
English: The citric acid cycle, also known as the tricarboxylic acid cycle (TCA cycle) or the Krebs cycle. Produced at WikiPathways. (Photo credit: Wikipedia)
Expanding the Genetic Alphabet and Linking the Genome to the Metabolome
Reporter& Curator: Larry Bernstein, MD, FCAP
Unlocking the diversity of genomic expression within tumorigenesis and “tailoring” of therapeutic options
1. Reshaping the DNA landscape between diseases and within diseases by the linking of DNA to treatments
In the NEW York Times of 9/24,2012 Gina Kolata reports on four types of breast cancer and the reshaping of breast cancer DNA treatment based on the findings of the genetically distinct types, which each have common “cluster” features that are driving many cancers. The discoveries were published online in the journal Nature on Sunday (9/23). The study is considered the first comprehensive genetic analysis of breast cancer and called a roadmap to future breast cancer treatments. I consider that if this is a landmark study in cancer genomics leading to personalized drug management of patients, it is also a fitting of the treatment to measurable “combinatorial feature sets” that tie into population biodiversity with respect to known conditions. The researchers caution that it will take years to establish transformative treatments, and this is clearly because in the genetic types, there are subsets that have a bearing on treatment “tailoring”. In addition, there is growing evidence that the Watson-Crick model of the gene is itself being modified by an expansion of the alphabet used to construct the DNA library, which itself will open opportunities to explain some of what has been considered junk DNA, and which may carry essential information with respect to metabolic pathways and pathway regulation. The breast cancer study is tied to the “Cancer Genome Atlas” Project, already reported. It is expected that this work will tie into building maps of genetic changes in common cancers, such as, breast, colon, and lung. What is not explicit I presume is a closely related concept, that the translational challenge is closely related to the suppression of key proteomic processes tied into manipulating the metabolome.
Saha S. Impact of evolutionary selection on functional regions: The imprint of evolutionary selection on ENCODE regulatory elements is manifested between species and within human populations. 9/12/2012. PharmaceuticalIntelligence.Wordpress.com
Hawrylycz MJ, Lein ES, Guillozet-Bongaarts AL, Shen EH, Ng L, et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature Sept 14-20, 2012
Sarkar A. Prediction of Nucleosome Positioning and Occupancy Using a Statistical Mechanics Model. 9/12/2012. PharmaceuticalIntelligence.WordPress.com
Heijden et al. Connecting nucleosome positions with free energy landscapes. (Proc Natl Acad Sci U S A. 2012, Aug 20 [Epub ahead of print]). http://www.ncbi.nlm.nih.gov/pubmed/22908247
2. Fiddling with an expanded genetic alphabet – greater flexibility in design of treatment (pharmaneogenesis?)

Diagram of DNA polymerase extending a DNA strand and proof-reading. (Photo credit: Wikipedia)
A clear indication of this emerging remodeling of the genetic alphabet is a new
study led by scientists at The Scripps Research Institute appeared in the
June 3, 2012 issue of Nature Chemical Biology that indicates the genetic code as
we know it may be expanded to include synthetic and unnatural sequence pairing (Study Suggests Expanding the Genetic Alphabet May Be Easier than Previously Thought, Genome). They infer that the genetic instructions for living organisms
that is composed of four bases (C, G, A and T)— is open to unnatural letters. An expanded “DNA alphabet” could carry more information than natural DNA, potentially coding for a much wider range of molecules and enabling a variety of powerful applications. The implications of the application of this would further expand the translation of portions of DNA to new transciptional proteins that are heretofore unknown, but have metabolic relavence and therapeutic potential. The existence of such pairing in nature has been studied in Eukariotes for at least a decade, and may have a role in biodiversity. The investigators show how a previously identified pair of artificial DNA bases can go through the DNA replication process almost as efficiently as the four natural bases. This could as well be translated into human diversity, and human diseases.
The Romesberg laboratory collaborated on the new study and his lab have been trying to find a way to extend the DNA alphabet since the late 1990s. In 2008, they developed the efficiently replicating bases NaM and 5SICS, which come together as a complementary base pair within the DNA helix, much as, in normal DNA, the base adenine (A) pairs with thymine (T), and cytosine (C) pairs with guanine (G). It had been clear that their chemical structures lack the ability to form the hydrogen bonds that join natural base pairs in DNA. Such bonds had been thought to be an absolute requirement for successful DNA replication, but that is not the case because other bonds can be in play.
The data strongly suggested that NaM and 5SICS do not even approximate the edge-to-edge geometry of natural base pairs—termed the Watson-Crick geometry, after the co-discoverers of the DNA double-helix. Instead, they join in a looser, overlapping, “intercalated” fashion that resembles a ‘mispair.’ In test after test, the NaM-5SICS pair was efficiently replicable even though it appeared that the DNA polymerase didn’t recognize it. Their structural data showed that the NaM-5SICS pair maintain an abnormal, intercalated structure within double-helix DNA—but remarkably adopt the normal, edge-to-edge, “Watson-Crick” positioning when gripped by the polymerase during the crucial moments of DNA replication. NaM and 5SICS, lacking hydrogen bonds, are held together in the DNA double-helix by “hydrophobic” forces, which cause certain molecular structures (like those found in oil) to be repelled by water molecules, and thus to cling together in a watery medium.
The finding suggests that NaM-5SICS and potentially other, hydrophobically bound base pairs could be used to extend the DNA alphabet and that Evolution’s choice of the existing four-letter DNA alphabet—on this planet—may have been developed allowing for life based on other genetic systems.
3. Studies that consider a DNA triplet model that includes one or more NATURAL nucleosides and looks closely allied to the formation of the disulfide bond and oxidation reduction reaction.
This independent work is being conducted based on a similar concep. John Berger, founder of Triplex DNA has commented on this. He emphasizes Sulfur as the most important element for understanding evolution of metabolic pathways in the human transcriptome. It is a combination of sulfur 34 and sulphur 32 ATMU. S34 is element 16 + flourine, while S32 is element 16 + phosphorous. The cysteine-cystine bond is the bridge and controller between inorganic chemistry (flourine) and organic chemistry (phosphorous). He uses a dual spelling, using sulfphur to combine the two referring to the master catalyst of oxidation-reduction reactions. Various isotopic alleles (please note the duality principle which is natures most important pattern). Sulfphur is Methionine, S adenosylmethionine, cysteine, cystine, taurine, gluthionine, acetyl Coenzyme A, Biotin, Linoic acid, H2S, H2SO4, HSO3-, cytochromes, thioredoxin, ferredoxins, purple sulfphur anerobic bacteria prokaroytes, hydrocarbons, green sulfphur bacteria, garlic, penicillin and many antibiotics; hundreds of CSN drugs for parasites and fungi antagonists. These are but a few names which come to mind. It is at the heart of the Krebs cycle of oxidative phosphorylation, i.e. ATP. It is also a second pathway to purine metabolism and nucleic acids. It literally is the key enzymes between RNA and DNA, ie, SH thiol bond oxidized to SS (dna) cysteine through thioredoxins, ferredoxins, and nitrogenase. The immune system is founded upon sulfphur compounds and processes. Photosynthesis Fe4S4 to Fe2S3 absorbs the entire electromagnetic spectrum which is filtered by the Allen belt some 75 miles above earth. Look up chromatium vinosum or allochromatium species. There is reasonable evidence it is the first symbiotic species of sulfphur anerobic bacteria (Fe4S4) with high potential mvolts which drives photosynthesis while making glucose with H2S.
He envisions a sulfphur control map to automate human metabolism with exact timing sequences, at specific three dimensional coordinates on Bravais crystalline lattices. He proposes adding the inosine-xanthosine family to the current 5 nucleotide genetic code. Finally, he adds, the expanded genetic code is populated with “synthetic nucleosides and nucleotides” with all kinds of customized functional side groups, which often reshape nature’s allosteric and physiochemical properties. The inosine family is nature’s natural evolutionary partner with the adenosine and guanosine families in purine synthesis de novo, salvage, and catabolic degradation. Inosine has three major enzymes (IMPDH1,2&3 for purine ring closure, HPGRT for purine salvage, and xanthine oxidase and xanthine dehydrogenase.
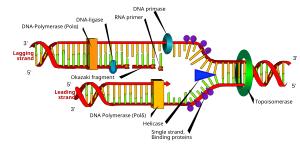
English: DNA replication or DNA synthesis is the process of copying a double-stranded DNA molecule. This process is paramount to all life as we know it. (Photo credit: Wikipedia)
3. Nutritional regulation of gene expression, an essential role of sulfur, and metabolic control
Finally, the research carried out for decades by Yves Ingenbleek and the late Vernon Young warrants mention. According to their work, sulfur is again tagged as essential for health. Sulfur (S) is the seventh most abundant element measurable in human tissues and its provision is mainly insured by the intake of methionine (Met) found in plant and animal proteins. Met is endowed with unique functional properties as it controls the ribosomal initiation of protein syntheses, governs a myriad of major metabolic and catalytic activities and may be subjected to reversible redox processes contributing to safeguard protein integrity.
Consuming diets with inadequate amounts of methionine (Met) are characterized by overt or subclinical protein malnutrition, and it has serious morbid consequences. The result is reduction in size of their lean body mass (LBM), best identified by the serial measurement of plasma transthyretin (TTR), which is seen with unachieved replenishment (chronic malnutrition, strict veganism) or excessive losses (trauma, burns, inflammatory diseases). This status is accompanied by a rise in homocysteine, and a concomitant fall in methionine. The ratio of S to N is quite invariant, but dependent on source. The S:N ratio is typical 1:20 for plant sources and 1:14.5 for animal protein sources. The key enzyme involved with the control of Met in man is the enzyme cystathionine-b-synthase, which declines with inadequate dietary provision of S, and the loss is not compensated by cobalamine for CH3- transfer.
As a result of the disordered metabolic state from inadequate sulfur intake (the S:N ratio is lower in plants than in animals), the transsulfuration pathway is depressed at cystathionine-β-synthase (CβS) level triggering the upstream sequestration of homocysteine (Hcy) in biological fluids and promoting its conversion to Met. They both stimulate comparable remethylation reactions from homocysteine (Hcy), indicating that Met homeostasis benefits from high metabolic priority. Maintenance of beneficial Met homeostasis is counterpoised by the drop of cysteine (Cys) and glutathione (GSH) values downstream to CβS causing reducing molecules implicated in the regulation of the 3 desulfuration pathways
4. The effect on accretion of LBM of protein malnutrition and/or the inflammatory state: in closer focus
Hepatic synthesis is influenced by nutritional and inflammatory circumstances working concomitantly and liver production of TTR integrates the dietary and stressful components of any disease spectrum. Thus we have a depletion of visceral transport proteins made by the liver and fat-free weight loss secondary to protein catabolism. This is most accurately reflected by TTR, which is a rapid turnover protein, but it is involved in transport and is essential for thyroid function (thyroxine-binding prealbumin) and tied to retinol-binding protein. Furthermore, protein accretion is dependent on a sulfonation reaction with 2 ATP. Consequently, Kwashiorkor is associated with thyroid goiter, as the pituitary-thyroid axis is a major sulfonation target. With this in mind, it is not surprising why TTR is the sole plasma protein whose evolutionary patterns closely follow the shape outlined by LBM fluctuations. Serial measurement of TTR therefore provides unequaled information on the alterations affecting overall protein nutritional status. Recent advances in TTR physiopathology emphasize the detecting power and preventive role played by the protein in hyper-homocysteinemic states.
Individuals submitted to N-restricted regimens are basically able to maintain N homeostasis until very late in the starvation processes. But the N balance study only provides an overall estimate of N gains and losses but fails to identify the tissue sites and specific interorgan fluxes involved. Using vastly improved methods the LBM has been measured in its components. The LBM of the reference man contains 98% of total body potassium (TBK) and the bulk of total body sulfur (TBS). TBK and TBS reach equal intracellular amounts (140 g each) and share distribution patterns (half in SM and half in the rest of cell mass). The body content of K and S largely exceeds that of magnesium (19 g), iron (4.2 g) and zinc (2.3 g).
TBN and TBK are highly correlated in healthy subjects and both parameters manifest an age-dependent curvilinear decline with an accelerated decrease after 65 years. Sulfur Methylation (SM) undergoes a 15% reduction in size per decade, an involutive process. The trend toward sarcopenia is more marked and rapid in elderly men than in elderly women decreasing strength and functional capacity. The downward SM slope may be somewhat prevented by physical training or accelerated by supranormal cytokine status as reported in apparently healthy aged persons suffering low-grade inflammation or in critically ill patients whose muscle mass undergoes proteolysis.
5. The results of the events described are:
- Declining generation of hydrogen sulfide (H2S) from enzymatic sources and in the non-enzymatic reduction of elemental S to H2S.
- The biogenesis of H2S via non-enzymatic reduction is further inhibited in areas where earth’s crust is depleted in elemental sulfur (S8) and sulfate oxyanions.
- Elemental S operates as co-factor of several (apo)enzymes critically involved in the control of oxidative processes.
Combination of protein and sulfur dietary deficiencies constitute a novel clinical entity threatening plant-eating population groups. They have a defective production of Cys, GSH and H2S reductants, explaining persistence of an oxidative burden.
6. The clinical entity increases the risk of developing:
- cardiovascular diseases (CVD) and
- stroke
in plant-eating populations regardless of Framingham criteria and vitamin-B status.
Met molecules supplied by dietary proteins are submitted to transmethylation processes resulting in the release of Hcy which:
- either undergoes Hcy — Met RM pathways or
- is committed to transsulfuration decay.
Impairment of CβS activity, as described in protein malnutrition, entails supranormal accumulation of Hcy in body fluids, stimulation of activity and maintenance of Met homeostasis. The data show that combined protein- and S-deficiencies work in concert to deplete Cys, GSH and H2S from their body reserves, hence impeding these reducing molecules to properly face the oxidative stress imposed by hyperhomocysteinemia.
Although unrecognized up to now, the nutritional disorder is one of the commonest worldwide, reaching top prevalence in populated regions of Southeastern Asia. Increased risk of hyperhomocysteinemia and oxidative stress may also affect individuals suffering from intestinal malabsorption or westernized communities having adopted vegan dietary lifestyles.
Ingenbleek Y. Hyperhomocysteinemia is a biomarker of sulfur-deficiency in human morbidities. Open Clin. Chem. J. 2009 ; 2 : 49-60.
7. The dysfunctional metabolism in transitional cell transformation
A third development is also important and possibly related. The transition a cell goes through in becoming cancerous tends to be driven by changes to the cell’s DNA. But that is not the whole story. Large-scale techniques to the study of metabolic processes going on in cancer cells is being carried out at Oxford, UK in collaboration with Japanese workers. This thread will extend our insight into the metabolome. Otto Warburg, the pioneer in respiration studies, pointed out in the early 1900s that most cancer cells get the energy they need predominantly through a high utilization of glucose with lower respiration (the metabolic process that breaks down glucose to release energy). It helps the cancer cells deal with the low oxygen levels that tend to be present in a tumor. The tissue reverts to a metabolic profile of anaerobiosis. Studies of the genetic basis of cancer and dysfunctional metabolism in cancer cells are complementary. Tomoyoshi Soga’s large lab in Japan has been at the forefront of developing the technology for metabolomics research over the past couple of decades (metabolomics being the ugly-sounding term used to describe research that studies all metabolic processes at once, like genomics is the study of the entire genome).
Their results have led to the idea that some metabolic compounds, or metabolites, when they accumulate in cells, can cause changes to metabolic processes and set cells off on a path towards cancer. The collaborators have published a perspective article in the journal Frontiers in Molecular and Cellular Oncology that proposes fumarate as such an ‘oncometabolite’. Fumarate is a standard compound involved in cellular metabolism. The researchers summarize that shows how accumulation of fumarate when an enzyme goes wrong affects various biological pathways in the cell. It shifts the balance of metabolic processes and disrupts the cell in ways that could favor development of cancer. This is of particular interest because “fumarate” is the intermediate in the TCA cycle that is converted to malate.
Animation of the structure of a section of DNA. The bases lie horizontally between the two spiraling strands. (Photo credit: Wikipedia)
The Keio group is able to label glucose or glutamine, basic biological sources of fuel for cells, and track the pathways cells use to burn up the fuel. As these studies proceed, they could profile the metabolites in a cohort of tumor samples and matched normal tissue. This would produce a dataset of the concentrations of hundreds of different metabolites in each group. Statistical approaches could suggest which metabolic pathways were abnormal. These would then be the subject of experiments targeting the pathways to confirm the relationship between changed metabolism and uncontrolled growth of the cancer cells.
Related articles
Read Full Post »



