Understanding of OVO-like proteins (OVOL), which are members of the zinc finger protein family, serve as transcription factors to regulate gene expression in various differentiation processes and are involved in epithelial development and differentiation in a wide variety of organisms. Thus, comparative genomic analysis among three different OVOL genes (OVOL1-3) in vertebrates may shed a light onto this crucial gene for development of molecular diagnostics and targeted therapies.
The Figure is from:
Genomics. Author manuscript; available in PMC 2010 June 29.
Published in final edited form as: Genomics. 2002 September; 80(3): 319–325.
Analysis of mouse and human OVOL2 gene products. (A) The 5′ end sequences of the mouse Ovol2B cDNA and the deduced OVOL2B protein. The “#” symbol indicates the position of an internal methionine previously mistaken as the initiation codon [14]. (B) Deduced amino acid sequences of OVOL2A proteins in mouse (mOvol2A) and human (hOvol2A). The “*” symbol indicates amino acid identity. The four C2H2 zinc fingers are underlined. The predicted NLS sequences are boxed. Sequences common to mouse OVOL2A and OVOL2B start at the brackets in (A) and (B). Human OVOL2B starts at the internal methionine (bold). Shown in bold and italics are positions where our predicted sequence differs from the previously reported sequence [14]. (C) Phylogenetic analysis of OVO proteins. cOvo, C. elegans OVO (GenBank acc. no. AF134806); dOvo, Drosophila OVO (GenBank acc. no. X59772); mOvol1, mouse OVOL1 (GenBank acc. no. AF134804); hOvol1, human OVOL1 (GenBank acc. no. AF016045); mOvol2, mouse OVOL2 (GenBank acc. no.AY090537); hOvol2, human OVOL2 (GenBank acc. no. AK022284); mOvol3, mouse OVOL3 (GenBank acc. no. BF714064); hOvol3, human OVOL3 (GenBank acc. no. AD001527).
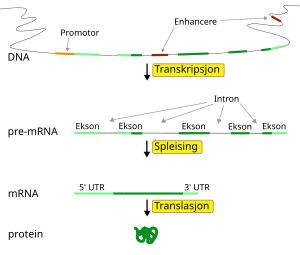
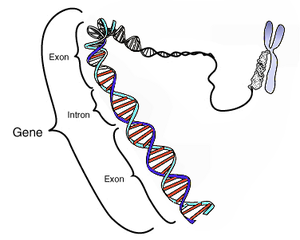
The Ovo gene family encodes evolutionarily conserved proteins contain four DNA-binding C2H2 zinc fingers at the C termini and possess transcriptional regulatory activities in diverse array of organisms from Caenorhabditis elegans, Drosophila, Zebrafish, chick, and mammals. Drosophila ovo, the founding member of the family, acts genetically downstream of Wg (fly Wnt homolog) and DER (fly epidermal growth factor receptor homolog) signaling pathways and is required for epidermal denticle formation and oogenesis.
OVOL proteins are characterized by the presence of hypervariable ID regions.
A. Mouse OVOL1 has ID residues in the first 100 amino acids. B. Mouse OVOL2 possesses ID residues in the first 50 amino acids with a glycine-rich and serine rich region as marked in red color. C. Mouse OVOL3 has ID segments within the N-terminal 100 residues. D. Drosophila OVO is intrinsically disordered with large patches of residue biasness as indicated by the red color. We used DISOPRED2 software [47] for the prediction of ID regions. The horizontal line indicates the ordered/disordered threshold for the default false positive rate of 5%. The ‘filter’ curve represents the outputs from DISOPRED2 and the ‘output’ curve represents the outputs from a linear support vector machine (SVM) classifier (DISOPREDsvm). The outputs from DISOPREDsvm are included to indicate shorter as low confidence predictions of disorder.
doi:10.1371/journal.pone.0039399.g001, http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0039399)
This gene family is identified as mammalian Ovol (Ovo-like) genes, including Ovol1(movo1), Ovol2 (movo2), and Ovol3 (movo3) in mice and OVOL1, OVOL2, andOVOL3 in humans. Ovol1 is the most studied compared to Ovol2 and Ovol3.
Kumar A, Bhandari A, Sinha R, Sardar P, et al. (2012) Molecular Phylogeny of OVOL Genes Illustrates a Conserved C2H2 Zinc Finger Domain Coupled by Hypervariable Unstructured Regions. PLoS ONE 7(6): e39399. doi:10.1371/journal.pone.0039399
http://www.plosone.org/article/info:doi/10.1371/journal.pone.0039399
Phylogenetic history of OVOL proteins using the Bayesian method. A. Full-length OVOL proteins. B. Selected region of OVOL proteins.
Posterior probabilities scores are depicted by various color balls. The placozoan OVOL protein (e_gw1.4.509.1) was used as the outgroup in this phylogenetic tree. Red x indicates sequence position, which did not accord with species phylogeny. BFL: B. floridae (lancelet), SPU: S. purpuratus (sea urchin), NVE: N. vectensis (sea anemone), HRO: H. robusta (annelids), LGI: L. gigantean (molluscs) and TAD: T. adhaerens(placozoan). Trees in figures 7A and 7B are generated using the MrBayes 3.2 [53] from alignments supplied in supplementary Files S1 and S2, respectively. (Figure 7 of doi:10.1371/journal.pone.0039399.g007, http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0039399)
“Fine scale analysis of gene expression in Drosophila melanogaster gonads reveals Programmed cell death 4 promotes the differentiation of female germline stem cells” Amy C Cash and Justen Andrews* BMC Developmental Biology 2012, 12:4 doi:10.1186/1471-213X-12-4
”Regulatory and functional interactions between ovarian tumor and ovo during Drosophila oogenesis. “ Shannon Hinson, Janette Pettus, Rod N Nagoshi Mechanisms of Development Volume 88, Issue 1, 1 October 1999, Pages 3–14 http://www.sciencedirect.com/science/article/pii/S0925477399001677
“Molecular phylogeny of OVOL genes illustrates a conserved C2H2 zinc finger domain coupled by hypervariable unstructured regions.” Kumar A, Bhandari A, Sinha R, Sardar P, Sushma M, Goyal P, Goswami C, Grapputo A. PLoS One. 2012;7(6):e39399. doi: 10.1371/journal.pone.0039399. Epub 2012 Jun 21.
“Gapped BLAST and PSI-BLAST: a new generation of protein database search programs.” Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ.
Nucleic Acids Res. 1997 Sep 1; 25(17):3389-402.
Ovol1:
Chromosomal localization of OVOL1 gene from selected vertebrates, flanked by a set of conserved marker genes. SIPA1: signal-induced proliferation-associated 1; RELA: v-rel reticuloendotheliosis viral oncogene homolog A (avian); KAT5: K (lysine) acetyltransferase; SNX32: sorting nexin 32; MUS81: MUS81 endonuclease homolog (S. cerevisiae); BANF1: barrier to autointegration factor 1; EXOC6B: exocyst complex component 6B; DYSF: dysferlin, limb girdle muscular dystrophy 2B; COL4A5: collagen, type IV, alpha 5; DAK: dihydroxyacetone kinase 2 S. cerevisiae homolog. doi:10.1371/journal.pone.0039399.g003, http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0039399)
Drosophila ovo/svb (dovo) is required for epidermal cuticle/denticle differentiation and is genetically downstream of the wg signaling pathway, so does a mouse homolog of dovo, movo1. Also, Li group showed that movo1 promoter is activated by the lymphoid enhancer factor 1 (LEF1)/β-catenin complex, a transducer of wnt signaling. Simply these data showed movo1 is a developmental target of wnt signaling during hair morphogenesis in mice, and there is a conserved regulatory pathway at wg/wnt-ovolink in epidermal appendage. human OVOL1 has been identified as a gene that is responsive to TGF-β1/BMP7 treatment via a Smad4-dependent pathway (Kowanetz et al., 2004).
Ovo1 li“Characterization of a human homolog (OVOL1) of the Drosophila ovo gene, which maps to chromosome 11q13.” Chidambaram A, Allikmets R, Chandrasekarappa S, Guru SC, Modi W, Gerrard B, Dean M. Mamm Genome. 1997 Dec;8(12):950-1.nks Wnt signaling with N-cadherin localization during neural crest migrationDevelopment 2010 137 (12) 1981-1990.
“Id2 and Id3 define the potency of cell proliferation and differentiation responses to transforming growth factor beta and bone morphogenetic protein.” Marcin Kowanetz,
Ulrich Valcourt, Rosita Bergström‡,Carl-Henrik Heldin and Aristidis Moustakas*
. Mol. Cell. Biol., 24 (2004), pp. 4241–4254 http://mcb.asm.org/content/24/10/4241
“The LEF1/β-catenin complex activates movo1, a mouse homolog of Drosophila ovo required for epidermal appendage differentiation” Baoan Li, Douglas R. Mackay, Qian Dai, Tony W. H. Li, Mahalakshmi Nair, Magid Fallahi, Christopher P. Schonbaum, Judith Fantes, Anthony P. Mahowald, Marian L. Waterman,Elaine Fuchs, and Xing DaiPNAS vol. 99 no. 9 Baoan Li, 6064–6069. http://www.pnas.org/content/99/9/6064.abstract?ijkey=a7d2985635ef09ca63982ae4397ef325aa46252b&keytype2=tf_ipsecsha
“Expression of murine novel zinc finger proteins highly homologous to Drosophila ovo gene product in testis.” Masu Y, Ikeda S, Okuda-Ashitaka E, Sato E, Ito S. FEBS Lett. 1998 Jan 16;421(3):224-8.
“The ovo gene required for cuticle formation and oogenesis in flies is involved in hair formation and spermatogenesis in mice” Xing Dai, Christopher Schonbaum, Linda Degenstein, Wenyu Bai,Anthony Mahowald, and Elaine Fuchs. (1998) Genes Dev. 12, 3452–3463 http://www.ncbi.nlm.nih.gov/pubmed/9808631
OvoL2 at the Junction of Decisions:
Brain development is fascinating and complex since cranial neurulation is an integral component of brain morphogenesis and there are factors present outside of the neuroepithelium can also affect the morphogenesis of the cranial neural tube. Previous studies revealed Ovol2 expression in brain, testis, and epithelial tissues such as skin and intestine of adult mice (Li et al., 2002a).
OVOL2 orthologs identified in vertebrates by comparing chromosomal localization.
RRBP1: ribosome binding protein 1 homolog; BANF2: barrier to autointegration factor 2; SNX5: sorting nexin 5; CSRP2BP: CSRP2 binding protein; SEC23B: protein transport protein Sec23B; POLR3F: polymerase (RNA) III (DNA directed) polypeptide F; RBBP9: Retinoblastoma-binding protein 9; DTD1: D-tyrosyl-tRNA deacylase 1. doi:10.1371/journal.pone.0039399.g004, http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0039399)
Neural/non-neural cell fate decisions is carried by bone morphogenetic protein (BMP) signaling, which inhibits precocious neural differentiation and allows for proper differentiation of mesoderm, endoderm, and epidermis, during early embryonic development. There are many unknown in this mechanism yet the expression of Ovol2, which encodes an evolutionarily conserved zinc finger transcription factor, is down-regulated during neural differentiation of mouse embryonic stem cells since null Ovol2 in embryonic stem cells facilitates neural conversion and inhibits mesendodermal differentiation, whereas Ovol2 overexpression gives rise to the opposite phenotype. Furthermore, the studies also prove BMP4 and ovo2 interacted to rescue these changes. If BMP4 is provided, Ovol2 knockdown partially rescues the neural inhibition. Mechanism studies show the regulation pattern between these BMP and Ovol2. BMP4 directly regulates Ovol2 expression through the binding of Smad1/5/8 to the second intron of the Ovol2 gene. Thus, Ovol2 acts downstream of BMP pathway. In addition, in vivo chick studies presented that when Ovol2 is ectopically expressed the prospective neural plate represses the expression of the definitive neural plate marker cSox2In the chick embryo. Also, lack of Ovol2 prevented increase BMP4 expression. During early germ layer development there is an important comment between neuroectoderm and mesendoderm provided by Ovol2.
In addition, Ovol2 acts in downstream of key developmental signaling pathways including Wg/Wnt and BMP/TGF-β. Based on findings from chromatin immunoprecipitation, luciferase reporter, and functional rescue assays, Wells group demonstrated that Ovol2 directly represses two critical downstream targets, c-Mycand Notch1. Hence, this action suppresses keratinocyte transient proliferation and terminal differentiation. Like a twilight zone to choose when to proliferate and when to resist differentiation.
“Ovol
“Ovol2, a Mammalian Homolog of Drosophila ovo: Gene Structure, Chromosomal Mapping, and Aberrant Expression in Blind-Sterile Mice.” Baoan Li, Qian Dai, Ling Li, Mahalakshmi Nair, Douglas R. Mackay, Xing DaiGenomics Volume 80, Issue 3, September 2002, Pages 319–325.
Wells J, Lee B, Cai AQ, Karapetyan A, Lee WJ, Rugg E, Sinha S, Nie Q, Dai X.
J Biol Chem. 2009 Oct 16;284(42):29125-35. doi: 10.1074/jbc.M109.008847. Epub 2009 Aug 21.
“The zinc finger transcription factor Ovol2 acts downstream of the bone morphogenetic protein pathway to regulate the cell fate decision between neuroectoderm and mesendoderm.” Zhang T, Zhu Q, Xie Z, Chen Y, Qiao Y, Li L, Jing N. J Biol Chem. 2013 Mar 1;288(9):6166-77. doi: 10.1074/jbc.M112.418376. Epub 2013 Jan 14.
“The mouse Ovol2 gene is required for cranial neural tube development” Douglas R. Mackaya, Ming Hua, Baoan Lia, Catherine Rhéaumea, Xing Dai.l Developmental Biology Volume 291, Issue 1, 1 March 2006, Pages 38–52.
Ovol3
While tracing the OVOL genes, we identified a third OVOL gene, OVOL3, in a wide array of mammals including humans (chromosome 19), chimpanzees (chromosome 19), mice (chromosome 7), rats (chromosome 1), cows (chromosome 18), pigs (chromosome 6), and opossums (chromosome 4) with a conserved synteny. The conserved synteny comprises an octet of genes, LIN37-PRODH2-KIRREL2-APLP11-NKF3ID-LPFN3-SDHAF1-CLIF3,on one side and POLR2L-CAPSN1-COX7A1 on the other side of OVOL3 in a region of about 400 kb (Figure 5 of http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0039399).
Synteny analysis of OVOL3 genes illustrates the loss of OVOL3a after duplication event and maintenance of paralogous OVOL3b in fishes.
LIN37: lin-37 homolog (C. elegans); PRODH2: proline dehydrogenase (oxidase) 2; KIRREL2: kin of IRRE like 2 (Drosophila); APLP1: amyloid beta (A4) precursor-like protein 1; NFKBID: nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, delta; LRFN3: leucine rich repeat and fibronectin type III domain containing 3; SDHAF1: succinate dehydrogenase complex assembly factor 1; CLIP3: CAP-GLY domain containing linker protein 3; POLR2I: polymerase (RNA) II (DNA directed) polypeptide I, 14.5 kDa; CAPNS1: calpain, small subunit 1; COX7A1: cytochrome c oxidase subunit VIIa polypeptide 1 (muscle); DMPK: dystrophia myotonica-protein kinase; HLCS: holocarboxylase synthetase; AMOT: angiomotin; REXO2: REX2 RNA exonuclease 2 homolog (S. cerevisiae). (Reference from doi:10.1371/journal.pone.0039399.g005 , http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0039399)
There are not many studies on Ovol 3but my interest in this region lies on Chromosome 19 since “it has the highest gene density of all human chromosomes and the large clustered gene families, corresponding high G + C content, CpG islands and density of repetitive DNA indicate a chromosome rich in biological and evolutionary significance.”
“The DNA sequence and biology of human chromosome 19.” Grimwood J, et al. Nature. 2004 Apr 1;428(6982):529-35.