Renal tumor macrophages linked to recurrence are identified using single-cell protein activity analysis
Curator and Reporter: Dr. Premalata Pati, Ph.D., Postdoc
When malignancy returns after a period of remission, it is called a cancer recurrence. After the initial or primary cancer has been treated, this can happen weeks, months, or even years later. The possibility of recurrence is determined by the type of primary cancer. Because small patches of cancer cells might stay in the body after treatment, cancer might reoccur. These cells may multiply and develop large enough to cause symptoms or cause cancer over time. The type of cancer determines when and where cancer recurs. Some malignancies have a predictable recurrence pattern.
Even if primary cancer recurs in a different place of the body, recurrent cancer is designated for the area where it first appeared. If breast cancer recurs distantly in the liver, for example, it is still referred to as breast cancer rather than liver cancer. It’s referred to as metastatic breast cancer by doctors. Despite treatment, many people with kidney cancer eventually develop cancer recurrence and incurable metastatic illness.
The most frequent type of kidney cancer is Renal Cell Carcinoma (RCC). RCC is responsible for over 90% of all kidney malignancies. The appearance of cancer cells when viewed under a microscope helps to recognize the various forms of RCC. Knowing the RCC subtype can help the doctor assess if the cancer is caused by an inherited genetic condition and help to choose the best treatment option. The three most prevalent RCC subtypes are as follows:
- Clear cell RCC
- Papillary RCC
- Chromophobe RCC
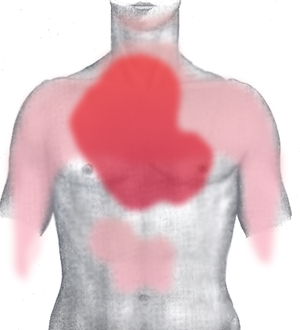
Clear Cell RCC (ccRCC) is the most prevalent subtype of RCC. The cells are clear or pale in appearance and are referred to as the clear cell or conventional RCC. Around 70% of people with renal cell cancer have ccRCC. The rate of growth of these cells might be sluggish or rapid. According to the American Society of Clinical Oncology (ASCO), clear cell RCC responds favorably to treatments like immunotherapy and treatments that target specific proteins or genes.

Image Source: Moffitt Cancer Center https://moffitt.org/for-healthcare-professionals/clinical-perspectives/clinical-perspectives-story-archive/moffitt-study-seeks-to-inform-improved-treatment-strategies-for-clear-cell-renal-cell-carcinoma/
Researchers at Columbia University’s Vagelos College of Physicians and Surgeons have developed a novel method for identifying which patients are most likely to have cancer relapse following surgery.
The study
Their findings are detailed in a study published in the journal Cell entitled, “Single-Cell Protein Activity Analysis Identifies Recurrence-Associated Renal Tumor Macrophages.” The researchers show that the presence of a previously unknown type of immune cell in kidney tumors can predict who will have cancer recurrence.

Image Source: https://marlin-prod.literatumonline.com/cms/attachment/a1208337-6b5b-4ef3-996d-227eb2536553/fx1_lrg.jpg
According to co-senior author Charles Drake, MD, PhD, adjunct professor of medicine at Columbia University Vagelos College of Physicians and Surgeons and the Herbert Irving Comprehensive Cancer Center,
the findings imply that the existence of these cells could be used to identify individuals at high risk of disease recurrence following surgery who may be candidates for more aggressive therapy.
As Aleksandar Obradovic, an MD/PhD student at Columbia University Vagelos College of Physicians and Surgeons and the study’s co-first author, put it,
it’s like looking down over Manhattan and seeing that enormous numbers of people from all over travel into the city every morning. We need deeper details to understand how these different commuters engage with Manhattan residents: who are they, what do they enjoy, where do they go, and what are they doing?
To learn more about the immune cells that invade kidney cancers, the researchers employed single-cell RNA sequencing. Obradovic remarked,
In many investigations, single-cell RNA sequencing misses up to 90% of gene activity, a phenomenon known as gene dropout.
The researchers next tackled gene dropout by designing a prediction algorithm that can identify which genes are active based on the expression of other genes in the same family. “Even when a lot of data is absent owing to dropout, we have enough evidence to estimate the activity of the upstream regulator gene,” Obradovic explained. “It’s like when playing ‘Wheel of Fortune,’ because I can generally figure out what’s on the board even if most of the letters are missing.”
The meta-VIPER algorithm is based on the VIPER algorithm, which was developed in Andrea Califano’s group. Califano is the head of Herbert Irving Comprehensive Cancer Center’s JP Sulzberger Columbia Genome Center and the Clyde and Helen Wu professor of chemistry and systems biology. The researchers believe that by including meta-VIPER, they will be able to reliably detect the activity of 70% to 80% of all regulatory genes in each cell, eliminating cell-to-cell dropout.
Using these two methods, the researchers were able to examine 200,000 tumor cells and normal cells in surrounding tissues from eleven patients with ccRCC who underwent surgery at Columbia’s urology department.
The researchers discovered a unique subpopulation of immune cells that can only be found in tumors and is linked to disease relapse after initial treatment. The top genes that control the activity of these immune cells were discovered through the VIPER analysis. This “signature” was validated in the second set of patient data obtained through a collaboration with Vanderbilt University researchers; in this second set of over 150 patients, the signature strongly predicted recurrence.
These findings raise the intriguing possibility that these macrophages are not only markers of more risky disease, but may also be responsible for the disease’s recurrence and progression,” Obradovic said, adding that targeting these cells could improve clinical outcomes
Drake said,
Our research shows that when the two techniques are combined, they are extremely effective at characterizing cells within a tumor and in surrounding tissues, and they should have a wide range of applications, even beyond cancer research.
Main Source
Single-cell protein activity analysis identifies recurrence-associated renal tumor macrophages
https://www.cell.com/cell/fulltext/S0092-8674(21)00573-0
Other Related Articles published in this Open Access Online Scientific Journal include the following:
Machine Learning (ML) in cancer prognosis prediction helps the researcher to identify multiple known as well as candidate cancer diver genes
Curator and Reporter: Dr. Premalata Pati, Ph.D., Postdoc
Renal (Kidney) Cancer: Connections in Metabolism at Krebs cycle and Histone Modulation
Curator: Demet Sag, PhD, CRA, GCP
Artificial Intelligence: Genomics & Cancer
https://pharmaceuticalintelligence.com/ai-in-genomics-cancer/
Bioinformatic Tools for Cancer Mutational Analysis: COSMIC and Beyond
Curator: Stephen J. Williams, Ph.D.
Deep-learning AI algorithm shines new light on mutations in once obscure areas of the genome
Reporter: Aviva Lev-Ari, PhD, RN
Premalata Pati, PhD, PostDoc in Biological Sciences, Medical Text Analysis with Machine Learning