When Clinical Application of miRNAs?
Author: Larry H Bernstein, MD, FCAP
Clinical Application of miRNAs Remains a Ways Off
When its time comes, prognostic tests will be first.
Patricia Fitzpatrick Dimond, Ph.D GEN Insight & Intelligence
It’s still early to tell how well microRNAs (miRNAs) will prove clinically useful. Preclinical research findings indicate their central role in controlling cellular pathways.
This novel class of nucleotides, about 20–25 nucleotides in length, affects
gene expression by interacting with
messenger RNAs. But unlike Small Interfering RNA,
siRNAs, miRNAs are encoded in the human genome and function as natural regulators of global gene expression.
Each of the more than 1,500 encoded miRNAs appears to regulate the expression of tens to hundreds of different genes, on-off switches, regulating multiple cellular functions including
- growth and
- proliferation.
miRNAs regulate the translation of genes through
- sequence-specific binding to mRNA.
Depending on the degree of sequence complimentarity, they can inhibit
- the translation and/or degradation of their target mRNAs.
Because of their role in controlling “suites” of genes and, ultimately, pathway function, these molecules have attracted considerable scientific and investor interest in the control of diseases ranging from cardiovascular diseases to cancer.
miRNAs target numerous biomolecules that play a role in carcinogenesis,
- functioning as both tumor promoters or suppressors.
Aberrant expression of miRNAs
- correlates with the development and progression of tumors;
inhibition of their expression can
- modulate the cancer phenotype,
- suggesting their potential as anticancer drug targets.
Further supporting their potential use as drug targets, miRNA expression profiling in a variety of tissue, cell, and disease types has revealed
- a “miRNA signature” specific to those cell types or disease states.
Research
In studying the global changes of the miRNA repertoire along the transitions defining breast cancer progression, the scientists found that
- let-7d, miR-210, and miR-221 were downregulated in the in situ and
- upregulated in the invasive transition, thus
- featuring an expression reversal along the cancer progression path.
- in addition, miRNAs for overall survival and time to metastasis.
Dr. Croce posed that targeted prognostic tests using miRNA will be available within the next two years.
- the problem he suggests is validating the signature in a large enough cohort of patients.
They used deep sequencing, an extremely sensitive approach to the determination of miRNAs because
you count the molecules. Studies have used microarrays and
RT-PCR, and his group used general microarrays and
validated RT-PCR. Their method avoided the possibility of artifacts (by counting). Sequencing permits counts of molecules to provide good data.
John F. McDonald, Ph.D., CSO Ovarian Cancer Institute, and colleagues at the Georgia Institute of Technology
- separately transfected two miRNAs (miR-7 and miR-128) into the ovarian cancer cell line (HEY) and
- then monitored global changes in gene expression levels.
- 20% of the changes in expression patterns of hundreds to thousands of genes
- could be attributed to direct miRNA–mRNA interactions, but
- the majority of the changes were indirect,
involving the downstream consequences of miRNA-mediated changes in regulatory gene expression.
The pathways most significantly affected by miR-7 transfection, are involved with
- cell adhesion and
- developmental networks previously associated with epithelial-mesenchymal transitions and
- processes linked with metastasis.
http://www.genengnews.com/insight-and-intelligenceand153/clinical-application-of-mirnas-remains-a-ways-off/77899650/
MicroRNAs
From Basic Mechanisms to Clinical Application in Cardiovascular Medicine
Christian Weber, Ludwig-Maximilians-Univ and German Centre for Cardiovasc Res, Munich, Germany
MicroRNAs (miRs) are small noncoding RNAs (≈23 nucleotides) that regulate gene expression at a posttranscriptional level by degradation or translational inhibition of target mRNAs. Initially discovered as regulators of development in plants, worms, and fruitflies,
miRs are emerging as
- pivotal modulators of cardiovascular biology and disease in mice and men.
Besides a cell-specific transcription factor profile,
- cell-specific miR-regulated gene expression is integral to cell fate and activation decisions.
Thus, the cell types involved in
- atherosclerosis,
- vascular disease, and
- its myocardial sequelae may be
- differentially regulated by distinct miRs, thereby
- controlling highly complex processes
- smooth muscle cell phenotype and
- inflammatory responses of endothelial cells or macrophages.
The generation of mature miR strands requires several steps of processing of the primary miR gene transcript, including
- cleavage of the terminal loop of miR-precursors by the RNase III enzyme,Dicer, to produce miR duplexes.
Although either strand of the miR duplex can be stably associated with an Argonaute (Ago) family protein,
- preferential loading of a specific strand (ie, the guide strand) onto the miR-induced silencing complex (RISC) is common.
The strand that is not loaded into the RISC (ie, the passenger strand or miR*) is typically degraded.3 Strand selection may be tissue-specific, and an accumulation observed for both strands implies that
- each strand can separately enter the silencing complex.4
Because of the often imperfect complementary binding of the miR seed sequence to the mRNA recognition element,
- an individual miR can affect the expression of hundreds of target mRNAs.
http://atvb.ahajournals.org/content/33/2/168.extract
Life’s Tiniest Architects Pinpointed by Yale Researchers
If a genome is the blueprint for life, then the chief architects are
- tiny slices of genetic material that orchestrate how we are assembled and function.
The study pinpoints the molecular regulators of epigenetics — the process by which unchanging genes along our DNA are switched on and off at precisely right time and place.
“Our genome is like a landscape with lakes, mountains, and rivers, but it is not yet a community or a city full of buildings,” said Haifan Lin, director of the Yale Stem Cell Center and senior author of the study. “What this system does is decide where and when to send out the masons, carpenters, and electricians to build a city or a community.”
In the past 20 years, scientists have discovered that some proteins, called epigenetic factors, traverse the static genome and turn the genes on or off. The staggering number of potential combinations of active and inactive genes explains why a relatively small number of genes can carry out such a wide range of functions.
What guides these epigenetic factors to their target? The answer:
- specialized RNAs called piRNAs.
In the latest study, the Yale team discovered that
- piRNAs guide epigenetic factors to numerous sites throughout the genome of the fruit fly Drosophila, where
- these switches work to turn genes on or off.
The dramatic change in gene expression patterns found illustrated
- piRNAs key role in coordinating biological activity.
“This is the first major mechanism discovered that controls where epigenetic factors —the gene switches — are to be placed in the genome,” Lin said.
Several types of cancers appeared to be
- triggered when the wrong kinds of piRNAs guide epigenetic factors to activate the wrong genes.
Blocking the action of these piRNAs should become a new opportunity to treat cancers, Lin said.
Xiao A. Huang and Hang Yin of Yale are co-lead authors of the paper.
The research was funded by a National Institutes of Health Pioneer Award to Haifan Lin and a grant from Connecticut Stem Cell Research Fund to
Lin and former Yale professor and co-author Michael Snyder, now of Stanford University.

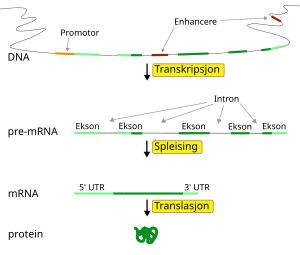
English: A diagram showing at which stages in the DNA-mRNA-protein pathway expression can be controlled. (Photo credit: Wikipedia)

Virus-Encoded microRNAs (Photo credit: AJC1)

English: A Tet-ON doxycycline inducible transgene expression system. (Photo credit: Wikipedia)
Like this:
Like Loading...
Read Full Post »