesis and degradation are equal. Although turnover of protein results in energy dissipation, regulation at the level of protein degradation effectively controls protein levels.
Intracellular proteins to be degraded in the lysosomes can get access to these organelles by the following processes:
- macroautophagy,
- microautophagy,
- crinophagy and selective,
- chaperonin mediated, direct uptake of proteins.
Overview of the involvement of signal transduction in the regulation of macroautophagic proteolysis by amino acids and cell swelling.
- Amino acids (AA) stimulate a protein kinase cascade via a plasma membrane receptor.
- Receptor activation results in activation of PtdIns 3-kinase (PI3K), possibly via a heterotrimeric Gái3 protein.
- followed by activation of PKC-æ, PKB/Akt, p70S6 kinase (p70S6k) and finally phosphorylation of ribosomal protein S6 (S6P).
- The GDP-bound form of Gái3 is required for autophagic sequestration, whereas the GTP-bound form is inhibitory.
- The constitutively formed phosphatidylinositol 3-phosphate (PI3P) is also required for autophagic sequestration. Therefore,
inhibition of PtdIns 3-kinase activity by
- wortmannin (W),
- LY294002 (LY) or
- 3-methyladenine (3MA) prevents autophagic sequestration.
Activation of PKC-æ and PKB/Akt is mediated by the 3,4- and 3,4,5-phosphate forms of phosphatidylinositol (PI3,4P2 and PI3,4,5P3) that are produced upon activation of PtdIns 3-kinase.
As a result of this, the first step of the macroautophagic pathway is
- inhibited by components of the cascade that are downstream of PtdIns 3-kinase.
- inhibition of this downstream cascade by rapamycin (RAPA) accelerates autophagic sequestration.
- cell swelling potentiates the effect of amino acids via a change in the receptor owing to membrane stretch.
Furthermore, the site of action of the different effectors of the cytoskeleton (okadaic acid, cytochalasin, nocodazole, vinblastin and colchicine) are indicated.
- AVi,
- initial autophagic vacuole;
- AVd,
- mature degradative autophagic vacuole,
- ER, endoplasmic reticulum.
The rate of proteolysis , an important determinant of the intracellular protein content, and part of its degradation occurs in the lysosomes and is mediated by macroautophagy. In liver, macroautophagy is very active and almost completely accounts for starvation-induced proteolysis. Factors inhibiting this process include
- amino acids,
- cell swelling and
- insulin.
In the mechanisms controlling macroautophagy, protein phosphorylation plays an important role.
- Activation of a signal transduction pathway, ultimately
- leading to phosphorylation of ribosomal protein S6,
- accompanies Inhibition of macroautophagy.
Components of this pathway may include
- a heterotrimeric Gi3-protein,
- phosphatidylinositol 3-kinase and
- p70S6 kinase.
Selectivity of Autophagy
It has been assumed for a long time that macroautophagy is a non-selective process, in which macromolecules are randomly degraded in the same ratio as they occur in the cytoplasm . However, recent observations strongly suggest that this may not always be the case, and that macroautophagy can be selective. Lysosomal protein degradation can selectively occur via ubiquitin-dependent and -independent pathways. In the perfused liver, although autophagic breakdown of protein and RNA (mainly ribosomal RNA) is sensitive to inhibition by amino acids and insulin, glucagon accelerates proteolysis but has no effect on RNA degradation.
Another example of selective autophagy is the degradation of superfluous peroxisomes in hepatocytes from clofibrate-treated rats. When hepatocytes from these rats, in which the number of peroxisomes is greatly increased, are incubated in the absence of amino acids to ensure maximal flux through the macroautophagic pathway, peroxisomes are degraded at a relative rate that exceeds that of any other component in the liver cell. The accelerated degradation of peroxisomes was sensitive to inhibition by 3-methyladenine, a specific autophagic sequestration inhibitor. Interestingly, the accelerated removal of peroxisomes was prevented by long-chain but not short-chain fatty acids. Since long-chain fatty acids are substrates for peroxisomal â-oxidation, this indicates that these organelles are removed by autophagy when they are functionally redundant. Our hypothesis is that acylation (palmitoylation?) of a peroxisomal membrane protein protects the peroxisome against autophagic sequestration.
Under normal conditions macroautophagy may be largely unselective and serves, for example, to produce amino acids for gluconeogenesis and the synthesis of essential proteins in starvation. When cell structures are functionally redundant or when they become damaged, the autophagic system is able to recognize this and is able to degrade the structure concerned. As yet, nothing is known about the recognition signals. A possibility is that ubiquitination of membrane proteins is required to mark the structure to be degraded for autophagic sequestration.
Ubiquitin may be involved in macroautophagy
Ubiquitin not only contributes to extralysosomal proteolysis but is also involved in autophagic protein degradation. Thus, in fibroblasts ubiquitin–protein conjugates can be found in the lysosomes, as shown by immunohistochemistry and immunogold electron microscopy. Free ubiquitin can also be found inside lysosomes. Accumulations of ubiquitin–protein conjugates in filamentous, presumably lysosomal, structures are also found in a large number of neurodegenerative diseases. Mallory bodies in the liver of alcoholics also contain ubiquitin–protein conjugates.
This presence of ubiquitin–protein conjugates in filamentous inclusions in neurons and other cells can be caused by a defect in the extralysosomal ubiquitin-dependent proteolytic pathway. However, it is also possible that these filamentous inclusions represent an attempt of the cell to get rid of unwanted material (proteins, organelles) via autophagy. Direct evidence that ubiquitin may be involved in the control of macroautophagy came from experiments with CHO cells with a temperature-sensitive mutation in the ubiquitin-activating enzyme E1. Wild-type cells increased their rate of proteolysis in response to stress (amino acid depletion, increased temperature). This was prevented by the acidotropic agent ammonia or by the autophagic sequestration inhibitor 3-methyladenine, indicating that the accelerated proteolysis occurred by autophagy. In the mutant cells, there was no such increase in proteolysis in response to stress at the restrictive temperature.
Autophagy and carcinogenesis
In cancer development, cell growth is mainly induced by inhibition of protein degradation, since differences in the rate of protein synthesis between tumorigenic cells and their normal counterparts are rather small. A striking example of how reduced autophagic proteolysis can contribute to cell growth can be found in the development of liver carcinogenesis. This decrease in autophagic flux results from a decrease in the rate of autophagic sequestration and is already detectable in the early preneoplastic stage. Autophagic flux is then hardly inhibitable by amino acids nor is it inducible by catabolic stimuli
and declines in the more advanced stage of cancer development to a rate of less than 20% of that seen in normal hepatocytes. The fact that the addition of 3-methyladenine to hepatocytes from normal rats increased hepatocyte viability to the same level as observed for the tumour cells strongly suggests that the fall in autophagic proteolysis contributes to the rapid growth rate of these cells and gives them a selective advantage over the normal hepatocytes.
Underlying control mechanisms for autophagy are gradually being unravelled. It is perhaps not surprising that protein phosphorylation and signal transduction are key elements in these mechanisms. The discovery of an amino acid receptor in the plasma membrane of the hepatocyte with a signal transduction pathway coupled to it may have important repercussions, not only for the control of macroautophagy but also for the control of other pathways.
It remains to be seen whether the details of the mechanisms controlling the process in yeast are similar to those in mammalian cells. For example, it is not known whether amino acids are able to control the process as they do in mammalian cells.
Blommaart EFC, Luiken JJFP, Meijer AJ. Autophagic proteolysis: control and specificity. Histochemical Journal (1997); 29:365–385.
A Novel Type of Selective Autophagy
Eukaryotic cells use autophagy and the ubiquitin–proteasome system (UPS) as their major protein degradation pathways. Whereas the UPS is required for the rapid degradation of proteins when fast adaptation is needed, autophagy pathways selectively remove protein aggregates and damaged or excess organelles. However, little is known about the targets and mechanisms that provide specificity to this process. Here we show that mature ribosomes are rapidly degraded by autophagy upon nutrient starvation in Saccharomyces cerevisiae. Surprisingly, this degradation not only occurs by a nonselective mechanism, but also involves a novel type of selective autophagy, which we term ‘ribophagy’. A genetic screen revealed that selective degradation of ribosomes requires catalytic activity of the Ubp3p/Bre5p ubiquitin protease. Although Ubp3p and Bre5p cells strongly accumulate 60S ribosomal particles upon starvation, they are proficient in starvation sensing and in general trafficking and autophagy pathways. Moreover, ubiquitination of several ribosomal subunits and/or ribosome associated proteins was specifically enriched in Ubp3p cells, suggesting that the regulation of ribophagy by ubiquitination may be direct. Interestingly, Ubp3p cells are sensitive to rapamycin and nutrient starvation, implying that selective degradation of ribosomes is functionally important in vivo. Taken together, our results suggest a link between ubiquitination and the regulated degradation of mature ribosomes by autophagy.
Kraft C, Deplazes A, Sohrmann M,Peter M. Mature ribosomes are selectively degraded upon starvation by an autophagy pathway requiring the Ubp3p/Bre5p ubiquitin protease. Nature Cell Biology 2008; 10(5): 603-609. DOI: 10.1038/ncb1723. www.nature.com/naturecellbiology
Mitochondrial Failure and Protein Degradation
Progressive mitochondrial failure is tightly associated with the the development of most age-related human diseases including neurodegenerative diseases, cancer, and type 2 diabetes.
This tight connection results from the double-edged sword of mitochondrial respiration, which is responsible for generating both ATP and ROS, as well as from risks that are inherent to mitochondrial biogenesis. To prevent and treat these diseases, a precise understanding of the mechanisms that maintain functional mitochondria is necessary. Mitochondrial protein quality control is one of the mechanisms that protect mitochondrial integrity, and increasing evidence implicates the cytosolic ubiquitin/proteasome system (UPS) as part of this surveillance network. In this review, we will discuss our current understanding of UPS-dependent mitochondrial protein degradation, its roles in diseases progression, and insights into future studies.
While mitochondria have their own genome, about 99% of the roughly 1000 mitochondrial proteins are encoded in the nuclear genome. Most mitochondrial proteins are therefore
- synthesized in the cytoplasm,
- unfolded,
- transported across one or both mitochondrial membranes,
- then refolded and/or assembled into complexes (Tatsuta, 2009).
Failure of this complex series of events generates unfolded or misfolded proteins within mitochondria, often disrupting critical functions.
Mitochondrial oxidative phosphorylation generates usable cellular energy in the form of ATP, but also produces reactive oxygen species (ROS) . ROS tend to react quickly, so their predominant sites of damage are mitochondrial macromolecules that are localized nearby the source of ROS production.
Exposure to oxidative stress facilitates misfolding and aggregation of these mitochondrial proteins, leading to disassembly of protein complexes and eventual loss of mitochondrial integrity.
The clearance of misfolded and aggregated proteins is constantly needed to maintain functional mitochondria.
There are several systems promoting this turnover.
- Mitophagy, a selective mitochondrial autophagy, mediates a bulk removal of damaged mitochondria.
- mitochondria intrinsically contain proteases in each of their compartments and these proteases recognize misfolded mitochondrial proteins and mediate their degradation.
Accumulating evidence shows that the ubiquitin proteasome system (UPS) plays an important role in mitochondrial protein degradation. At various cellular sites, the UPS is involved in protein degradation. With the help of ubiquitin E1–E2–E3 enzyme cascades, target proteins destined for destruction are marked by conjugation of K48-linked poly-ubiquitin chain. This poly-ubiquitinated protein is then targeted to the proteasome for degradation.
Cells treated with proteasome inhibitors exhibit elevated levels of ubiquitinated mitochondrial proteins, suggesting the potentially important roles of the proteasome on mitochondrial protein degradation. Studies have also identified mitochondrial substrates of the UPS.
- Fzo1, an outer mitochondrial membrane (OMM) protein involved in mitochondrial fusion, is partially dependent on the proteasome for its degradation in yeast.
- The F box protein Mdm30 mediates ubiquitination of Fzo1 by Skp1-Cullin-F-boxMdm30 ligase, which leads to proteasomal degradation.
The UPS has also been implicated in mitochondrial protein degradation in higher organisms. In mammals,
- the OMM proteins mitofusin 1 and 2 (Mfn1/2; the mammalian orthologs of Fzo1) and Mcl1 are polyubiquitinated and degraded by the proteasome.
- VDAC1, Tom20 and Tom70 were also suggested as targets of proteasomal degradation as they are stabilized by proteasome inhibition.
- inactivation of the proteasome also induces accumulation of intermembrane space (IMS) proteins and, consistent with this, the proteasome plays a role in degradation of the IMS protein, Endonuclease G.
Turnover of some inner mitochondrial membrane (IMM) proteins is also dependent upon the proteasome. Uncoupling proteins (UCPs) 2 and 3 exhibit an unusually short half-life compared with other IMM proteins, and Brand and colleagues showed that inactivation of the proteasome prevents their turnover in vivo and in a reconstituted in vitro system. Finally, mitochondrial matrix proteins can also be degraded by the proteasome.
Cdc48/p97 is involved in many cellular processes through its role in protein degradation and is targeted to different subcellular sites by adaptor proteins. For example, Cdc48/p97 is recruited to the endoplasmic reticulum with the help of two adaptor proteins, Npl4 and Ufd1. This implies the existence of specific adaptors that recruit Cdc48/p97 to mitochondria. Consistent with this notion, the authors recently identified a mitochondrial adaptor protein for Cdc48, which we named Vms1 (VCP/Cdc48-associated mitochondrial stress responsive 1). Vms1 interacts with Cdc48/p97 and Npl4, but not with Ufd1, which indicates that the Cdc48/p97–Npl4–Ufd1 complex functions in ER protein degradation while the Vms1–Cdc48/p97–Npl4 complex acts in mitochondria. In agreement with this notion, overexpression of Cdc48 or Npl4 rescues the Vms1 mutant phenotype while Ufd1 has no effect.
Normally, Vms1 is cytoplasmic. Upon mitochondrial stress, however, Vms1 recruits Cdc48 and Npl4 to mitochondria. In agreement with the role of Cdc48/p97 in OMM protein degradation, loss of the Vms1 system results in accumulation of ubiquitin-conjugated proteins in purified mitochondria as well as stabilization of Fzo1 under mitochondrial stress conditions. Accumulation of damaged and misfolded mitochondrial proteins disturbs the normal physiology of the mitochondria, leading to mitochondrial dysfunction. As expected, the Vms1 mutants progressively lose mitochondrial respiratory activity, eventually leading to cell death. The VMS1 gene is broadly conserved in eukaryotes, implying an important functional role in a wide range of organisms. The C. elegans Vms1 homolog exhibits a similar pattern of mitochondrial stress responsive translocation and is required for normal lifespan. Additionally, mammalian Vms1 also forms a stable complex with p97. Combining these observations, the authors conclude that Vms1 is a conserved component of the UPS-dependent mitochondrial protein quality control system.
The UPS regulates mitochondrial dynamics and initiation of mitophagy
The UPS regulates mitochondrial dynamics. Major proteins involved in mitochondrial fission or fusion (e.g. Mfn1/2, Drp1 and Fis1) are degraded by the UPS. MITOL, a mitochondrial E3 ubiquitin ligase, is required for Drp1-dependent mitochondrial fission as depletion or inactivation of MITOL blocks mitochondrial fragmentation. Moreover, knockdown of USP30, an OMM-localized deubiquitinating enzyme, induces an elongated mitochondrial morphology, suggesting a defect in fission. Through this regulatory process, the UPS controls mitochondrial dynamics. Parkin, an E3 ligase involved in mitophagy, utilizes the UPS to enhance mitochondrial fission through degradation of components of the fusion machinery. By facilitating fragmentation of damaged mitochondria, which is essential for initiation of mitophagy, Parkin stimulates mitophagy. The underlying mechanisms linking the UPS to the regulation of mitochondrial dynamics remain unclear.
Accumulation of aberrant proteins and human diseases
In neurodegenerative diseases wherein aberrant pathological proteins accumulate throughout the cell, including sites in mitochondria. Amyloid precursor protein (APP), a protein associated with Alzheimer’s disease, accumulates within mitochondria and is implicated in blockade of mitochondrial protein import. A, a neurotoxic APP cleavage product, can also facilitate the formation of the mitochondrial permeability transition pore (mPTP) by binding to mPTP components VDAC1, CypD and ANT, which provokes cell death.
-Synuclein, a protein associated with the development of Parkinson’s disease, is targeted to the IMM where it binds to the mitochondrial respiratory complex I and impairs its function. -Synuclein interferes with mitochondrial dynamics as its unique interaction with the mitochondrial membrane disturbs the fusion process. Finally, in Huntington’s disease, increased association of the mutant huntingtin protein with mitochondria can impair mitochondrial trafficking. Moreover, accumulation of mutant huntingtin protein disrupts cristae structure and it facilitates mitochondrial fragmentation by activation of Drp1. These examples demonstrate the crucial importance of prompt removal of dysfunctional and/or aberrant proteins in maintaining functional mitochondria.
UPS-mediated mitochondrial protein degradation.
Misfolded and/or damaged mitochondrial proteins destined for proteasomal degradation in the cytosol are recruited to the outer mitochondrial membrane (OMM) from each mitochondrial compartment by unknown mechanisms. Upon reaching the OMM, these proteins are presented to the proteasome through a series of events. They are K48 polyubiquitinated by the cytoplasmic (e.g. Parkin) or mitochondrial ubiquitin E3 ligases. For proteasomal degradation, polyubiquitinated mitochondrial substrate proteins need to be retrotranslocated to the cytoplasm, probably, either by the proteasome per se or by the help of UPS components such as Vms1, Cdc48/p97 and Npl4. Following dislocation to the cytoplasm, these substrate proteins are degraded by the proteasome.
Treatment of diseases that arise from defects in protein quality control will depend on greater depth in our understanding of this process, which could contribute to the development of novel therapeutic approaches. For instance, both mutant SOD1, a misfolded mitochondrial protein associated with the onset of amyotrophic lateral sclerosis, and polyglutamine expanded ataxin-3, a pathogenic protein causing Machado-Joseph disease, are ubiquitinated by MITOL and then degraded by the proteasome. Facilitating the proteasomal degradation of these aberrant proteins might therefore efficiently control diseases progression and, eventually, cure the diseases. Answering these questions would partially unveil the mysterious physiology of mitochondria, which, in turn, would facilitate the development of therapeutics to prevent and cure devastating human diseases.
Heo JM, Rutter J. Ubiquitin-dependent mitochondrial protein degradation. The International Journal of Biochemistry & Cell Biology 2011; 43:1422– 1426. http://www.elsevier.com/locate/biocel
UPS Inhibitors and Apoptotic Machinery
Over the past decade, the promising results of UPSIs (UPS inhibitors) in eliciting apoptosis in various cancer cells, and the approval of the first UPSI (Bortezomib/Velcade/PS-341) for the treatment of multiple myeloma have raised interest in assessing the death program activated upon proteasomal blockage. Several reports indicate that UPSIs stimulate apoptosis in malignant cells by operating at multiple levels, possibly by inducing different types of cellular stress. Normally cellular stress signals converge on the core elements of the apoptotic machinery to trigger the cellular demise. In addition to eliciting multiple stresses, UPSIs can directly operate on the core elements of the apoptotic machinery to control their abundance. Alterations in the relative levels of anti and pro-apoptotic factors can render cancer cells more prone to die in response to other anti-cancer treatments. Aim of the present review is to discuss those core elements of the apoptotic machinery that are under the control of the UPS.
The UPS (Ubquitin-Proteasome System)
To fulfill the protein-degradation process two branches, operating at different levels, principally comprise the UPS.
- The first branch is formed by the enzymatic activities responsible for delivering the substrate to the degradative machinery: the targeting branch.
- The second branch is represented by the proteolytic machinery, which ultimately fragments the protein substrate into small oligopeptides.
Oligopeptides are further digested to single amino acids by cytosolic proteases.
It is important to remember that conjugation of ubiquitin to a specific protein is not sufficient to determine its degradation. In fact, mono-ubiquitination or poly-monoubiquitination and in certain cases also poly-ubiquitination of proteins are post-translational modifications related to various cellular functions including DNA repair or membrane trafficking . To deliver polypeptides for proteasomal degradation poly-ubiquitin chains of more than 4 ubiquitins must be assembled through lysine-48 linkages.
There are 3 catalytic sites for each polyubiquitin chain. These sites show specific requirements in terms of substrate specificities and catalytic activities, and they are identified as
- trypsin-like, which prefer to cleave after hydrophobic bonds, chymotrypsin-like, which cleave at basic residues and
- postglutamyl peptide hydrolase-like or
- caspase-like activities, which cut after acidic amino acid.
Each proteasome active site uses the side chain hydroxyl group of an NH2-terminal threonine as the catalytic nucleophile, a mechanism that distinguishes the proteasome from other cellular proteases. The presence of substrate proteolysis small size peptides ranging from 3 to 22 residues are generated. Alternative catalytic sites guarantees the efficient processing of several different substrates.
UPS Inhibitors
By UPS inhibitors (UPSI) we mean small molecules that share the ability to target and inhibit specific activities of the UPS, causing the accumulation of poly-ubiquitinated proteosomal substrates. UPSIs are heterogeneous compounds and among them bortezomib is the only one used in clinical practice.
PR-171, a modified peptide related to the natural product epoxomicin, is composed of two key elements:
- a peptide portion that selectively binds with high affinity in the substrate binding pocket(s) of the proteasome and
- an epoxyketone pharmacophore that stereospecifically interacts with the catalytic threonine residue and irreversibly inhibits enzyme activity.
In comparison to bortezomib, PR-171 exhibits equal potency, but greater selectivity, for the chymotrypsin-like activity of the proteasome. In cell culture PR-171 is more cytotoxic than bortezomib. In mice PR-171 is well tolerated and shows stronger anti-tumor activity when compared with bortezomib . Clinical studies are in progress to test the safety of PR-171 at different dose levels on some hematological cancers.
Cell Death by UPSI
In vitro experiments have unambiguously established that incubation of neoplastic cells with UPSIs including bortezomib triggers their death. Apoptosis or type I cell death relies on the timed activation of caspases, a group of cysteine proteases, which cleave selected cellular substrates after aspartic residues. Two main apoptotic pathways keep in check caspase activation.
The turnover of a large number of cellular proteins is under the control of the UPS. Thus in principle any proteosomal substrate could contribute directly or indirectly to the cell death phenotype. This is perfectly exemplified by two master regulators of cell life and death, p53 and NFkB. UPSIs cause
- NF-kB inhibition through reduced IkB degradation and,
- in opposition; they promote stabilization and accumulation of p53.
c-FLIP is the most important element of the extrinsic pathway under the direct control of the UPS. Two different FLIP isoforms exist:
- c-FLIPL (Long) and
- c-FLIPS (Short).
c-FLIPL is highly homologus to caspase-8 and contains two tandem repeat Death Effector Domains (DED) and a catalytically inactive caspase-like domain. Both FLIPs can be degraded by the UPS; however they display distinct half-lives and the unique C terminus of c-FLIPS possesses a destabilizing function. The regulation of c-FLIP levels in response to UPSIs is rather controversial. Some reports indicate that UPSIs can reduce c-FLIP levels and in this manner synergize with TRAIL to promote apoptosis.
UPSIs activate multiple cellular responses and different stress signals that ultimately cause cell death. For this reason they represent broad inducers of apoptosis. In addition, since many of the available UPSIs alter the proteolytic activity of the proteasome, they represent non-specific modulators of the expression/activity of various components of the apoptotic machinery. Paradoxically they can simultaneously favor the accumulation of pro- and anti-apoptotic factors.
Brancolini C. Inhibitors of the Ubiquitin-Proteasome System and the Cell Death Machinery: How Many Pathways are Activated? Current Molecular Pharmacology, 2008; 1:24-37.
Mitochondrial Quality Control
The PINK1–Parkin pathway plays a critical role in mitochondrial quality control by selectively targeting damaged mitochondria for autophagy. The AAA-type ATPase p97 acts downstream of PINK1 and Parkin to segregate fusion-incompetent mitochondria for turnover. [Tanaka et al. (2010. J. Cell Biol. doi: 10.1083/jcb.201007013)]. p97 acts by targeting the mitochondrial fusion-promoting factor mitofusin for degradation through an endoplasmic reticulum–associated degradation (ERAD)-like mechanism.
Pallanck LJ. Culling sick mitochondria from the herd. J Cell Biol 2012;191(7):1225–1227. http://www.jcb.org/cgi/doi/10.1083/jcb.201011068
PINK1 and Parkin and Parkinson’s Disease
Studies of the familial Parkinson disease-related proteins PINK1 and Parkin have demonstrated that these factors promote the fragmentation and turnover of mitochondria following treatment of cultured cells with mitochondrial depolarizing agents. Whether PINK1 or Parkin influence mitochondrial quality control under normal physiological conditions in dopaminergic neurons, a principal cell type that degenerates in Parkinson disease, remains unclear. To address this matter, we developed a method to purify and characterize neural subtypes of interest from the adult Drosophila brain.
Using this method, we find that dopaminergic neurons from Drosophila parkin mutants accumulate enlarged, depolarized mitochondria, and that genetic perturbations that promote mitochondrial fragmentation and turnover rescue the mitochondrial depolarization and neurodegenerative phenotypes of parkin mutants. In contrast, cholinergic neurons from parkin mutants accumulate enlarged depolarized mitochondria to a lesser extent than dopaminergic neurons, suggesting that a higher rate of mitochondrial damage, or a deficiency in alternative mechanisms to repair or eliminate damaged mitochondria explains the selective vulnerability of dopaminergic neurons in Parkinson disease.
Our study validates key tenets of the model that PINK1 and Parkin promote the fragmentation and turnover of depolarized mitochondria in dopaminergic neurons. Moreover, our neural purification method provides a foundation to further explore the pathogenesis of Parkinson disease, and to address other neurobiological questions requiring the analysis of defined neural cell types.
Burmana JL, Yua S, Poole AC, Decala RB , Pallanck L. Analysis of neural subtypes reveals selective mitochondrial dysfunction in dopaminergic neurons from parkin mutants.
Autophagy in Parkinson’s Disease.
Parkinson’s disease is a common neurodegenerative disease in the elderly. To explore the specific role of autophagy and the ubiquitin-proteasome pathway in apoptosis, a specific proteasome inhibitor and macroautophagy inhibitor and stimulator were selected to investigate pheochromocytoma (PC12) cell lines transfected with human mutant (A30P) and wildtype (WT) -synuclein.
The apoptosis ratio was assessed by flow cytometry. LC3, heat shock protein 70 (hsp70) and caspase-3 expression in cell culture were determined by Western blot. The hallmarks of apoptosis and autophagy were assessed with transmission electron microscopy. Compared to the control group or the rapamycin (autophagy stimulator) group, the apoptosis ratio in A30P and WT cells was significantly higher after treatment with inhibitors of the proteasome and macroautophagy. The results of Western blots for caspase-3 expression were similar to those of flow cytometry; hsp70 protein was significantly higher in the proteasome inhibitor group than in control, but in the autophagy inhibitor and stimulator groups, hsp70 was similar to control. These findings show that inhibition of the proteasome and autophagy promotes apoptosis, and the macroautophagy stimulator rapamycin reduces the apoptosis ratio. And inhibiting or stimulating autophagy has less impact on hsp70 than the proteasome pathway.
In conclusion, either stimulation or inhibition of macroautophagy, has less impact on hsp70 than on the proteasome pathway. This study found that rapamycin decreased apoptotic cells in A30P cells independent of caspase-3 activity. Although several lines of evidence recently demonstrated crosstalk between autophagy and caspase-independent apoptosis, we could not confirm that autophagy activation protects cells from caspase-independent cell death. Undoubtedly, there are multiple connections between the apoptotic and autophagic processes.
Inhibition of autophagy may subvert the capacity of cells to remove damaged organelles or to remove misfolded proteins, which would favor apoptosis. However, proteasome inhibition activated macroautophagy and accelerated apoptosis. A likely explanation is inhibition of the proteasome favors oxidative reactions that trigger apoptosis, presumably through
1. a direct effect on mitochondria, and
2. the absence of NADPH2 and ATP
which may deinhibit the activation of caspase-2 or MOMP. Another possibility is that aggregated proteins induced by proteasome inhibition increase apoptosis.
Yang F, Yanga YP, Maoa CJ, Caoa BY, et al. Role of autophagy and proteasome degradation pathways in apoptosis of PC12 cells overexpressing human -synuclein. Neuroscience Letters 2009; 454:203–208. doi:10.1016/j.neulet.2009.03.027. http://www.elsevier.com/locate/neulet
Parkin-dependent Ubiquitination of Endogenous Bax
Autosomal recessive loss-of-function mutations within the PARK2 gene functionally inactivate the E3 ubiquitin ligase parkin, resulting in neurodegeneration of catecholaminergic neurons and a familial form of Parkinson disease. Current evidence suggests both a mitochondrial function for parkin and a neuroprotective role, which may in fact be interrelated. The antiapoptotic effects of Parkin have been widely reported, and may involve fundamental changes in the threshold for apoptotic cytochrome c release, but the substrate(s) involved in Parkin dependent protection had not been identified. Here, we demonstrate the Parkin-dependent ubiquitination of endogenous Bax comparing primary cultured neurons from WT and Parkin KO mice and using multiple Parkin-overexpressing cell culture systems. The direct ubiquitination of purified Bax was also observed in vitro following incubation with recombinant parkin. The authors found that Parkin prevented basal and apoptotic stress induced translocation of Bax to the mitochondria. Moreover, an engineered ubiquitination-resistant form of Bax retained its apoptotic function, but Bax KO cells complemented with lysine-mutant Bax did not manifest the antiapoptotic effects of Parkin that were observed in cells expressing WT Bax. These data suggest that Bax is the primary substrate responsible for the antiapoptotic effects of Parkin, and provide mechanistic insight into at least a subset of the mitochondrial effects of Parkin.
Johnson BN, Berger AK, Cortese GP, and LaVoie MJ. The ubiquitin E3 ligase Parkin regulates the proapoptotic function of Bax. PNAS 2012, pp 6. http://www.pnas.org/cgi/doi/10.1073/pnas.1113248109
Parkin Promotes Mitochondrial Loss in Autophagy
Parkin, an E3 ubiquitin ligase implicated in Parkinson’s disease, promotes degradation of dysfunctional mitochondria by autophagy. Using proteomic and cellular approaches, we show that upon translocation to mitochondria, Parkin activates the ubiquitin–proteasome system (UPS) for widespread degradation of outer membrane proteins. This is evidenced by an increase in K48-linked polyubiquitin on mitochondria, recruitment of the 26S proteasome and rapid degradation of multiple outer membrane proteins. The degradation of proteins by the UPS occurs independently of the autophagy pathway, and inhibition of the 26S proteasome completely abrogates Parkin-mediated mitophagy in HeLa, SH-SY5Y and mouse cells. Although the mitofusins Mfn1 and Mfn2 are rapid degradation targets of Parkin, degradation of additional targets is essential for mitophagy. These results indicate that remodeling of the mitochondrial outer membrane proteome is important for mitophagy, and reveal a causal link between the UPS and autophagy, the major pathways for degradation of intracellular substrates.
Chan NC, Salazar AM, Pham AH, Sweredoski MJ, et al. Broad activation of the ubiquitin–proteasome system by Parkin is critical for mitophagy. Human Molecular Genetics 2011; 20(9): 1726–1737. doi:10.1093/hmg/ddr048.
TRAP1 and TBP7 Interaction in Refolding of Damaged Proteins
TRAP1 is a mitochondrial antiapoptotic heat shock protein. The information available on the TRAP1 pathway describes just a few well-characterized functions of this protein in mitochondria. However, our group’s use of mass spectrometry analysis identified TBP7, an AAA-ATPase of the 19S proteasomal subunit, as a putative TRAP1-interacting protein. Surprisingly, TRAP1 and TBP7 co-localize in the endoplasmic reticulum (ER), as demonstrated by biochemical and confocal/electron microscopy analyses, and directly interact, as confirmed by FRET analysis. This is the first demonstration of TRAP1 presence in this cellular compartment. TRAP1 silencing by shRNAs, in cells exposed to thapsigargin-induced ER stress, correlates with up-regulation of BiP/Grp78, thus suggesting a role of TRAP1 in the refolding of damaged proteins and in ER stress protection. Consistently, TRAP1 and/or TBP7 interference enhanced stress-induced cell death and increased intracellular protein ubiquitination. These experiments led us to hypothesize an involvement of TRAP1 in protein quality control for mistargeted/misfolded mitochondria-destined proteins, through interaction with the regulatory proteasome protein TBP7. Remarkably, the expression of specific mitochondrial proteins decreased upon TRAP1 interference as a consequence of increased ubiquitination. The proposed TRAP1 network has an impact in vivo, since it is conserved in human colorectal cancers, is controlled by ER-localized TRAP1 interacting with TBP7 and provides a novel model of ER-mitochondria crosstalk.
Amoroso MR, Matassa DS, Laudiero G, Egorova AV. TRAP1 AND THE PROTEASOME REGULATORY PARTICLE TBP7/Rpt3 INTERACT IN THE ENDOPLASMIC RETICULUM AND CONTROL CELLULAR UBIQUITINATION OF SPECIFIC MITOCHONDRIAL PROTEINS. Cell Death and Differentiation 2012; pp? DOI : 10.1038/cdd.2011.128
VMS1 and Mitochondrial Protein Degradation
We show that Ydr049 (renamed VCP/Cdc48-associated mitochondrial stress-responsive—Vms1), a member of an unstudied pan-eukaryotic protein family, translocates from the cytosol to mitochondria upon mitochondrial stress. Cells lacking Vms1 show progressive mitochondrial failure, hypersensitivity to oxidative stress, and decreased chronological life span. Both yeast and mammalian Vms1 stably interact with Cdc48/VCP/p97, a component of the ubiquitin/proteasome system with a well-defined role in endoplasmic reticulum-associated protein degradation (ERAD), wherein misfolded ER proteins are degraded in the cytosol. We show that oxidative stress triggers mitochondrial localization of Cdc48 and this is dependent on Vms1. When this system is impaired by mutation of Vms1,
- ubiquitin-dependent mitochondrial protein degradation,
- mitochondrial respiratory function,and
- cell viability are compromised.
We demonstrate that Vms1 is a required component of an evolutionarily conserved system for mitochondrial protein degradation, which is
necessary to maintain
- mitochondrial,
- cellular, and
- organismal viability.
Heo JM, Livnat-Levanon N, Taylor EB, Jones KT. A Stress-Responsive System
for Mitochondrial Protein Degradation. Molecular Cell 2010; 40:465–480.
DOI 10.1016/j.molcel.2010.10.021
Mitochondrial Protein Degradation
The biogenesis of mitochondria and the maintenance of mitochondrial functions depends on an autonomous proteolytic system in the organelle which is highly conserved throughout evolution. Components of this system include processing
- peptidases and
- ATP-dependent proteases, as well as
- molecular chaperone proteins and
- protein complexes with apparently regulatory functions.
While processing peptidases mediate maturation of nuclear-encoded mitochondrial preproteins, quality control within various subcompartments of mitochondria is ensured by ATP-dependent proteases which selectively remove non-assembled or misfolded polypeptides. Moreover, these proteases appear to control the activity- or steady-state levels of specific regulatory proteins and thereby ensure mitochondrial genome integrity, gene expression and protein assembly.
Kaser M and Langer T. Protein degradation in mitochondria. CELL & DEVELOPMENTAL BIOLOGY 2000; 11:181–190. doi: 10.1006/10.1006/scdb.2000.0166.
RING finger E3s
Ubiquitin-ligases or E3s are components of the ubiquitin proteasome system (UPS) that coordinate the transfer of ubiquitin to the target protein. A major class of ubiquitin-ligases consists of RING-finger domain proteins that include the substrate recognition sequences in the same polypeptide; these are known as single-subunit RING finger E3s. We are studying a particular family of RING finger E3s, named ATL, that contain a transmembrane domain and the RING-H2 finger domain; none of the member of the family contains any other previously described domain. Although the study of a few members in A. thaliana and O. sativa has been reported, the role of this family in the life cycle of a plant is still vague.
To provide tools to advance on the functional analysis of this family we have undertaken a phylogenetic analysis of ATLs in twenty-four plant genomes. ATLs were found in all the 24 plant species analyzed, in numbers ranging from 20–28 in two basal species to 162 in soybean. Analysis of ATLs arrayed in tandem indicates that sets of genes are expanding in a species-specific manner. To
get insights into the domain architecture of ATLs we generated 75 pHMM LOGOs from 1815 ATLs, and unraveled potential protein-protein interaction regions by means of yeast two-hybrid assays. Several ATLs were found to interact with DSK2a/ubiquilin through a region at the amino-terminal end, suggesting that this is a widespread interaction that may assist in the mode of action of ATLs; the region was traced to a distinct sequence LOGO. Our analysis provides significant observations on the evolution and expansion of the ATL family in addition to information on the domain structure of this class of ubiquitin-ligases that may be involved in plant adaptation to environmental stress.
Aguilar-Hernandez V, Aguilar-Henonin L, Guzman P. Diversity in the Architecture of ATLs, a Family of Plant Ubiquitin-Ligases, Leads to Recognition and Targeting of Substrates in Different Cellular Environments. PLoS ONE 2011; 6(8): e23934. doi:10.1371/journal.pone.0023934
UPS Proteolytic Function Inadequate in Proteinopathies
Proteinopathies are a family of human disease caused by toxic aggregation-prone proteins and featured by the presence of protein aggregates in the affected cells. The ubiquitin-proteasome system (UPS) and autophagy are two major intracellular protein degradation pathways. The UPS mediates the targeted degradation of most normal proteins after performing their normal functions as well as the removal of abnormal, soluble proteins. Autophagy is mainly responsible for degradation of defective organelles and the bulk degradation of cytoplasm during starvation. The collaboration between the UPS and autophagy appears to be essential to protein quality control in the cell.
UPS proteolytic function often becomes inadequate in proteinopathies which may lead to activation of autophagy, striving to remove abnormal proteins especially the aggregated forms. HADC6, p62, and FoxO3 may play an important role in mobilizing this proteolytic consortium. Benign measures to enhance proteasome function are currently lacking; however, enhancement of autophagy via pharmacological intervention and/or lifestyle change has shown great promise in alleviating bona fide proteinopathies in the cell and animal models. These pharmacological interventions are expected to be applied clinically to treat human proteinopathies in the near future.
Zheng Q, Li J, Wang X. Interplay between the ubiquitin-proteasome system and
autophagy in proteinopathies. Int J Physiol Pathophysiol Pharmacol 2009;1:127-142. http://www.ijppp.org/IJPPP904002
Ubiquitin-associated Protein-Protein Interactions
Applicability of in vitro biotinylated ubiquitin for evaluation of endogenous ubiquitin conjugation and analysis of ubiquitin-associated protein-protein interactions has been investigated. Incubation of rat brain mitochondria with biotinylated ubiquitin followed by affinity chromatography on avidin-agarose, intensive washing, tryptic digestion of proteins bound to the affinity sorbent and their mass spectrometry analysis resulted in reliable identification of 50 proteins belonging to mitochondrial and extramitochondrial compartments. Since all these proteins were bound to avidin-agarose only after preincubation of the mitochondrial fraction with biotinylated ubiquitin, they could therefore be referred to as specifically bound proteins. A search for specific
ubiquitination signature masses revealed several extramitochondrial and intramitochondrial ubiquitinated proteins representing about 20% of total number of proteins bound to avidin-agarose. The interactome analysis suggests that the identified non-ubiquitinated proteins obviously form tight complexes either with ubiquitinated proteins or with their partners and/or mitochondrial membrane components. Results of the present study demonstrate that the use of biotinylated ubiquitin may be considered as the method of choice for in vitro evaluation of endogenous ubiquitin-conjugating machinery in particular
subcellular organelles and changes in ubiquitin/organelle associated interactomes. This may be useful for evaluation of changes in interactomes induced by protein ubiquitination.
Buneeva OA, Medvedeva MV, Kopylov AT, Zgoda VG, Medvedev AE. Use of Biotinylated Ubiquitin for Analysis of Rat Brain Mitochondrial Proteome and Interactome. Int J Mol Sci 2012; 13: 11593-11609; doi:10.3390/ijms130911593
IL-6 regulation on mitochondrial remodeling/dysfunction
Muscle protein turnover regulation during cancer cachexia is being rapidly defined, and skeletal muscle mitochondria function appears coupled to processes regulating muscle wasting. Skeletal muscle oxidative capacity and the expression of proteins regulating mitochondrial biogenesis and dynamics are disrupted in severely cachectic ApcMin/+ mice. It has not been determined if these changes occur at the onset of cachexia and are necessary for the progression of muscle wasting. Exercise and anti-cytokine therapies have proven effective in preventing cachexia development in tumor bearing mice, while their effect on mitochondrial content, biogenesis and dynamics is not well understood.
The purposes of this study were to
1) determine IL-6 regulation on mitochondrial remodeling/dysfunction during the progression of cancer cachexia and
2) to determine if exercise training can attenuate mitochondrial dysfunction and the induction of proteolytic pathways during IL-6 induced cancer cachexia.
ApcMin/+ mice were examined during the progression of cachexia, after systemic interleukin (IL)-6r antibody treatment, or after IL-6 over-expression with or without exercise. Direct effects of IL-6 on mitochondrial remodeling were examined in cultured C2C12 myoblasts.
Mitochondrial content was not reduced during the initial development of cachexia, while muscle PGC-1α and fusion (Mfn1, Mfn2) protein expression was repressed.
With progressive weight loss mitochondrial content decreased, PGC-1α and fusion proteins were further suppressed, and fission protein (FIS1) was induced.
IL-6 receptor antibody administration after the onset of cachexia
- improved mitochondrial content,
- PGC-1α,
- Mfn1/Mfn2 and
- FIS1 protein expression.
IL-6 over-expression in pre-cachectic mice
- accelerated body weight loss and muscle wasting, without reducing mitochondrial content,
- while PGC-1α and Mfn1/Mfn2 protein expression was suppressed
- and FIS1 protein expression induced.
Exercise normalized these IL-6 induced effects. C2C12 myotubes administered IL-6 had
- increased FIS1 protein expression,
- increased oxidative stress, and
- reduced PGC-1α gene expression
- without altered mitochondrial protein expression.
Altered expression of proteins regulating mitochondrial biogenesis and fusion are early events in the initiation of cachexia regulated by IL-6, which precede the loss of muscle mitochondrial content. Furthermore, IL-6 induced mitochondrial remodeling and proteolysis can be rescued with moderate exercise training even in the presence of high circulating IL-6 levels.
White JP, Puppa MJ, Sato S, Gao S. IL-6 regulation on skeletal muscle mitochondrial remodeling during cancer cachexia in the ApcMin/+ mouse. Skeletal Muscle 2012; 2:14-30.
http://www.skeletalmusclejournal.com/content/2/1/14
Starvation-induced Autophagy
Upon starvation cells undergo autophagy, a cellular degradation pathway important in the turnover of whole organelles and long lived proteins. Starvation-induced protein degradation has been regarded as an unspecific bulk degradation process. We studied global protein dynamics during amino acid starvation-induced autophagy by quantitative mass spectrometry and were able to record nearly 1500 protein profiles during 36 h of starvation. Cluster analysis of the recorded protein profiles revealed that cytosolic proteins were degraded rapidly, whereas proteins annotated to various complexes and organelles were degraded later at different time periods. Inhibition of protein degradation pathways identified the lysosomal/autophagosomal system as the main degradative route.
Thus, starvation induces degradation via autophagy, which appears to be selective and to degrade proteins in an ordered fashion and not completely arbitrarily as anticipated so far.
Kristensen AR, Schandorff S, Høyer-Hansen M, Nielsen MO, et al. Ordered Organelle Degradation during Starvation-induced Autophagy. Molecular & Cellular Proteomics 2008; 7:2419–2428.
Skeletal Muscle Macroautophagy
Skeletal muscles are the agent of motion and one of the most important tissues responsible for the control of metabolism. Coordinated movements are allowed by the highly organized structure of the cytosol of muscle fibers (or myofibers), the multinucleated and highly specialized cells of skeletal muscles involved in contraction. Contractile proteins are assembled into repetitive structures, the basal unit of which is the sarcomere, that are well packed into the myofiber cytosol. Myonuclei are located at the edge of the myofibers, whereas the various organelles such as mitochondria and sarcoplasmic reticulum are embedded among the myofibrils. Many different changes take place in the cytosol of myofibers during catabolic conditions:
- proteins are mobilized
- organelles networks are reorganized for energy needs
- the setting of myonuclei can be modified.
Further,
- strenuous physical activity,
- improper dietary regimens and
- aging
lead to mechanical and metabolic damages of myofiber organelles, especially mitochondria, and contractile proteins. During aging the protein turnover is slowed down, therefore it is easier to accumulate aggregates of dysfunctional proteins. Therefore, a highly dynamic tissue such as skeletal muscle requires a rapid and efficient system for the removal of altered organelles, the elimination of protein aggregates, and the disposal of toxic products.
The two major proteolytic systems in muscle are the ubiquitin-proteasome and the autophagy-lysosome pathways. The proteasome system requires
- the transcription of the two ubiquitin ligases (atrogin-1 and MuRF1) and
- the ubiquitination of the substrates.
Therefore, the ubiquitin-proteasome system can provide the rapid elimination of single proteins or small aggregates. Conversely, the autophagic system is able to degrade entire organelles and large proteins aggregates. In the autophagy-lysosome system, double-membrane vesicles named autophagosomes are able to engulf a portion of the cytosol and fuse with lysosomes, where their content is completely degraded by lytic enzymes.
The autophagy flux can be biochemicaly monitored following LC3 lipidation and p62 degradation. LC3 is the mammalian homolog of the yeast Atg8 gene, which is lipidated when recruited for the double-membrane commitment and growth. p62 (SQSTM-1) is a polyubiquitin-binding protein involved in the proteasome system and that can either reside free in the cytosol and nucleus or occur within autophagosomes and lysosomes. The GFP-LC3 transgenic mouse model allows easy detection of autophagosomes by simply monitoring the presence of bright GFP-positive puncta inside the myofibrils and beneath the plasma membrane of the myofibers, thus investigate the activation of autophagy in skeletal muscles with different contents of slow and fast-twitching myofibers and in response to stimuli such as fasting. For example, in the fast-twiching extensor digitorum longus muscle few GFP-LC3 dots were observed before starvation, while many small GFP-LC3 puncta appeared between myofibrils and in the perinuclear regions after 24 h starvation. Conversely, in the slow-twitching soleus muscle, autophagic puncta were almost absent in standard condition and scarcely induced after 24 h starvation.
Autophagy in Muscle Homeostasis
The autophagic flux was found to be increased during certain catabolic conditions, such as fasting, atrophy , and denervation , thus contributing to protein breakdown. Food deprivation is one of the strongest stimuli known to induce autophagy in muscle. Indeed skeletal muscle, after the liver, is the most responsive tissue to autophagy activation during food deprivation. Since muscles are the biggest reserve of amino acids in the body, during fasting autophagy has the vital role to maintain the amino acid pool by digesting muscular protein and organelles. In mammalian cells, mTORC1, which consists of
is the nutrient sensor that negatively regulates autophagy.
During atrophy, protein breakdown is mediated by atrogenes, which are under the forkhead box O (FoxO) transcription factors control, and activation of autophagy seems to aggravate muscle loss during atrophy. In vivo and in vitro studies demonstrated that several genes coding for components of the autophagic machinery, such as
- LC3,
- GABARAP,
- Vps34,
- Atg12 and
- Bnip3,
are controlled by FoxO3 transcription factor. FoxO3 is able to regulate independently the ubiquitin-proteasome system and the autophagy-lysosome machinery in vivo and in vitro. Denervation is also able to induce autophagy in skeletal muscle, although at a slower rate than fasting. This effect is mediated by RUNX1, a transcription factor upregulated during autophagy; the lack of RUNX1 results in excessive autophagic flux in denervated muscle and leads to atrophy. The generation of Atg5 and Atg7 muscle-specific knockout mice have shown that with suppression of autophagy both models display muscle weakness and atrophy and a significant reduction of weight, which is correlated with the important loss of muscle tissue due to an atrophic condition. An unbalanced autophagy flux is highly detrimental for muscle, as too much induces atrophy whereas too little leads to muscle weakness and degeneration. Muscle wasting associated with autophagy inhibition becomes evident and symptomatic only after a number of altered proteins and dysfunctional organelles are accumulated, a condition that becomes evident after months or even years. On the other hand, the excessive increase of autophagy flux is able to induce a rapid loss of muscle mass (within days or weeks).
Alterations of autophagy are involved in the pathogenesis of several myopathies and dystrophies.
The maintenance of muscle homeostasis is finely regulated by the balance between catabolic and anabolic process. Macroautophagy (or autophagy) is a catabolic process that provides the degradation of protein aggregation and damaged organelles through the fusion between autophagosomes and lysosomes. Proper regulation of the autophagy flux is fundamental for the homeostasis of skeletal muscles during physiological situations and in response to stress. Defective as well as excessive autophagy is harmful for muscle health and has a pathogenic role in several forms of muscle diseases.
Grumati P, Bonaldo P. Autophagy in Skeletal Muscle Homeostasis and in Muscular Dystrophies. Cells 2012, 1, 325-345; doi:10.3390/cells1030325. ISSN 2073-4409. http://www.mdpi.com/journal/cells
Parkinson’s Disease Mutations
Mutations in parkin, a ubiquitin ligase, cause early-onset familial Parkinson’s disease (AR-JP). How Parkin suppresses Parkinsonism remains unknown. Parkin was recently shown to promote the clearance of impaired mitochondria by autophagy, termed mitophagy. Here, we show that Parkin promotes mitophagy by catalyzing mitochondrial ubiquitination, which in turn recruits ubiquitin-binding autophagic components, HDAC6 and p62, leading to mitochondrial clearance.
During the process, juxtanuclear mitochondrial aggregates resembling a protein aggregate-induced aggresome are formed. The formation of these “mito-aggresome” structures requires microtubule motor-dependent transport and is essential for efficient mitophagy. Importantly, we show that AR-JP–causing Parkin mutations are defective in supporting mitophagy due to distinct defects at
- recognition,
- transportation, or
- ubiquitination of impaired mitochondria,
thereby implicating mitophagy defects in the development of Parkinsonism. Our results show that impaired mitochondria and protein aggregates are processed by common ubiquitin-selective autophagy machinery connected to the aggresomal pathway, thus identifying a mechanistic basis for the prevalence of these toxic entities in Parkinson’s disease.
Lee JY,Nagano Y, Taylor JP,Lim KL, and Yao TP. Disease-causing mutations in Parkin impair mitochondrial ubiquitination, aggregation, and HDAC6-dependent mitophagy. J Cell Biol 2010; 189(4):671-679. http://www.jcb.org/cgi/doi/10.1083/jcb.201001039
Drosophila Parkin Requires PINK1
Loss of the E3 ubiquitin ligase Parkin causes early onset Parkinson’s disease, a neurodegenerative disorder of unknown etiology. Parkin has been linked to multiple cellular processes including
- protein degradation,
- mitochondrial homeostasis, and
- autophagy;
however, its precise role in pathogenesis is unclear. Recent evidence suggests that Parkin is recruited to damaged mitochondria, possibly affecting
- mitochondrial fission and/or fusion,
- to mediate their autophagic turnover.
The precise mechanism of recruitment and the ubiquitination target are unclear. Here we show in Drosophila cells that PINK1 is required to recruit Parkin to dysfunctional mitochondria and promote their degradation. Furthermore, PINK1 and Parkin mediate the ubiquitination of the profusion factor Mfn on the outer surface of mitochondria. Loss of Drosophila PINK1 or parkin causes an increase in Mfn abundance in vivo and concomitant elongation of mitochondria. These findings provide a molecular mechanism by which the PINK1/Parkin pathway affects mitochondrial fission/fusion as suggested by previous genetic interaction studies. We hypothesize that Mfn ubiquitination may provide a mechanism by which terminally damaged mitochondria are labeled and sequestered for degradation by autophagy.
Ziviani E, Tao RN, and Whitworth AJ. Drosophila Parkin requires PINK1 for mitochondrial translocation and ubiquitinates Mitofusin. PNAS 2010. Pp6 http://www.pnas.org/cgi/doi/10.1073/pnas.0913485107
Dynamin-related protein 1 (Drp1) in Parkinson’s
Mutations in Parkin, an E3 ubiquitin ligase that regulates protein turnover, represent one of the major causes of familial Parkinson’s disease (PD), a neurodegenerative disorder characterized by the loss of dopaminergic neurons and impaired mitochondrial functions. The underlying mechanism by which pathogenic parkin mutations induce mitochondrial abnormality is not fully understood. Here we demonstrate that Parkin interacts with and subsequently ubiquitinates dynamin-related protein 1 (Drp1), for promoting its proteasome-dependent degradation. Pathogenic mutation or knockdown of Parkin inhibits the ubiquitination and degradation of Drp1, leading to an increased level of Drp1 for mitochondrial fragmentation. These results identify Drp1 as a novel substrate of Parkin and suggest a potential mechanism linking abnormal Parkin expression to mitochondrial dysfunction in the pathogenesis of PD.
Wang H, Song P, Du L, Tian W. Parkin ubiquitinates Drp1 for proteasome-dependent degradation: implication of dysregulated mitochondrial dynamics in Parkinson’s disease.
JBC Papers in Press. Published on February 3, 2011 as Manuscript M110.144238. http://www.jbc.org/cgi/doi/10.1074/jbc.M110.144238
Pink1, Parkin, and DJ-1 Form a Complex
Mutations in the genes PTEN-induced putative kinase 1 (PINK1), PARKIN, and DJ-1 cause autosomal recessive forms of Parkinson disease (PD), and the Pink1/Parkin pathway regulates mitochondrial integrity and function. An important question is whether the proteins encoded by these genes function to regulate activities of other cellular compartments. A study in mice, reported by Xiong et al. in this issue of the JCI, demonstrates that Pink1, Parkin, and DJ-1 can form a complex in the cytoplasm, with Pink1 and DJ-1 promoting the E3 ubiquitin ligase activity of Parkin to degrade substrates via the proteasome (see the related article, doi:10.1172/ JCI37617).
This protein complex in the cytosol may or may not be related to the role of these proteins in regulating mitochondrial function or oxidative stress in vivo.
Three models for the role of the PPD complex. In this issue of the JCI, Xiong et al. report that Pink1, Parkin, and DJ-1 bind to each other and form a PPD E3 ligase complex in which Pink1 and DJ-1 modulate Parkin-dependent ubiquitination and subsequent degradation of substrates via the proteasome. Previous work suggests that the Pink1/Parkin pathway regulates mitochondrial integrity and promotes mitochondrial fission in Drosophila.
(A) Parkin and DJ-1 may be recruited to the mitochondrial outer membrane during stress and interact with Pink1. These interactions may facilitate the ligase activity of Parkin, thereby facilitating the turnover of molecules that regulate mitochondrial dynamics and mitophagy. The PPD complex may have other roles in the cytosol that result in degradative ubiquitination and/or relay information from mitochondria to other cellular compartments.
(B) Alternatively, Pink1 may be released from mitochondria after cleavage to interact with DJ-1 and Parkin in the cytosol.
A and B differ in the site of action of the PPD complex and the cleavage status of Pink1.
The complex forms on the mitochondrial outer membrane potentially containing full-length Pink1 in A, and in the cytosol with cleaved Pink1 in B.
Lack of DJ-1 function results in phenotypes that are distinct from the mitochondrial phenotypes observed in null mutants of Pink1 or Parkin in Drosophila. Thus, although the PPD complex is illustrated here as regulating mitochondrial fission, the role of DJ-1 in vivo remains to be clarified.
(C) It is also possible that the action occurs in the cytosol and is independent of the function of Pink1/Parkin in regulating mitochondrial integrity and function.
The Xiong et al. study offers an entry point for explorations of the role of Pink1, Parkin, and DJ-1 in the cytoplasm. It remains to be shown whether Parkin, in complex with Pink1 and DJ-1, carries out protein degradation in vivo.
Li H, and Guo M. Protein degradation in Parkinson disease revisited: it’s complex. commentaries. J Clin Invest. doi:10.1172/JCI38619. http://www.jci.org
Xiong, H., et al. Parkin, PINK1, and DJ-1 form a ubiquitin E3 ligase complex promoting unfolded protein degradation. J. Clin. Invest. 2009; 119:650–660.
Mitochondrial Ubiquitin Ligase, MITOL, protects neuronal cells
Nitric oxide (NO) is implicated in neuronal cell survival. However, excessive NO production mediates neuronal cell death, in part via mitochondrial dysfunction. Here, we report that the mitochondrial ubiquitin ligase, MITOL, protects neuronal cells from mitochondrial damage caused by accumulation of S-nitrosylated microtubule associated protein 1B-light chain 1 (LC1). S-nitrosylation of LC1 induces a conformational change that serves both to activate LC1 and to promote its ubiquination by MITOL, indicating that microtubule
stabilization by LC1 is regulated through its interaction with MITOL. Excessive NO production can inhibit MITOL, and MITOL inhibition resulted in accumulation of S-nitrosylated LC1 following stimulation of NO production by calcimycin and N-methyl-D-aspartate. LC1 accumulation under these conditions resulted in mitochondrial dysfunction and neuronal cell death. Thus, the balance between LC1 activation by S-nitrosylation and down-regulation by MITOL is critical for neuronal cell survival. Our findings may contribute significantly to an understanding of the mechanisms of neurological diseases caused by nitrosative stress-mediated mitochondrial dysfunction.
Yonashiro R, Kimijima Y, Shimura T, Kawaguchi K, et al. Mitochondrial ubiquitin ligase MITOL blocks S-nitrosylated MAP1B-light chain 1-mediated mitochondrial dysfunction and neuronal cell death. PNAS; 2012. pp 6. http://www.pnas.org/cgi/doi/10.1073/pnas.1114985109
Ubiquitin–Proteasome System in Neurodegeneration
A common histopathological hallmark of most neurodegenerative diseases is the presence of aberrant proteinaceous inclusions inside affected neurons. Because these protein aggregates are detected using antibodies against components of the ubiquitin–proteasome system (UPS), impairment of this machinery for regulated proteolysis has been suggested to be at the root of neurodegeneration. This hypothesis has been difficult to prove in vivo owing to the lack of appropriate tools. The recent report of transgenic mice with ubiquitous expression of a UPS-reporter protein should finally make it possible to test in vivo the role of the UPS in neurodegeneration.
Hernandez F, Dıaz-Hernandez M, Avila J and Lucas JJ. Testing the ubiquitin–proteasome hypothesis of neurodegeneration in vivo. TRENDS in Neurosciences 2004; 27(2): 66-68.
ALP in Parkinson’s
The ubiquitin-proteasome system (UPS) and autophagy-lysosome pathway (ALP) are the two most important mechanisms that normally repair or remove abnormal proteins. Alterations in the function of these systems to degrade misfolded and aggregated proteins are being increasingly recognized as playing a pivotal role in the pathogenesis of many neurodegenerative disorders such as Parkinson’s disease. Dysfunction of the UPS has been already strongly implicated in the pathogenesis of this disease and, more recently, growing interest has been shown in identifying the role of ALP in neurodegeneration. Mutations of a-synuclein and the increase of intracellular concentrations of non-mutant a-synuclein have been associated with Parkinson’s disease phenotype.
The demonstration that a-synuclein is degraded by both proteasome and autophagy indicates a possible linkage between the dysfunction of the UPS or ALP and the occurrence of this disorder.The fact that mutant a-synucleins inhibit ALP functioning by tightly binding to the receptor on the lysosomal membrane for autophagy pathway further supports the assumption that impairment of the ALP may be related to the development of Parkinson’s disease.
In this review, we summarize the recent findings related to this topic and discuss the unique role of the ALP in this neurogenerative disorder and the putative therapeutic potential through ALP enhancement.
Pan Y, Kondo S, Le W, Jankovic J. The role of autophagy-lysosome pathway in
neurodegeneration associated with Parkinson’s disease. Brain 2008; 131: 1969-1978. doi:10.1093/brain/awm318.
Ubiquitin-Proteasome System in Parkinson’s
There is growing evidence that dysfunction of the mitochondrial respiratory chain and failure of the cellular protein degradation machinery, specifically the ubiquitin-proteasome system, play an important role in the pathogenesis of Parkinson’s disease. We now show that the corresponding pathways of these two systems are linked at the transcriptomic level in Parkinsonian substantia nigra. We examined gene expression in medial and lateral substantia nigra (SN) as well as in frontal cortex using whole genome DNA oligonucleotide microarrays. In this study, we use a hypothesis-driven approach in analysing microarray data to describe the expression of mitochondrial and ubiquitin-proteasomal system (UPS) genes in Parkinson’s disease (PD).
Although a number of genes showed up-regulation, we found an overall decrease in expression affecting the majority of mitochondrial and UPS sequences. The down-regulated genes include genes that encode subunits of complex I and the Parkinson’s-disease-linked UCHL1. The observed changes in expression were very similar for both medial and lateral SN and also affected the PD cerebral cortex. As revealed by “gene shaving” clustering analysis, there was a very significant correlation between the transcriptomic profiles of both systems including in control brains.
Therefore, the mitochondria and the proteasome form a higher-order gene regulatory network that is severely perturbed in Parkinson’s disease. Our quantitative results also suggest that Parkinson’s disease is a disease of more than one cell class, i.e. that it goes beyond the catecholaminergic neuron and involves glia as well.
Duke DC, Moran LB, Kalaitzakis ME, Deprez M, et al. Transcriptome analysis reveals link between proteasomal and mitochondrial pathways in Parkinson’s disease. Neurogenetics 2006; 7:139-148.
Bax Degradation a Novel Mechanism for Survival in Bcl-2 overexpressed cancer cells
The authors previously reported that proteasome inhibitors were able to overcome Bcl-2-mediated protection from apoptosis, and now show that inhibition of the proteasome activity in Bcl-2-overexpressing cells accumulates the proapoptotic Bax protein to mitochondrial cytoplasm, where it interacts to Bcl-2 protein. This event was followed by release of mitochondrial cytochrome c into the cytosol and activation of caspase-mediated apoptosis. In contrast, proteasome inhibition did not induce any apparent changes in Bcl-2 protein levels. In addition, treatment with a proteasome inhibitor increased levels of ubiquitinated forms of Bax protein, without any effects on Bax mRNA expression. They also established a cell-free Bax degradation assay in which an in vitro-translated, 35S-labeled Bax protein can be degraded by a tumor cell protein extract, inhibitable by addition of a proteasome inhibitor or depletion of the proteasome or ATP. The Bax degradation activity can be reconstituted in the proteasome-depleted supernatant by addition of a purified 20S proteasome or proteasome-enriched fraction. Finally, by using tissue samples of human prostate adenocarcinoma, they demonstrated that increased levels of Bax degradation correlated well with decreased levels of Bax protein and increased Gleason scores of prostate cancer. These studies strongly suggest that ubiquitin-proteasome-mediated Bax degradation is a novel survival mechanism in human cancer cells and that selective targeting of this pathway should provide a unique approach for treatment of human cancers, especially those overexpressing Bcl-2.
In the current study, These investigators report that
- (i) proteasome inhibition results in Bax accumulation before release of cytochrome c and induction of apoptosis, which is associated with the ability of proteasome inhibitors to overcome Bcl-2-mediated antiapoptotic function;
- (ii) Bax is regulated by an ATP-ubiquitin-proteasome-dependent degradation pathway; and
- (iii) decreased levels of Bax protein correlate with increased levels of Bax degradation in aggressive human prostate cancer.
Li B and Dou QP. Bax degradation by the ubiquitin-proteasome-dependent pathway: Involvement in tumor survival and progression. PNAS 2000; 97(8): 3851-3855. http://www.pnas.org
p97 and DBeQ, ATP-competitive p97 inhibitor
A major limitation to current studies on the biological functions of p97/Cdc48 is that there is no method to rapidly shut off its ATPase activity. Given the range of cellular processes in which Cdc48 participates, it is difficult to determine whether any particular phenotype observed in the existing mutants is due to a direct or indirect effect. Moreover, inhibition of p97 activity in animal cells by siRNA or expression of a dominant-negative version is challenged by its high abundance and is not suited to evaluating proximal phenotypic effects of p97 loss of function.
A specific small-molecule inhibitor of p97 would provide an important tool to investigate diverse functions of this essential ATPase associated with diverse cellular activities (AAA) ATPase and to evaluate its potential to be a therapeutic target in human disease. Cancer cells may be particularly sensitive to killing by suppression of protein degradation mechanisms, because they may exhibit a heightened dependency on these mechanisms to clear an elevated burden of quality-control substrates. For example, some cancers produce high levels of a specific protein that is a prominent quality-control substrate (e.g., Ig light chains in multiple myeloma) or produce high levels of reactive oxygen species, which can result in excessive protein damage via oxidation. Therefore, a specific p97 inhibitor would be a valuable research tool to investigate p97 function in cells.
We carried out a high-throughput screen to identify inhibitors of p97 ATPase activity. Dual-reporter cell lines that simultaneously express p97-dependent and p97-independent proteasome substrates were used to stratify inhibitors that emerged from the screen. N2,N4-dibenzylquinazoline-2,4-diamine (DBeQ) was identified as a selective,potent, reversible, and ATP-competitive p97 inhibitor.
DBeQ blocks multiple processes that have been shown by RNAi to depend on p97, including degradation of ubiquitin fusion degradation and endoplasmic reticulum-associated degradation pathway reporters, as well as autophagosome maturation. DBeQ also potently inhibits cancer cell growth and is more rapid than a proteasome inhibitor at mobilizing the executioner caspases-3 and -7.
Simultaneous inhibition of proteasome and histone deacetylase 6 (HDAC6) [which is required for autophagy results in synergistic killing of multiple myeloma cells]. Interestingly, more than one dozen human clinical trials (www.clinicaltrials.gov) combine bortezomib with the broad-spectrum HDAC inhibitor vorinostat, which is active toward HDAC6. Targeting p97
may provide an alternative route to achieving the same objective. Our results provide a rationale for targeting p97 in cancer therapy. Future work will provide molecular insight into how inhibition of p97 activity by DBeQ results in apoptosis and could strengthen the rationale for a p97-targeted cancer therapeutic.
Chou TF, Brown SJ, Minond D, Nordin BE, et al. Reversible inhibitor of p97, DBeQ, impairs both ubiquitin-dependent and autophagic protein clearance pathways. PNAS 2011; pp 6 http://www.pnas.org/cgi/doi/10.1073/pnas.1015312108
The causes of various neurodegenerative diseases, particularly sporadic cases, remain unknown, but increasing evidence suggests that these diseases may share similar molecular and cellular mechanisms of pathogenesis. One prominent feature common to most neurodegenerative diseases is the accumulation of misfolded proteins in the form of insoluble protein aggregates or inclusion bodies. Although these aggregates have different protein compositions, they all contain ubiquitin and proteasome subunits, implying a failure of the ubiquitin-proteasome system (UPS) in the removal of misfolded proteins.
A direct link between UPS dysfunction and neurodegeneration has been
provided by recent findings that genetic mutations in UPS components cause several rare, familial forms of neurodegenerative diseases. Furthermore, it is becoming increasingly clear that oxidative stress, which results from aging or exposure to environmental toxins, can directly damage UPS components, thereby contributing to the pathogenesis of sporadic forms of neurodegenerative diseases.
Aberrations in the UPS often result in defective proteasome-mediated protein degradation, leading to accumulation of toxic proteins and eventually to neuronal cell death. Interestingly, emerging evidence has begun to suggest that impairment in substrate-specific components of the UPS, such as E3 ubiquitin-protein ligases, may cause aberrant ubiquitination and neurodegeneration in a proteasome-independent manner. This provides an overview of the molecular components of the UPS and their impairment in familial and sporadic forms of neurodegenerative diseases, and summarizes present knowledge about the pathogenic mechanisms of UPS dysfunction in neurodegeneration.
Molecular mechanisms of protein ubiquitination and degradation by the UPS. Ubiquitination involves a highly specific enzyme cascade in which
- ubiquitin (Ub) is first activated by the ubiquitinactivating enzyme (E1),
- then transferred to an ubiquitin-conjugating enzyme (E2), and
- finally covalently attached to the substrate by an ubiquitin-protein ligase (E3).
Ubiquitination is a reversible posttranslational modification in which the removal of Ub is mediated by a deubiquitinating enzyme (DUB).
- Substrate proteins can be either monoubiquitinated or polyubiquitinated through successive conjugation of Ub moieties to an internal lysine residue in Ub.
- K48-linked poly-Ub chains are recognized by the 26S proteasome, resulting in degradation of the substrate and recycling of Ub.
- Monoubiquitination or K63-linked polyubiquitination plays a number of regulatory roles in cells that are proteasome-independent.
Parkin
Loss-of-function mutations in parkin, a 465-amino-acid RING-type E3 ligase, were first identified as the cause for autosomal recessive juvenile Parkinsonism (AR-JP) and subsequently found to account for ~50% of all recessively transmitted early-onset PD cases. Interestingly, patients with parkin mutations do not exhibit Lewy body pathology.
Possible pathogenic mechanisms by which impaired UPS components cause neurodegeneration. Genetic mutations or oxidative stress from aging and/or exposure to environmental toxins have been shown to impair the ubiquitination machinery (particularly E3 ubiquitin-protein ligases) and deubiquitinating enzymes (DUBs), resulting in abnormal ubiquitination. Depending on the type of ubiquitination affected, the impairment could cause neurodegeneration through two different mechanisms.
In the first model, aberrant K48-linked polyubiquitination resulting from impaired E3s or DUBs alters protein degradation by the proteasome, leading to accumulation of toxic proteins and subsequent neurodegeneration. The proteasomes could be directly damaged by oxidative stress or might be inhibited by protein aggregation, which exacerbates the neurotoxicity.
In the second model, aberrant monoubiquitination or K63-linked polyubiquitination resulting from impaired E3s or DUBs alters crucial non-proteasomal functions, such as gene transcription and protein trafficking, thereby causing neurodegeneration without protein aggregation.
These two models are not mutually exclusive because a single E3 or DUB enzyme, such as parkin or UCH-L1, could regulate more than one type of ubiquitination. In addition, abnormal ubiquitination and neurodegeneration could also result from mutation or oxidative stress-induced structural changes in the protein substrates that alter their recognition and degradation by the UPS.
Lian Li and Chin LS. IMPAIRMENT OF THE UBIQUITIN-PROTEASOME SYSTEM: A COMMON PATHOGENIC MECHANISM IN NEURODEGENERATIVE DISORDERS. In The Ubiquitin Proteasome System…Chapter 23. (Eds: Eds: Mario Di Napoli and Cezary Wojcik) 553-577 © 2007 Nova Science Publishers, Inc. ISBN 978-1-60021-749-4.
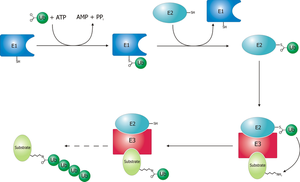
filedesc Schematic diagram of the ubiquitylation system. Created by Roger B. Dodd (Photo credit: Wikipedia)
Current Noteworthy Work
Nassif M and Hetz C. Autophagy impairment: a crossroad between neurodegeneration and tauopathies. BMC Biology 2012; 10:78. http://www.biomedcentral.com/1741-7007/10/78
Impairment of protein degradation pathways such as autophagy is emerging as a consistent and transversal pathological phenomenon in neurodegenerative diseases, including Alzheimer´s, Huntington´s, and Parkinson´s disease. Genetic inactivation of autophagy in mice has demonstrated a key role of the pathway in maintaining protein homeostasis in the brain, triggering massive neuronal loss and the accumulation of abnormal protein inclusions. A paper in Molecular Neurodegeneration from Abeliovich´s group now suggests a role for phosphorylation of Tau and the activation of glycogen synthase kinase 3β (GSK3β) in driving neurodegeneration in autophagy-deficient neurons. We discuss the implications of this study for understanding the factors driving neurofibrillary tangle formation in Alzheimer´s disease and tauopathies.
Cajee UF, Hull R and Ntwasa M. Modification by Ubiquitin-Like Proteins: Significance in Apoptosis and Autophagy Pathways. Int. J. Mol. Sci. 2012, 13, 11804-11831; doi:10.3390/ijms130911804
Ubiquitin-like proteins (Ubls) confer diverse functions on their target proteins. The modified proteins are involved in various biological processes, including DNA replication, signal transduction, cell cycle control, embryogenesis, cytoskeletal regulation,
metabolism, stress response, homeostasis and mRNA processing. Modifiers such as SUMO, ATG12, ISG15, FAT10, URM1, and UFM have been shown to modify proteins thus conferring functions related to programmed cell death, autophagy and regulation of
the immune system. Putative modifiers such as Domain With No Name (DWNN) have been identified in recent times but not fully characterized. In this review, we focus on cellular processes involving human Ubls and their targets.
Aloy P. Shaping the future of interactome networks. (A report of the third Interactome Networks Conference, Hinxton, UK, 29 August-1 September 2007). Genome Biology 2007; 8:316 (doi:10.1186/gb-2007-8-10-316)
Complex systems are often networked, and biology is no exception. Following on from the genome sequencing projects,
experiments show that proteins in living organisms are highly connected, which helps to explain how such great complexity
can be achieved by a comparatively small set of gene products. At a recent conference on interactome networks held outside
Cambridge, UK, the most recent advances in research on cellular networks were discussed. This year’s conference focused on
identifying the strengths and weaknesses of currently resolved interaction networks and the techniques used to determine
them – reflecting the fact that the field of mapping interaction networks is maturing.
Peroutka RJ, Orcutt SJ, Strickler JE, and Butt TR. SUMO Fusion Technology for Enhanced Protein Expression and Purification in Prokaryotes and Eukaryotes. Chapter 2. in T.C. Evans, M.-Q. Xu (eds.), Heterologous Gene Expression in E. coli, Methods in Molecular Biology 705:15-29. DOI 10.1007/978-1-61737-967-3_2, © Springer Science+Business Media, LLC 2011
The preparation of sufficient amounts of high-quality protein samples is the major bottleneck for structural proteomics. The use of recombinant proteins has increased significantly during the past decades. The most commonly used host, Escherichia coli, presents many challenges including protein misfolding, protein degradation, and low solubility. A novel SUMO fusion technology appears to enhance protein expression and solubility (www.lifesensors.com). Efficient removal of the SUMO tag by SUMO protease in vitro facilitates the generation of target protein with a native N-terminus. In addition to its physiological relevance in eukaryotes, SUMO can be used as a powerful biotechnology tool for enhanced functional protein expression in prokaryotes and eukaryotes.
Juang YC, Landry MC, et al. OTUB1 Co-opts Lys48-Linked Ubiquitin Recognition to Suppress E2 Enzyme Function. Molecular Cell 2012; 45: 384–397. DOI 10.1016/j.molcel.2012.01.011
Ubiquitylation entails the concerted action of E1, E2, and E3 enzymes. We recently reported that OTUB1, a deubiquitylase, inhibits the DNA damage response independently of its isopeptidase activity. OTUB1 does so by blocking ubiquitin transfer by UBC13, the cognate E2 enzyme for RNF168. OTUB1 also inhibits E2s of the UBE2D and UBE2E families. Here we elucidate the structural mechanism by which OTUB1 binds E2s to inhibit ubiquitin transfer. OTUB1 recognizes ubiquitin-charged E2s through contacts with both donor ubiquitin and the E2 enzyme. Surprisingly, free ubiquitin associates with the canonical distal ubiquitin-binding site on OTUB1 to promote formation of the inhibited E2 complex. Lys48 of donor ubiquitin lies near the OTUB1 catalytic site and the C terminus of free ubiquitin, a configuration that mimics the products of Lys48-linked ubiquitin chain cleavage. OTUB1 therefore co-opts Lys48-linked ubiquitin chain recognition to suppress ubiquitin conjugation and the DNA damage response.
Hunter T. The Age of Crosstalk: Phosphorylation, Ubiquitination, and Beyond. Molecular Cell 2007; 28:730-738. DOI 10.1016/ j.molcel.2007.11.019.
Crosstalk between different types of posttranslational modification is an emerging theme in eukaryotic biology. Particularly prominent are the multiple connections between phosphorylation and ubiquitination, which act either positively or negatively in both directions to regulate these processes.
Tu Y, Chen C, et al. The Ubiquitin Proteasome Pathway (UPP) in the regulation of cell cycle control and DNA damage repair and its implication in tumorigenesis. Int J Clin Exp Pathol 2012;5(8):726-738. www.ijcep.com /ISSN:1936-2625/IJCEP1208018
Accumulated evidence supports that the ubiquitin proteasome pathway (UPP) plays a crucial role in protein
metabolism implicated in the regulation of many biological processes such as cell cycle control, DNA damage
response, apoptosis, and so on. Therefore, alterations for the ubiquitin proteasome signaling or functional impairments
for the ubiquitin proteasome components are involved in the etiology of many diseases, particularly in cancer
development.The authors discuss the ubiquitin proteasome pathway in the regulation of cell cycle control and DNA
damage response, the relevance for the altered regulation of these signaling pathways in tumorigenesis, and finally
assess and summarize the advancement for targeting the ubiquitin proteasome pathway in cancer therapy.
Cebollero E , Reggiori F and Kraft C. Ribophagy: Regulated Degradation of Protein Production Factories. Int J Cell Biol. 2012; 2012: 182834. doi: 10.1155/2012/182834 (online).
During autophagy, cytosol, protein aggregates, and organelles are sequestered into double-membrane vesicles called autophagosomes and delivered to the lysosome/vacuole for breakdown and recycling of their basic components. In all eukaryotes this pathway is important for adaptation to stress conditions such as nutrient deprivation, as well as to regulate intracellular homeostasis by adjusting organelle number and clearing damaged structures. For a long time, starvation-induced autophagy has been viewed as a nonselective transport pathway; however, recent studies have revealed that autophagy is able to selectively engulf specific structures, ranging from proteins to entire organelles. In this paper, we discuss recent findings on the mechanisms and physiological implications of two selective types of autophagy: ribophagy, the specific degradation of ribosomes, and reticulophagy, the selective elimination of portions of the ER.
Lee JH, Yu WH,…, Nixon RA. Lysosomal Proteolysis and Autophagy Require Presenilin 1 and Are Disrupted by Alzheimer-Related PS1 Mutations. Cell 2010; 141, 1146–1158. DOI 10.1016/j.cell.2010.05.008.
Macroautophagy is a lysosomal degradative pathway essential for neuron survival. Here, we show that macroautophagy requires the Alzheimer’s disease (AD)-related protein presenilin-1 (PS1). In PS1 null blastocysts, neurons from mice hypomorphic for PS1 or
conditionally depleted of PS1, substrate proteolysis and autophagosome clearance during macroautophagy are prevented as a result of a selective impairment of autolysosome acidification and cathepsin activation. These deficits are caused by failed PS1-dependent
targeting of the v-ATPase V0a1 subunit to lysosomes. N-glycosylation of the V0a1 subunit, essential for its efficient ER-to-lysosome delivery, requires the selective binding of PS1 holoprotein to the unglycosylated subunit and the sec61alpha/ oligosaccharyltransferase complex. PS1 mutations causing early-onset AD produce a similar lysosomal/autophagy phenotype in
fibroblasts from AD patients. PS1 is therefore essential for v-ATPase targeting to lysosomes, lysosome acidification, and proteolysis during autophagy. Defective lysosomal proteolysis represents a basis for pathogenic protein accumulations and neuronal cell death in AD and suggests previously unidentified therapeutic targets.
Pohl C and Jentsch S. Midbody ring disposal by autophagy is a post-abscission event of cytokinesis. nature cell biology 2009; 11 (1): 65-70. DOI: 10.1038/ncb1813.
At the end of cytokinesis, the dividing cells are connected by an intercellular bridge, containing the midbody along with a single,
densely ubiquitylated, circular structure called the midbody ring (MR). Recent studies revealed that the MR serves as a target
site for membrane delivery and as a physical barrier between the prospective daughter cells. The MR materializes in telophase,
localizes to the intercellular bridge during cytokinesis, and moves asymmetrically into one cell after abscission. Daughter
cells rarely accumulate MRs of previous divisions, but how these large structures finally disappear remains unknown.
Here, we show that MRs are discarded by autophagy, which involves their sequestration into autophagosomes and delivery to
lysosomes for degradation. Notably, autophagy factors, such as the ubiquitin adaptor p62 and the ubiquitin-related protein Atg8 , associate with the MR during abscission, suggesting that autophagy is coupled to cytokinesis. Moreover, MRs accumulate in cells of patients with lysosomal storage disorders, indicating that defective MR disposal is characteristic of these diseases. Thus our findings suggest that autophagy has a broader role than previously assumed, and that cell renovation by clearing from superfluous large macromolecular assemblies, such as MRs, is an important autophagic function.
Hanai JI, Cao P, Tanksale P, Imamura S, et al. The muscle-specific ubiquitin ligase atrogin-1/MAFbx mediates statin-induced muscle toxicity. The Journal of Clinical Investigation 2007; 117(12):3930-3951. http://www.jci.org
Statins inhibit HMG-CoA reductase, a key enzyme in cholesterol synthesis, and are widely used to treat hypercholesterolemia.
These drugs can lead to a number of side effects in muscle, including muscle fiber breakdown; however, the mechanisms of muscle injury by statins are poorly understood. We report that lovastatin induced the expression of atrogin-1, a key gene involved in skeletal muscle atrophy, in humans with statin myopathy, in zebrafish embryos, and in vitro in murine skeletal muscle cells. In cultured mouse myotubes, atrogin-1 induction following lovastatin treatment was accompanied by distinct morphological changes, largely absent in
atrogin-1 null cells. In zebrafish embryos, lovastatin promoted muscle fiber damage, an effect that was closely mimicked by knockdown of zebrafish HMG-CoA reductase. Moreover, atrogin-1 knockdown in zebrafish embryos prevented lovastatin-induced muscle injury. Finally, overexpression of PGC-1α, a transcriptional coactivator that induces mitochondrial biogenesis and protects against the development of muscle atrophy, dramatically prevented lovastatin-induced muscle damage and abrogated atrogin-1 induction both in fish and in cultured mouse myotubes. Collectively, our human, animal, and in vitro findings shed light on the molecular mechanism of statin-induced myopathy and suggest that atrogin-1 may be a critical mediator of the muscle
damage induced by statins.
Inami Y, Waguri S, Sakamoto A, Kouno T, et al. Persistent activation of Nrf2 through p62 in hepatocellular carcinoma cells. J. Cell Biol. 2011; 193(2): 275–284. http://www.jcb.org/cgi/doi/10.1083/jcb.201102031
Macroautophagy (hereafter referred to as autophagy) is a cellular degradation system in which cytoplasmic components, including
organelles, are sequestered by double membrane structures called autophagosomes and the sequestered materials are
degraded by lysosomal hydrolases for supply of amino acids and for cellular homeostasis. Although autophagy has generally been considered nonselective, recent studies have shed light on another indispensable role for basal autophagy in cellular homeostasis, which is mediated by selective degradation of a specific substrate(s). p62 is a ubiquitously expressed cellular protein that is conserved in metazoa but not in plants and fungi, and recently it has been known as one of the selective substrates for autophagy.
This protein is localized at the autophagosome formation site and directly interacts with LC3, an autophagosome localizing protein . Subsequently, the p62 is incorporated into the autophagosome and then degraded. Therefore, impaired autophagy is accompanied by
accumulation of p62 followed by the formation of p62 and ubiquitinated protein aggregates because of the nature of both self- oligomerization and ubiquitin binding of p62.
Kima K, Khayrutdinov BI, Leeb CK, et al. Solution structure of the Zβ domain of human DNA-dependent activator of IFN-regulatory factors and its binding modes to B- and Z-DNAs. PNAS 2010; Early Edition ∣ pp 6. www.pnas.org/cgi/doi/10.1073/pnas.1014898107
The DNA-dependent activator of IFN-regulatory factors (DAI), also known as DLM-1/ZBP1, initiates an innate immune response by binding to foreign DNAs in the cytosol. For full activation of the immune response, three DNA binding domains at the N terminus are required: two Z-DNA binding domains (ZBDs), Zα and Zβ, and an adjacent putative B-DNA binding domain. The crystal structure of the Zβ domain of human DAI (hZβDAI) in complex with Z-DNA revealed structural features distinct from other known Z-DNA binding proteins, and it was classified as a group II ZBD. To gain structural insights into the DNA binding mechanism of hZβDAI, the solution structure of the free hZβDAI was solved, and its bindings to B- and Z-DNAs were analyzed by NMR spectroscopy. Compared to the Z-DNA–bound structure, the conformation of free hZβDAI has notable alterations in the α3 recognition helix, the “wing,” and Y145, which are critical in Z-DNA recognition. Unlike some other Zα domains, hZβDAI appears to have conformational flexibility, and structural adaptation is required for Z-DNA binding. Chemical-shift perturbation experiments revealed that hZβDAI also binds weakly to B-DNA via a different binding mode. The C-terminal domain of DAI is reported to undergo a conformational change on B-DNA binding; thus, it is possible that these changes are correlated. During the innate immune response, hZβDAI is likely to play an active role in binding to DNAs in both B and Z conformations in the recognition of foreign DNAs.
Epicrisis
This extensive review leaves little left unopened. We have seen the central role that the UPS system plays in normal organelle proteolysis in concert with autophagy. Impaired ubiquitination occurs from aging, and/or toxins, under oxidative stress involving E3s or DUBs.
This leads to altered gene transcripton, altered protein trafficking, and plays a role in neurodegenative disease, muscle malfunction, and cancer as well.

English: A cartoon representation of a lysine 48-linked diubiquitin molecule. The two ubiquitin chains are shown as green cartoons with each chain labelled. The components of the linkage are indicated and shown as orange sticks. Image was created using PyMOL from PDB id 1aar. (Photo credit: Wikipedia)

Different forms of protein ubiquitylation (Photo credit: Wikipedia)
Related articles
Read Full Post »




















 The Cardioid code developed by a team of Livermore and
The Cardioid code developed by a team of Livermore and 