
typical changes in CK-MB and cardiac troponin in Acute Myocardial Infarction (Photo credit: Wikipedia)
Reporter and curator:
Larry H Bernstein, MD, FCAP
This posting is a followup on two previous posts covering the design and handling of HIT to improve healthcare outcomes as well as lower costs from better workflow and diagnostics, which is self-correcting over time.
The first example is a non technology method designed by Lee Goldman (Goldman Algorithm) that was later implemented at Cook County Hospital in Chicago with great success. It has been known that there is over triage of patients to intensive care beds, adding to the costs of medical care. If the differentiation between acute myocardial infarction and other causes of chest pain could be made more accurate, the quantity of scare resources used on unnecessary admissions could be reduced. The Goldman algorithm was introduced in 1982 during a training phase at Yale-New Haven Hospital based on 482 patients, and later validated at the BWH (in Boston) on 468 patients.They demonstrated improvement in sensitivity as well as specificity (67% to 77%), and positive predictive value (34% to 42%). They modified the computer derived algorithm in 1988 to achieve better results in triage of patients to the ICU of patients with chest pain based on a study group of 1379 patients. The process was tested prospectively on 4770 patients at two university and 4 community hospitals. The specificity improved by 74% vs 71% in recognizing absence of AMI by the algorithm vs physician judgement. The sensitivity was not different for admission (88%). Decisions based solely on the protocol would have decreased admissions of patients without AMI by 11.5% without adverse effects. The study was repeated by Qamar et al. with equal success.
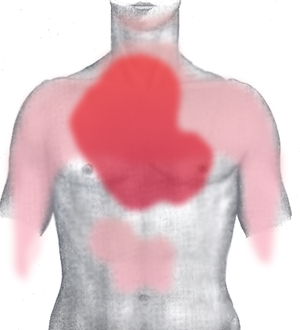
Pain in acute myocardial infarction (front) (Photo credit: Wikipedia)

An ECG showing pardee waves indicating acute myocardial infarction in the inferior leads II, III and aVF with reciprocal changes in the anterolateral leads. (Photo credit: Wikipedia)
Acute myocardial infarction with coagulative necrosis (4) (Photo credit: Wikipedia)
Goldman L, Cook EF, Brand DA, Lee TH, Rouan GW, Weisberg MC, et al. A computer protocol to predict myocardial infarction in emergency department patients with chest pain. N Engl J Med. 1988;318:797-803.
A Qamar, C McPherson, J Babb, L Bernstein, M Werdmann, D Yasick, S Zarich. The Goldman algorithm revisited: prospective evaluation of a computer-derived algorithm versus unaided physician judgment in suspected acute myocardial infarction. Am Heart J 1999; 138(4 Pt 1):705-709. ICID: 825629
The usual accepted method for determining the decision value of a predictive variable is the Receiver Operator Characteristic Curve, which requires a mapping of each value of the variable against the percent with disease on the Y-axis. This requires a review of every case entered into the study. The ROC curve is done to validate a study to classify data on leukemia markers for research purposes as shown by Jay Magidson in his demonstation of Correlated Component Regression (2012)(Statistical Innovations, Inc.) The test for the contribution of each predictor is measured by Akaike Information Criteria and Bayes Information Criteria, which have proved to be critically essential tests over the last 20 years.
I go back 20 years and revisit the application of these principles in clinical diagnostics, but the ROC was introduced to medicine in radiology earlier. A full rendering of this matter can be found in the following:
R A Rudolph, L H Bernstein, J Babb. Information induction for predicting acute myocardial infarction.Clin Chem 1988; 34(10):2031-2038. ICID: 825568.
Rypka EW. Methods to evaluate and develop the decision process in the selection of tests. Clin Lab Med 1992; 12:355
Rypka EW. Syndromic Classification: A process for amplifying information using S-Clustering. Nutrition 1996;12(11/12):827-9.
Christianson R. Foundations of inductive reasoning. 1964. Entropy Publications. Lincoln, MA.
Inability to classify information is a major problem in deriving and validating hypotheses from PRIMARY data sets necessary to establish a measure of outcome effectiveness. When using quantitative data, decision limits have to be determined that best distinguish the populations investigated. We are concerned with accurate assignment into uniquely verifiable groups by information in test relationships. Uncertainty in assigning to a supervisory classification can only be relieved by providing suffiuciuent data.
A method for examining the endogenous information in the data is used to determine decision points. The reference or null set is defined as a class having no information. When information is present in the data, the entropy (uncertainty in the data set) is reduced by the amount of information provided. This is measureable and may be referred to as the Kullback-Liebler distance, which was extended by Akaike to include statistical theory. An approach is devised using EW Rypka’s S-Clustering has been created to find optimal decision values that separate the groups being classified. Further, it is possible to obtain PRIMARY data on-line and continually creating primary classifications (learning matrices). From the primary classifications test-minimized sets of features are determined with optimal useful and sufficient information for accurately distinguishing elements (patients). Primary classifications can be continually created from PRIMARY data. More recent and complex work in classifying hematology data using a 30,000 patient data set and 16 variables to identify the anemias, moderate SIRS, sepsis, lymphocytic and platelet disorders has been published and recently presented. Another classification for malnutrition and stress hypermetabolism is now validated and in press in the journal Nutrition (2012), Elsevier.
G David, LH Bernstein, RR Coifman. Generating Evidence Based Interpretation of Hematology Screens via Anomaly Characterization. Open Clinical Chemistry Journal 2011; 4 (1):10-16. 1874-2416/11 2011 Bentham Open. ICID: 939928
G David; LH Bernstein; RR Coifman. The Automated Malnutrition Assessment. Accepted 29 April 2012.
http://www.nutritionjrnl.com. Nutrition (2012), doi:10.1016/j.nut.2012.04.017.
Keywords: Network Algorithm; unsupervised classification; malnutrition screening; protein energy malnutrition (PEM); malnutrition risk; characteristic metric; characteristic profile; data characterization; non-linear differential diagnosis
Summary: We propose an automated nutritional assessment (ANA) algorithm that provides a method for malnutrition risk prediction with high accuracy and reliability. The problem of rapidly identifying risk and severity of malnutrition is crucial for minimizing medical and surgical complications. We characterized for each patient a unique profile and mapped similar patients into a classification. We also found that the laboratory parameters were sufficient for the automated risk prediction.
We here propose a simple, workable algorithm that provides assistance for interpreting any set of data from the screen of a blood analysis with high accuracy, reliability, and inter-operability with an electronic medical record. This has been made possible at least recently as a result of advances in mathematics, low computational costs, and rapid transmission of the necessary data for computation. In this example, acute myocardial infarction (AMI) is classified using isoenzyme CKMB activity, total LD, and isoenzyme LD-1, and repeated studies have shown the high power of laboratory features for diagnosis of AMI, especially with NSTEMI. A later study includes the scale values for chest pain and for ECG changes to create the model.
LH Bernstein, A Qamar, C McPherson, S Zarich. Evaluating a new graphical ordinal logit method (GOLDminer) in the diagnosis of myocardial infarction utilizing clinical features and laboratory data. Yale J Biol Med 1999; 72(4):259-268. ICID: 825617
The quantitative measure of information, Shannon entropy treats data as a message transmission. We are interested in classifying data with near errorless discrimination. The method assigns upper limits of normal to tests computed from Rudolph’s maximum entropy definitions of group-based normal reference. Using the Bernoulli trial to determine maximum entropy reference, we determine from the entropy in the data a probability of a positive result that is the same for each test and conditionally independent of other results by setting the binary decision level for each test. The entropy of the discrete distribution is calculated from the probabilities of the distribution. The probability distribution of the binary patterns is not flat and the entropy decreases when there is information in the data. The decrease in entropy is the Kullback-Liebler distance.
The basic principle of separatory clustering is extracting features from endogenous data that amplify or maximize structural information into disjointed or separable classes. This differs from other methods because it finds in a database a theoretic – or more – number of variables with required VARIETY that map closest to an ideal, theoretic, or structural information standard. Scaling allows using variables with different numbers of message choices (number bases) in the same matrix, binary, ternary, etc (representing yes-no; small-modest, large, largest). The ideal number of class is defined by x^n. In viewing a variable value we think of it as low, normal, high, high high, etc. A system works with related parts in harmony. This frame of reference improves the applicability of S-clustering. By definition, a unit of information is log.r r = 1.
The method of creating a syndromic classification to control variety in the system also performs a semantic function by attributing a term to a Port Royal Class. If any of the attributes are removed, the meaning of the class is made meaningless. Any significant overlap between the groups would be improved by adding requisite variety. S-clustering is an objective and most desirable way to find the shortest route to diagnosis, and is an objective way of determining practice parameters.
Multiple Test Binary Decision Patterns where CK-MB = 18 u/l, LD-1 = 36 u/l, %LD1 = 32 u/l.
No. Pattern Freq P1 Self information Weighted information
0 000 26 0.1831 2.4493 0.4485
1 001 3 0.0211 5.5648 0.1176
2 010 4 0.0282 5.1497 0.1451
3 011 2 0.0282 6.1497 0.0866
4 100 6 0.0423 4.5648 0.1929
6 110 8 0.0563 4.1497 0.2338
7 111 93 0.6549 0.6106 0.3999
Entropy: sum of weighted information (average) 1.6243 bits
The effective information values are the least-error points. Non AMI patients exhibit patterns 0, 1, 2, 3, and 4: AMI patients are 6 and 7. There is 1 fp 4, and 1 fn 6. The error rate is 1.4%.
Summary:
A major problem in using quantitative data is lack of a justifiable definition of reference (normal). Our information model consists of a population group, a set of attributes derived from observations, and basic definitions using Shannon’s information measure entropy. In this model, the population set and its values for its variables are considered to be the only information available. The finding of a flat distribution with the Bernoulli test defines the reference population that has no information. The complementary syndromic group, treated in the same way, produces a distribution that is not flat and has a less than maximum information uncertainty.
The vector of probabilities – (1/2), (1/2), …(1/2), can be related to the path calculated from the Rypka-Fletcher equation, which
Ct = 1 – 2^-k/1 -2^-n
determines the theoretical maximum comprehension from the test of n attributes. We constructed a ROC curve from theoriginal IRIS data of R Fisher from four measurements of leaf and petal with a result obtained using information-based induction principles to determine discriminant points without the classification that had to be used for the discriminant analysis. The principle of maximum entropy, as formu;ated by Jaynes and Tribus proposes that for problems of statistical inference – which as defined, are problems of induction – the probabilities should be assigned so that the entropy function is maximized. Good proposed that maximum entropy be used to define the null hypothesis and Rudolph proposed that medical reference be defined as at maximum entropy.
Rudolph RA. A general purpose information processing automation: generating Port Royal Classes with probabilistic information. Intl Proc Soc Gen systems Res 1985;2:624-30.
Jaynes ET. Information theory and statistical mechanics. Phys Rev 1956;106:620-30.
Tribus M. Where do we stand after 30 years of maximum entropy? In: Levine RD, Tribus M, eds. The maximum entropy formalism. Cambridge, Ma: MIT Press, 1978.
Good IJ. Maximum entropy for hypothesis formulation, especially for multidimensional contingency tables. Ann Math Stat 1963;34:911-34.
The most important reason for using as many tests as is practicable is derived from the prominent role of redundancy in transmitting information (Noisy Channel Theorem). The proof of this theorem does not tell how to accomplish nearly errorless discrimination, but redundancy is essential.
In conclusion, we have been using the effective information (derived from Kullback-Liebler distance) provided by more than one test to determine normal reference and locate decision values. Syndromes and patterns that are extracted are empirically verifiable.
Related articles
Read Full Post »




