Endothelial Function and Cardiovascular Disease
Pathologist and Author: Larry H Bernstein, MD, FCAP
This discussion is a continuation of a series on Nitric Oxide, vascular relaxation, vascular integrity, and systemic organ dysfunctions related to inflammatory and circulatory disorders. In some of these, the relationships are more clear than others, and in other cases the vascular disorders are aligned with serious metabolic disturbances. This article, in particular centers on the regulation of NO production, NO synthase, and elaborates more on the assymetrical dimethylarginine (ADMA) inhibition brought up in a previous comment, and cardiovascular disease, including:
- Acute myocardial infarction
- Congestive heart failure
- Chronic renal disease and dialysis
- Hypertension
Recall, though, that in SIRS leading to septic shock, that there is a difference between the pulmonary circulation, the systemic circulation and the portal circulation in these events. The comment calls attention to:
Böger RH. Asymmetric dimethylarginine, an endogenous inhibitor of nitric oxide synthase, explains the ‘L-arginine paradox’ and acts as a novel cardiovascular risk factor. J Nutr 2004; 134: 2842S–7S.
This observer points out that ADMA inhibits vascular NO production at concentrations found in pathophysiological conditions (i.e., 3–15 μmol/l); ADMA also causes local vasoconstriction when it is infused intra-arterially. ADMA is increased in the plasma of humans with hypercholesterolemia, atherosclerosis, hypertension, chronic renal failure, and chronic heart failure.
Increased ADMA levels are associated with reduced NO synthesis as assessed by impaired endothelium-dependent vasodilation. We’ll go into that more with respect to therapeutic targets – including exercise, sauna, and possibly diet, as well as medical drugs.
It is remarkable how far we have come since the epic discovery of 17th century physician, William Harvey, by observing the action of the heart in small animals and fishes, proved that heart receives and expels blood during each cycle, and argued for the circulation in man. This was a huge lead into renaissance medicine. What would he think now?
Key Words: eNOS, NO, endothelin, ROS, oxidative stress, blood flow, vascular resistance, cardiovascular disease, chronic renal disease, hypertension, diabetes, atherosclerosis, MI, exercise, nutrition, traditional chinese medicine, statistical modeling for targeted therapy.
Endothelial Function
The endothelium plays a crucial role in the maintenance of vascular tone and structure by means of eNOS, producing the endothelium-derived vasoactive mediator nitric oxide (NO), an endogenous messenger molecule formed in healthy vascular endothelium from the amino acid precursor L-arginine. Nitric oxide synthases (NOS) are the enzymes responsible for nitric oxide (NO) generation. The generation and actions of NO under physiological and pathophysiological conditions are exquisitely regulated and extend to almost every cell type and function within the circulation. While the molecule mediates many physiological functions, an excessive presence of NO is toxic to cells.
The enzyme NOS, constitutively or inductively, catalyses the production of NO in several biological systems. NO is derived not only from NOS isoforms but also from NOS-independent sources. In mammals, to date, three distinct NOS isoforms have been identified:
- neuronal NOS (nNOS),
- inducible NOS (iNOS), and
- endothelial NOS (eNOS).
The molecular structure, enzymology and pharmacology of these enzymes have been well defined, and reveal critical roles for the NOS system in a variety of important physiological processes. The role of NO and NOS in regulating vascular physiology, through neuro-hormonal, renal and other non-vascular pathways, as well as direct effects on arterial smooth muscle, appear to be more intricate than was originally thought.
Vallance et al. described the presence of asymmetric dimethylarginine (ADMA) as an endogenous inhibitor of eNOS in 1992. Since then, the role of this molecule in the regulation of eNOS has attracted increasing attention.
Endothelins are 21-amino acid peptides, which are active in almost all tissues in the body. They are potent vasoconstrictors, mediators of cardiac, renal, endocrine and immune functions and play a role in bronchoconstriction, neurotransmitter regulation, activation of inflammatory cells, cell proliferation and differentiation.
Endothelins were first characterised by Yanagisawa et al. (1988). The three known endothelins ET-1, -2 and -3 are structurally similar to sarafotoxins from snake venoms. ET-1 is the major isoform generated in blood vessels and appears to be the isoform of most importance in the cardiovascular system with a major role in the maintenance of vascular tone.
The systemic vascular response to hypoxia is vasodilation. However, reports suggest that the potent vasoconstrictor endothelin-1 (ET-1) is released from the vasculature during hypoxia. ET-1 is reported to augment superoxide anion generation and may counteract nitric oxide (NO) vasodilation. Moreover, ET-1 was proposed to contribute to increased vascular resistance in heart failure by increasing the production of asymmetric dimethylarginine (ADMA).
A study investigated the role of ET-1, the NO pathway, the potassium channels and radical oxygen species in hypoxia-induced vasodilation of large coronary arteries and found NO contributes to hypoxic vasodilation, probably through K channel opening, which is reversed by addition of ET-1 and enhanced by endothelin receptor antagonism. These latter findings suggest that endothelin receptor activation counteracts hypoxic vasodilation.
Endothelial dysfunction
Patients with Raynaud’s Phemonenon had abnormal vasoconstrictor responses to cold pressor tests (CPT) that were similar in primary and secondary RP. There were no differences in median flow-mediated and nitroglycerin mediated dilation or CPT of the brachial artery in the 2 populations. Patients with secondary RP were characterized by abnormalities in microvascular responses to reactive hyperemia, with a reduction in area under the curve adjusted for baseline perfusion, but not in time to peak response or peak perfusion ratio.
Plasma ET-1, ADMA, VCAM-1, and MCP-1 levels were significantly elevated in secondary RP compared with primary RP. There was a significant negative correlation between ET-1 and ADMA values and measures of microvascular perfusion but not macrovascular endothelial function. Secondary RP is characterized by elevations in plasma ET-1 and ADMA levels that may contribute to alterations in cutaneous microvascular function.
ADMA inhibits vascular NO production within the concentration range found in patients with vascular disease. ADMA also causes local vasoconstriction when infused intra-arterially, and increases systemic vascular resistance and impairs renal function when infused systemically. Several recent studies have supplied evidence to support a pathophysiological role of ADMA in the pathogenesis of vascular dysfunction and cardiovascular disease. High ADMA levels were found to be associated with carotid artery intima-media-thickness in a study with 116 clinically healthy human subjects. Taking this observation further, another study performed with hemodialysis patients reported that ADMA prospectively predicted the progression of intimal thickening during one year of follow-up.
In a nested, case-control study involving 150 middle-aged, non-smoking men, high ADMA levels were associated with a 3.9-fold elevated risk for acute coronary events. Clinical and experimental evidence suggests elevation of ADMA can cause a relative L-arginine deficiency, even in the presence of “normal” L-arginine levels. As ADMA is a competitive inhibitor of eNOS, its inhibitory action can be overcome by increasing the concentration of the substrate, L-arginine. Elevated ADMA concentration is one possible explanation for endothelial dysfunction and decreased NO production in these diseases.
Metabolic Regulation of L-arginine and NO Synthesis
Methylation of arginine residues within proteins or polypeptides occurs through N-methyltransferases, which utilize S-adenosylmethionine as a methyl donor. After proteolysis of these proteins or polypeptides, free ADMA is present in the cytoplasm. ADMA can also be detected in circulating blood plasma. ADMA acts as an inhibitor of eNOS by competing with the substrate of this enzyme, L-arginine. The ensuing reduction in nitric oxide synthesis causes vascular endothelial dysfunction and, subsequently, atherosclerosis. ADMA is eliminated from the body via urinary excretion and via metabolism by the enzyme DDAH to citrulline and dimethylamine.
Supplementation with L-arginine in animals with experimentally-induced vascular dysfunction atherosclerosis improves endothelium-dependent vasodilation. Moreover, L-arginine supplementation results in enhanced endothelium-dependent inhibition of platelet aggregation, inhibition of monocyte adhesion, and reduced vascular smooth muscle proliferation. One mechanism that explains the occurrence of endothelial dysfunction is the presence of elevated blood levels of asymmetric dimethylarginine (ADMA) – an L-arginine analogue that inhibits NO formation and thereby can impair vascular function. Supplementation with L-arginine has been shown to restore vascular function and to improve the clinical symptoms of various diseases associated with vascular dysfunction.
Beneficial Effects of L-Arginine
- Angina
- Congestive Heart Failure
- Hypertension
- Erectile dysfunction
- Sickle Cell Disease and Pulmonary Hypertension
The ratio of L-arginine to ADMA is considered to be the most accurate measure of eNOS substrate availability. This ratio will increase during L-arginine supplementation, regardless of initial ADMA concentration. Due to the pharmacokinetics of oral L-arginine and the positive results from preliminary studies, it appears supplementation with a sustained-release L-arginine preparation will achieve positive therapeutic results at lower dosing levels.
Many prospective clinical trials have shown that the association between elevated ADMA levels and major cardiovascular events and total mortality is robust and extends to diverse patient populations. However, we need to define more clearly in the future who will profit from ADMA determination, in order to use this novel risk marker as a more specific diagnostic tool.
Elimination of ADMA by way of DDAH
Asymmetric dimethylarginine (ADMA) and monomethyl arginine (L-NMMA) are endogenously produced amino acids that inhibit all three isoforms of nitric oxide synthase (NOS). ADMA accumulates in various disease states, including renal failure, diabetes and pulmonary hypertension, and its concentration in plasma is strongly predictive of premature cardiovascular disease and death. Both LNMMA and ADMA are eliminated largely through active metabolism by dimethylarginine dimethylaminohydrolase (DDAH) and thus DDAH dysfunction may be a crucial unifying feature of increased cardiovascular risk. These investigators ask whether ADMA is the underlying issue related to the pathogenesis of the vascular disorder.
They identified the structure of human DDAH-1 and probed the function of DDAH-1 both by deleting the Ddah1 gene in mice and by using DDAH-specific inhibitors that is shown by crystallography, bind to the active site of human DDAH-1. The loss of DDAH-1 activity leads to accumulation of ADMA and reduction in NO signaling. This in turn causes vascular pathophysiology, including endothelial dysfunction, increased systemic vascular resistance and elevated systemic and pulmonary blood pressure. The results suggest that DDAH inhibition could be harnessed therapeutically to reduce the vascular collapse associated with sepsis.
Methylarginines are formed when arginine residues in proteins are methylated by the action of protein arginine methyltransferases (PRMTs), and free methylarginines are liberated following proteolysis. Clear demonstration of an effect of endogenous ADMA and L-NMMA on cardiovascular physiology would be of importance, not only because of the implications for disease, but also because it would expose a link between post-translational modification of proteins and signaling through a proteolytic product of these modified proteins.
Which is it? ADMA or DDHA: Intrusion of a Genetic alteration.
The study showed that loss of DDAH expression or activity causes endothelial dysfunction, we believe that DDAH inhibition could potentially be used therapeutically to limit excessive NO production, which can have pathological effects. They then showed treated cultured isolated blood vessels with lipopolysaccharide (LPS) induced expression of the inducible isoform of NO synthase (iNOS) and generated high levels of NO, which were blocked by the iNOS-selective inhibitor 1400W and by DDAH inhibitors. Treatment of isolated blood vessels with DDAH inhibitors significantly increased ADMA accumulation in the culture medium. Treatment of isolated blood vessels with bacterial LPS led to the expected hyporeactivity to the contractile effects of phenylephrine, which was reversed by treatment with a DDAH inhibitor. The effect of the DDAH inhibitor was large and stereospecific, and was reversed by the addition of L-arginine.
In conclusion, genetic and chemical-biology approaches provide compelling evidence that loss of DDAH-1 function results in increased ADMA concentrations and thereby disrupts vascular NO signaling. A broader implication of this study is that post-translational methylation of arginine residues in proteins may have downstream effects by affecting NO signaling upon hydrolysis and release of the free methylated amino acid. This signaling pathway seems to have been highly conserved through evolution.
The crucial role of nitric oxide (NO) for normal endothelial function is well known. In many conditions associated with increased risk of cardiovascular diseases such as hypercholesterolemia, hypertension, abdominal obesity, diabetes and smoking, NO biosynthesis is dysregulated, leading to endothelial dysfunction. The growing evidence from animal and human studies indicates that endogenous inhibitors of endothelial NO synthase such as asymmetric dimethylarginine (ADMA) and NG-monomethyl-L-arginine (L-NMMA) are associated with the endothelial dysfunction and potentially regulate NO synthase.
Asymmetric dimethylarginine (ADMA) is one of three known endogenously produced circulating methylarginines (i.e. ADMA, NG-monomethyl-L-arginine (L-NMMA) and symmetrically methylated NG, NG-dimethyl-L-arginine). ADMA is formed by the action of protein arginine methyltransferases that methylate arginine residues in proteins and after which free ADMA is released. ADMA and L-NMMA can competitively inhibit NO elaboration by displacing L-arginine from NO synthase (NOS). The amount of methylarginines is related to overall metabolic activity and the protein turnover rate of cells. Although methylarginines are excreted partly by the kidneys, the major route of elimination of ADMA in humans is metabolism by the dimethylarginine dimethylaminohydrolase enzymes[ dimethylarginine dimethylaminohydrolase-1 and -2 (DDAH)] enzymes. Inhibition of DDAH leads to the accumulation of ADMA and consequently to inhibition of NO-mediated endothelium dependent relaxation of blood vessels.
The potential role of ADMA in angina pectoris has been evaluated by Piatti and co-workers, who reported ADMA levels to be higher in patients with cardiac syndrome X (angina pectoris with normal coronary arteriograms) than in controls. According to preliminary results from the CARDIAC (Coronary Artery Risk Determination investigating the influence of ADMA Concentration) study, patients with coronary heart disease (n 816) had a higher median ADMA plasma concentration than age and sex matched controls (median 0.91 vs. 0.70 mol/l; p 0.0001). Further, in a prospective Chinese study, a high plasma ADMA level independently predicted subsequent cardiovascular adverse events (cardiovascular death, myocardial infarction, and repeated revascularization of a target vessel).
There are only few published findings concerning variations in human DDAH. However, polymorphisms in other genes potentially related to risk factors for endothelial dysfunction and cardiovascular events have been studied. Reduced NO synthesis has been implicated in the development of atherosclerosis. For example, there are some functionally important variants of the NOS that could affect individual vulnerability to atherosclerosis by changing the amount of NO generated by the endothelium.
There are probably several functional variations in genes coding DDAH enzymes in different populations. Some of them could confer protection against the harmful effects of elevated ADMA and others impair enzyme function causing accumulation of ADMA in cytosol and/or blood.
In a study of 16 men with either low or high plasma ADMA concentrations were screened to identify DDAH polymorphisms that could potentially be associated with increased susceptibility to cardiovascular diseases. In that study a novel functional mutation of DDAH-1 was identified; the mutation carriers had a significantly elevated risk for cardiovascular disease and a tendency to develop hypertension. These results confirmed the clinical role of DDAH enzymes in ADMA metabolism. Furthermore, it is possible that more common variants of DDAH genes contribute more widely to increased cardiovascular risk.
We found a rare variation in the DDAH-1 gene, which is associated with elevated plasma concentrations of ADMA in heterozygous mutation carriers. There was also an increased prevalence of CHD and a tendency to hypertension among individuals with this DDAH-1 mutation. These observations highlight the importance of ADMA as a possible risk factor and emphasize the essential role of DDAH in regulating ADMA levels.
ADMA Elevation and Coronary Artery Disease
Endothelial dysfunction may be considered as a systemic disorder and involves different vascular beds. Coronary endothelial dysfunction (CED) precedes the development of coronary. Endothelial dysfunction is characterized by a reduction in endogenous nitric oxide (NO) activity, which may be accompanied by elevated plasma asymmetric dimethylarginine (ADMA) levels. ADMA is a novel endogenous competitive inhibitor of NO synthase (NOS), an independent marker for cardiovascular risk.
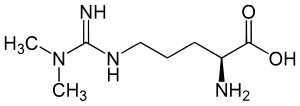
English: Structure of asymmetric dimethylarginine; ADMA; N,N-Dimethylarginine Deutsch: Asymmetrisches Dimethylarginin; N,N-Dimethyl-L-arginin; Guanidin-N,N-dimethylarginin (Photo credit: Wikipedia)
In a small study fifty-six men without obstructive coronary artery disease (CAD) who underwent coronary endothelial function testing were studied. Men with CED had significant impairment of erectile function (P ¼ 0.008) and significantly higher ADMA levels (0.50+0.06 vs. 0.45+0.07 ng/mL, P ¼ 0.017) compared with men with normal endothelial function. Erectile function positively correlated with coronary endothelial function. This correlation was independent of age, body mass index, high-density lipoprotein, C-reactive protein, homeostasis model assessment of insulin resistance index, and smoking status, suggesting that CED is independently associated with ED and plasma ADMA concentration in men with early coronary atherosclerosis.
ADMA and Chronic Renal Failure in Hepatorenal Syndrome
The concentration of SDMA was significantly higher in the patients with HRS compared to the patients without HRS and it was also higher than the values obtained from the healthy participants (1.76 ± 0.3 μmol/L; 1.01 ± 0.32 and 0.520 ± 0.18 μmol/L, respectively; p < 0.01). The concentrations of ADMA were higher in the cirrhotic patients with HRS than in those without this serious complication of cirrhosis. The concentration of ADMA in all the examined cirrhotic patients was higher than those obtained from healthy volunteers (1.35 ± 0.27 μmol/L, 1.05 ± 0.35 μmol/L and 0.76 ± 0.21 μmol/L, respectively). In the patients with terminal alcoholic liver cirrhosis, the concentrations
of ADMA and SDMA correlated with the progress of cirrhosis as well as with the development of cirrhosis complications. In the patients with HRS there was a positive correlation between creatinine and SDMA in plasma (r2 = 0.0756, p < 0.001) which was not found between creatinine and ADMA. The results demonstrate that the increase in SDMA concentration is proportionate to the progression of chronic damage of the liver and kidneys. Increased ADMA concentration can be a causative agent of renal insufficiency in patients with cirrhosis.
In patients with cirrhosis, ADMA, as well as SDMA could be markers for kidney insufficiency development. Accumulation of ADMA in plasma causes kidney
vasoconstriction and thereby retention of SDMA. Considering that ADMA has several damaging effects, it can be concluded that modulation of the activity of enzyme which participates in ADMA catabolism may represent a new therapeutic goal which is intended to reduce the progress of liver and kidney damage and thus the development of HRS.
ADMA Therapeutic Targets
Elevated plasma concentrations of the endogenous nitric oxide synthase
inhibitor asymmetric dimethylarginine (ADMA) are found in various clinical settings, including
- renal failure,
- coronary heart disease,
- hypertension,
- diabetes and
- preeclampsia.
In healthy people acute infusion of ADMA promotes vascular dysfunction,
and in mice chronic infusion of ADMA promotes progression of atherosclerosis.
Thus, ADMA may not only be a marker but also an active player in cardiovascular disease, which makes it a potential target for therapeutic interventions.
This review provides a summary and critical discussion of the presently available data concerning the effects on plasma ADMA levels of cardiovascular drugs, hypoglycemic agents, hormone replacement therapy, antioxidants, and vitamin supplementation.
We assess the evidence that the beneficial effects of drug therapies on vascular function can be attributed to modification of ADMA levels. To develop more specific ADMA-lowering therapies, mechanisms leading to elevation of plasma ADMA concentrations in cardiovascular disease need to be better understood.
ADMA is formed endogenously by degradation of proteins containing arginine residues that have been methylated by S-adenosylmethionine-dependent methyltransferases (PRMTs). There are two major routes of elimination: renal excretion and enzymatic degradation by the dimethylarginine dimethylaminohydrolases (DDAH-1 and -2).
Oxidative stress causing upregulation of PRMT expression and/or attenuation of DDAH activity has been suggested as a mechanism and possible drug target in clinical conditions associated with elevation of ADMA. As impairment of DDAH activity or capacity is associated with substantial increases in plasma ADMA concentrations, DDAH is likely to emerge as a prime target for specific therapeutic interventions.
Cardiovascular diseases (CVD) in diabetic patients have endothelial dysfunction as a key pathogenetic event. Asymmetric dimethylarginine (ADMA), an endogenous inhibitor of nitric oxide synthase (NOS), plays a pivotal role in endothelial dysfunction. Different natural polyphenols have been shown to preserve endothelial function and prevent CVD. Another study assessed the effect of silibinin, a widely used flavonolignan from milk thistle, on ADMA levels and endothelial dysfunction in db/db mice.
Plasma and aorta ADMA levels were higher in db/db than in control lean mice. Silibinin administration markedly decreased plasma ADMA; consistently, aorta ADMA was reduced in silibinin-treated animals. Plasma and aorta ADMA levels exhibited a positive correlation, whereas liver ADMA was inversely correlated with both plasma and aorta ADMA concentrations. Endothelium-(NO)-dependent vasodilatation to ACh was impaired in db/db mice and was restored in the silibinin group, in accordance with the observed reduction of plasma and vascular levels of ADMA. Endothelium-independent vasodilatation to SNP was not modified by silibinin administration.
Endothelin Inhibitors
Endothelins are potent vasoconstrictors and pressor peptides and are important mediators of cardiac, renal andendocrine functions. Increased ET-1 levels in disease states such as congestive heart failure, pulmonary hypertension, acute myocardial infarction, and renal failure suggest the endothelin system as an attractive target for pharmacotherapy. A non-peptidic, selective, competitive endothelin receptor antagonist with an affinity for the ETA receptor in the subnanomolar range was administered by continuous intravenous infusion to beagle dogs, rats, and Goettingen minipigs. It caused mild arteriopathy characterised by segmental degeneration in the media of mid- to large-size coronary arteries in the heart of dog, but not rat or minipig.
The lesions only occurred in the atrium and ventricle. Frequency and severity of the vascular lesions was not sex or dose related. No effects were noted in blood vessels in other organs or tissue. Plasma concentrations at steady state, and overall exposure in terms of AUC(0–24h) were higher in minipig and rat than the dog but did not cause cardiac arteriopathy. These findings concur with those caused by other endothelin anatagonists, vasodilators and positive inotropic: vasodilating drugs such as potassium channel openers, phosphodiesterase inhibitors and peripheral vasodilators.
Results by echocardiography indicate treatment-related local vasodilatation in the coronary arteries. These data suggest that the coronary arteriopathy may be the result of exaggerated pharmacology. Sustained vasodilatation in the coronary vascular bed may alter flow dynamics and lead to increased shear stress and tension on the coronary wall with subsequent microscopic trauma. In our experience with a number of endothelin receptor antagonists, the cardiac arteriopathy was only noted in studies with multiple daily or continuous intravenous infusion inviting speculation that sustained high plasma levels are needed for development of the lesions.
Up-regulation of vascular endothelin type B (ETB) receptors is implicated in the
pathogenesis of cardiovascular disease. Culture of intact arteries has been shown to induce similar receptor alterations and has therefore been suggested as a suitable method for, ex vivo, in detail delineation of the regulation of endothelin receptors. We hypothesize that mitogen-activated kinases (MAPK) and protein kinase C (PKC) are involved in the regulation of endothelin ETB receptors in human internal mammary arteries.
The endothelin-1-induced contraction (after endothelin ETB receptor desensitization) and the endothelin ETA receptor mRNA expression levels were not altered by culture. The sarafotoxin 6c contraction, endothelin ETB receptor protein and mRNA expression levels were increased. This increase was antagonized by;
PKC inhibitors (10 μM bisindolylmaleimide I and 10 μM Ro-32-0432), and
inhibitors of the p38, extracellular signal related kinases 1 and 2 (ERK1/2) and C-jun terminal kinase (JNK) MAPK pathways
Endothelin Receptor Antagonist Tezosentan
The effects of changes in the mean (Sm) and pulsatile (Sp) components of arterial wall shear stress on arterial dilatation of the iliac artery of the anaesthetized dog were examined in the absence and presence of the endothelin receptor antagonist tezosentan (10 mg kg_1 I.V.; Ro 61-0612; [5-isopropylpyridine-2-sulphonic acid 6-(2-hydroxy-ethoxy)-5-(2-methoxy-phenoxy)-2-(2-1H-tetrazol-5-ylpyridin-4-yl)-pyrimidin-4-ylamide]).
Changes in shear stress were brought about by varying local peripheral resistance and stroke volume using a distal infusion of acetylcholine and stimulation of the left ansa subclavia. An increase in Sm from 1.81 ± 0.3 to 7.29 ± 0.7 N m_2 (means ± S.E.M.) before tezosentan caused an endothelium-dependent arterial dilatation which was unaffected by administration of tezosentan for a similar increase in Sm from 1.34 ± 0.6 to 5.76 ± 1.4 N m_2 (means ± S.E.M.).
In contrast, increasing the Sp from 7.1 ± 0.8 to a maximum of 11.5 ± 1.1 N m_2 (means ± S.E.M.) before tezosentan reduced arterial diameter significantly. Importantly, after administration of tezosentan subsequent increases in Sp caused arterial dilatation for the same increase in Sp achieved prior to tezosentan, increasing from a baseline of 4.23 ± 0.4 to a maximum of 9.03 ± 0.9 N m_2 (means ± S.E.M.; P < 0.001). The results of this study provide the first in vivo evidence that pulsatile shear stress is a stimulus for the release of endothelin from the vascular endothelium.
Exercise and Diet
Vascular endotheliumis affected by plasma asymmetric dimethylarginine (ADMA), and it is induced by inflammatory cytokines of tumour necrosis factor (TNF)-a in vitro. Would a tight glycemic control restore endothelial function in patients with type-2 diabetes mellitus (DM) with modulation of TNF-a and/or reduction of ADMA level? In 24 patients with type-2 DM, the flow-mediated, endothelium-dependent dilation (FMD: %) of brachial arteries during reactive hyperaemia was determined by a high-resolution ultrasound method. Blood samples for glucose, cholesterol, TNF-a, and ADMA analyses were also collected from these patients after fasting. No significant glycemic or FMD changes were observed in 10 patients receiving the conventional therapy.
In 14 patients who were hospitalized and intensively treated, there was a significant decrease in glucose level after the treatment [from 190+55 to 117+21 (mean+SD) mg/dL, P , 0.01]. After the intensive control of glucose level, FMD increased significantly (from 2.5+0.9 to 7.2+3.0%), accompanied by a significant (P , 0.01) decrease in TNF-a (from 29+16 to 11+9 pg/dL) and ADMA (from 4.8+1.5 to 3.5+1.1 mM/L) levels. The changes in FMD after treatment correlated inversely with those in TNF-a (R ¼ 20.711, P , 0.01) and ADMA (R ¼ 20.717, P , 0.01) levels.
The exaggerated blood pressure response to exercise (EBPR) is an independent predictor of hypertension. Asymmetric dimethylarginine (ADMA) is an endogenous nitric oxide inhibitor and higher plasma levels of ADMA are related to increased cardiovascular risk. The aim of this study is to identify the relationship between ADMA and EBPR.
A total of 66 patients (36 with EBPR and 30 as controls) were enrolled in the study. EBPR is defined as blood pressure (BP) measurements ≥200/100 mmHg during the treadmill test. All the subjects underwent 24-h ambulatory BP monitoring. L-arginine and ADMA levels were measured using a high performance lipid chromatography technique.
The serum ADMA levels were increased in the EBPR group compared to the healthy controls (4.0±1.4 vs 2.6±1.1 μmol/L respectively, P=0.001), but L-arginine levels were similar in the 2 groups (P=0.19). The serum ADMA levels were detected as an independent predictor of EBPR (odds ratio 2.28; 95% confidence interval 1.22–4.24; P=0.002). Serum ADMA levels might play a role in EBPR to exercise.
Endothelial dysfunction occurs early in atherosclerosis in response to cardiovascular risk factors. The occurrence of endothelial dysfunction is primarily the result of reduced nitric oxide (NO) bioavailabilty. It represents an independent predictor of cardiovascular events and predicts the prognosis of the patient. Therefore, endothelial function has been identified as a target for therapeutic intervention. Regular exercise training is a nonpharmacological option to improve endothelial dysfunction in patients with cardiovascular disease by increasing NO bioavailability.
Peripheral Arterial Disease (PAD) is a cause of significant morbidity and mortality in the Western world. risk factor modification and endovascular and surgical revascularisation are the main treatment options at present. However, a significant number of patients still require major amputation. There is evidence that nitric oxide (NO) and its endogenous inhibitor asymmetric dimethylarginine (ADMA) play significant roles in the pathophysiology of PAD.
This paper reviews experimental work implicating the ADMA-DDAH-NO pathway in PAD, focusing on both the vascular dysfunction and both the vascular dysfunction and effects within the ischaemic muscle, and examines the potential of manipulating this pathway as a novel adjunct therapy in PAD.
In patients with CHF, the peripheral vascular resistance is increased via activation of the neurohormonal system, namely by autonomous sympathetic nervous system, rennin -angiotensin- aldosterone system (RAAS), and endothelin system. The vascular endothelial function in patients with CHF, mainly represented by the endothelium-dependent vasodilation, is altered.
Such alteration leads to increased vascular tone and remodeling of the blood vessels, reducing the peripheral blood flow. Hence, the amount of oxygen for the skeletal muscles is compromised, with progressive exercise intolerance. The vascular endothelial dysfunction in the CHF is mainly due to the decrease of the nitric oxide production induced by the reduced gene expression of eNOS and increased oxidative stress.
The endothelium-dependent vasodilation alteration has been virtually reported in all cardiovascular diseases. Using sauna bath as therapeutic option for CHF is not very recent, since in the 1950’s the first studies with CHF patients were conducted and the potential beneficial effect of sauna was suggested. However, some time later the studies emphasized especially its risks and recommended caution in its use for cardiac patients.
Frequently, sports medicine physicians are invited to evaluate the impact of the sauna on diseases and on health in general. Sauna can be beneficial or dangerous depending on its use. In the past few years the sauna is considered beneficial for the cardiovascular diseases’ patients, as the heart failure and lifestyle-related diseases, mainly by improving the peripheral endothelial function through the increase in cardiac output and peripheral vasodilation.
It is widely known that the vasodilators, such as angiotensin converting enzyme inhibitors, improve the CHF and increase the peripheral perfusion. Since the endothelial function is altered in CHF, the endothelium is considered as a new therapeutic target in heart failure. Hence, the angiotensin converting enzyme inhibitors and physical training improve the endothelial function in CHF patients. One of the proposed mechanisms for the alteration of the endothelium-dependent vasodilation would be through the decrease of the NO production in the peripheral vessels in CHF patients. The decrease of peripheral perfusion would decrease the shear stress. The shear stress is an important stimulus for NO production and eNOS expression. On the other hand, the heat increases the cardiac output and improves the peripheral perfusion in CHF patients. Consequently, with the cardiac output improvement in CHF patients, an increase of the shear stress, NO production and eNOS expression are expected.
Sauna bath
The sauna bath represents a heat load of 300-600 W/m2 of body surface area. The skin temperature rapidly increases to ± 40o-41oC and the thermoregulatory mechanisms are triggered. Evaporative heat transfer by sweating is the only effective body heat loss channel in dry sauna. The sweating begins rapidly and reaches its maximum level in ± 15 min. The total sweat secretion represents a heat loss of about 200 W/m2 of the body surface area. The body cannot compensate for the heat load and causing elevation of internal temperature. The skin circulation increases substantially. The skin blood flow, in the thermo-neutral condition (± 20oC) and in rest corresponding to ± 5-10% of the cardiac output, can reach ± 50-70% of the cardiac output.
Thermal therapy in 60oC produced systemic arterial, pulmonary arterial and venous vasodilation, reduced the preload and afterload and improved the cardiac output and the peripheral perfusion, clinical symptoms, life quality, and cardiac arrhythmias in CHF patients. In infants with severe CHF secondary to ventricular septal defect, the sauna therapy decreased the systemic vascular resistance and increased the cardiac output. The sauna benefits in CHF patients are possibly caused by the improvement of the vascular endothelial function and normalization of the neurohormonal system .
Ikeda et al. discovered that the observed improvements in the sauna therapy are due to the eNOS expression increase in the arterial endothelium. They later showed that the thermal therapy with sauna improves the survival of the TO-2 cardiomyopathic hamsters with CHF and, more recently, showed that the repetitive therapy with sauna increases the eNOS expression and the nitric oxide production in artery endothelium of TO-2 cardiomyopathic hamsters with CHF.
Whether n-3 polyunsaturated fatty acid (PUFA) supplementation and/or diet intervention might have beneficial influence on endothelial function was assessed using plasma levels of ADMA and L-arginine. A male population (n = 563, age 70 ± 6 yrs) with long-standing hyperlipidemia, characterized as high risk individuals in 1970–72, was included, randomly allocated to receive placebo n-3 PUFA capsules (corn oil) and no dietary advice (control group), dietary advice (Mediterranean type), n-3 PUFA capsules, or dietary advice and n-3 PUFA combined and followed for 3 years. Fasting blood samples were drawn at baseline and the end of the study.
Compliance with both intervention regimens were demonstrated by changes in serum fatty acids and by recordings from a food frequency questionnaire. No influence of either regimens on ADMA levels were obtained. However, n-3 PUFA supplementation was accompanied by a significant increase in L-arginine levels, different from the decrease observed in the placebo group (p < 0.05). In individuals with low body mass index (<26 kg/m2), the decrease in L-arginine on placebo was strengthened (p = 0.01), and the L-arginine/ADMA ratio was also significantly reduced (p = 0.04). In this rather large randomized intervention study, ADMA levels were not influenced by n-3 PUFA supplementation or dietary counselling. n-3 PUFA did, however, counteract the age related reduction in L-arginine seen on placebo, especially in lean individuals, which might be considered as an improvement of endothelial function.
Traditional Chinese Medicine
Traditional Chinese Medicine (TCM) involves a broad range of empirical testing and refinement and plays an important role in the health maintenance for people all over the world. However, due to the complexity of Chinese herbs, a full understanding of TCM’s action mechanisms is still unavailable despite plenty of successful applications of TCM in the treatment of various diseases, including especially cardiovascular diseases (CVD), one of the leading causes of death.
An integrated system of TCM has been constructed to uncover the underlying action mechanisms of TCM by incorporating the chemical predictors, target predictors and network construction approaches from three representative Chinese herbs, i.e., Ligusticum chuanxiong Hort., Dalbergia odorifera T. Chen and Corydalis yanhusuo WT Wang widely used in CVD treatment, by combined use of drug absorption, distribution, metabolism and excretion (ADME) screening and network pharmacology techniques. These studies have generated 64 bioactive ingredients and identified 54 protein targets closely associated with CVD, to clarify some of the common conceptions in TCM, and provide clues to modernize such specific herbal medicines.
Ligusticum chuanxiong Hort., Dalbergia odorifera T. Chen and Corydalis yanhusuo WT Wang
Twenty-two of 194 ingredients in Ligusticum chuanxiong demonstrate good bioavailability (60%) after oral administration. Interestingly, as the most abundant bioactive compound of Chuanxiong, Ligustilide (M120) only has an adequate OB of 50.10%, although it significantly inhibits the vasoconstrictions induced by norepinephrine bitartrate (NE) and calcium chloride (CaCl2). Indeed, this compound can be metabolized to butylidenephthalide, senkyunolide I (M156), and senkyunolide H (M155) in vivo.
The three natural ingredients produce various pharmacological activities in cerebral blood vessels, the general circulatory system and immune system including spasmolysis contraction effects, inhibitory effects of platelet aggregation and anti-proliferative activity, and thus improve the therapeutic effect on patients. Cnidilide (M93, OB = 77.55%) and spathulenol (M169, OB = 82.37%) also closely correlate with the smooth muscle relaxant action, and thereby have the strongest spasmolytic activity. Carotol (M8) and Ferulic acid (M105) with an OB of 149.03% and 86.56%, respectively, demonstrate better bioavailability compared with cnidilide and spathulenol, which show strong antifungal, antioxidant and anti-inflammatory activity.
The pharmacological activity of ferulic acid results in the improvement of blood fluidity and the inhibition of platelet aggregation, which may offer beneficial effects against cancer, CVD, diabetes and Alzheimer’s disease. As for 3-n-butylphthalide (M85, OB = 71.28%), this compound is not only able to inhibit platelet aggregation, but also decreases the brain infarct volume and enhances microcirculation, thus benefiting patients with ischemic stroke. Platelet aggregation represents a multistep adhesion process involving distinct receptors and adhesive ligands, with the contribution of individual receptor-ligand interactions to the aggregation process depending on the prevailing blood flow conditions, implying that the rheological (blood flow) conditions are an important impact factor for platelet aggregation. Moreover, thrombosis, the pathological formation of platelet aggregates and one of the biggest risk factors for CVD, occludes blood flow causing stroke and heart attack. This explains why the traditional Chinese herb Ligusticum chuanxiong that inhibits platelet aggregates forming and promotes blood circulation can be used in treatment of CVD.
Twenty-six percent (24 of 93) of the ingredients in Dalbergia odorifera meet the OB > 60% criterion irrespective of the pharmacological activity. Relatively high bioavailability values were predicted for the mainly basic compounds odoriflavene (M275, OB = 84.49%), dalbergin (M247, OB = 78.57%), sativanone (M281, OB = 73.01%), liquiritigenin (M262, OB = 67.19%), isoliquiritigenin (M259, OB = 61.38%) and butein (M241, OB = 78.38%). Interestingly, all of the six ingredients show obvious anti-inflammatory property. Butein, liquiritigenin and isoliquiritigenin inhibit cell inflammatory responses by suppressing the NF-κB activation induced by various inflammatory agents and carcinogens, and by decreasing the NF-κB reporter activity. Inflammation occurs with CVD, and Dalbergia odorifera, one of the most potent anti-cardiovascular and anti-cerebrovascular agents, exerts great anti-inflammatory activity.
Corydalis yanhusuo has gained ever-increasing popularity in today’s world because of its therapeutic effects for the treatment of cardiac arrhythmia disease, gastric and duodenal ulcer and menorrhalgia. In our work, 21% (15 of 73) of chemicals in this Chinese herb display good OB (60% or even high), and the four main effective ingredients are natural alkaloid agents.
Dehydrocorydaline blocks the release of noradrenaline from the adrenergic nerve terminals in both the Taenia caecum and pulmonary artery, and thereby inhibits the relaxation or contraction of adrenergic neurons. As for dehydrocavidine with an OB of 47.59%, this alkaloid exhibits a significant spasmolytic effect, which acts via relaxing smooth muscle.
In recent years, CVD has been at the top list of the most serious health problems. Many different types of therapeutic targets have already been identified for the management and prevention of CVD, such as endothelin and others. The key question asked is
- what the interactions of the active ingredients of the Chinese herbs are with their protein targets in a systematic manner and
- how do the corresponding targets change under differential perturbation of the chemicals?
The study used an unbiased approach to probe the proteins that bind to the small molecules of interest in CVD on the basis of the Random Forest (RF) and Support Vector Machine (SVM) methods combining the chemical, genomic and pharmacological information for drug targeting and discovery on a large scale. Applied to 64 ingredients derived from the three traditional Chinese medicines Dalbergia odorifera, Ligusticum chuanxiong and Corydalis yanhusuo, which show good OB, 261 ligand-target interactions have been constructed, 221 of which are enzymes, receptors, and ion channels. This indicates that chemicals with multiple relative targets are responsible for the high interconnectedness of the ligand-target interactions. The promiscuity of drugs has restrained the advance in recent TCM, because they were thought to be undesirable in favor of more target-specific drugs.
Target Identification and Validation
To validate the reliability of these target proteins, the researchers performed a docking analysis to select the ligand-protein interactions with a binding free energies of ≤−5.0 kcal/mol, which leads to the sharp reduction of the interaction number from 5982 to 760. These drug target candidates were subsequently subject to PharmGkb (available online: http://www.pharmgkb.org; accessed on 1 December 2011), a comprehensive disease-target database, to investigate whether they were related to CVD or not, and finally, 54 proteins were collected and retained.
Fourty-two proteins (76%) were identified as the targets of Ligusticum chuanxiong, such as dihydrofolate reductase (P150), an androgen receptor (P210) and angiotensin-converting enzyme (P209) that were involved in the development of CVD. Of the proteins, seven and two were recognized as those of Dalbergia odorifera and Corydalis yanhusuo, respectively. For Dalbergia odorifera, this Chinese herb has 48 potential protein targets, 13 of which have at least one link to other drugs.
The three herbs share 29 common targets, accounting for 52.7% of the total number. Indeed, as one of the most important doctrines of TCM
abstracted from direct experience and perception, “multiple herbal drugs for one disease” has played an undeniable role. These studies explored the targets of the three Chinese herbs, indicating that these drugs target the same targets simultaneously and exhibit similar pharmacological effects on CVD. This is consistent with the theory of “multiple herbal drugs for one disease”.
The three Chinese herbs possess specific targets. The therapeutic efficacy of a TCM depends on multiple components, targets and pathways. The complexity becomes a huge obstacle for the development and innovation of TCM. For example, the Chinese herb Ligusticum chuanxiong identifies the protein caspase-3 (P184), a cysteinyl aspartate-specific protease, as one of its specific targets, and exhibits inhibitory effects on the activity of this protease. In fact, connective tissue growth factor enables the activation of caspase-3 to induce apoptosis in human aortic vascular smooth muscle cells.
Thus, modulation of the activity of caspase-3 with Ligusticum chuanxiong suggests an efficient therapeutic approach to CVD. The Chinese herb Dalbergia odorifera has the α-2A adrenergic receptor (P216) as its specific target and probably blocks the release of this receptor, and thus influences its action. As for Corydalisyanhusuo, the protein tyrosine-protein kinase JAK2 (P9) is the only specific target of this Chinese herb. The results indicate different specific targets possessed by the three Chinese herbs.
Ligand-Candidate Target and Ligand-Potential Target Networks
Previous studies have already reported the relationships of the small molecules with CVD, which indicates the reliability of our results [45,46]. Regarding the candidate targets, we have found that prostaglandin G/H synthase 2 (P46) and prostaglandin G/H synthase 1 (P47) possess the largest number of connected ingredients. Following are nitric-oxide synthase, endothelial (P66) and tyrosine-protein phosphatase non-receptor type 1 (P8), which have 62 and 61 linked chemicals, respectively.
The 29 targets shared by the three traditional Chinese herbs exhibit a high degree of correlations with CVD, which further verifies their effectiveness for the treatment of CVD. These results provide a clear view of the relationships of the target proteins with CVD and other related diseases, which actually link the Chinese herbs and the diseases via the protein targets. This result further explains the theory of “multiple herbal drugs for one disease” based on molecular pharmacology.
Target-Pathway Network
Cells communicate with each other using a “language” of chemical signals. The cell grows, divides,or dies according to the signals it receives. Signals are generally transferred from the outside of the cell. Specialized proteins are used to pass the signal—a process known as signal transduction. Cells have a number of overlapping pathways to transmit signals to multiple targets. Ligand binding in many of the signaling proteins in the pathway can change the cellular communication and finally affect cell growth and proliferation. The authors extracted nine signal pathways closely associated with CVD in PharmGkb (available online: http://www.pharmgkb.org; accessed on 1 December 2011).
As the main components in the VEGF system, proto-oncogene tyrosine-protein kinase Src, eNOS, and hsp90-α is also recognized as common targets of Dalbergia odorifera, Ligusticum chuanxiong and Corydalis yanhusuo, which are efficient for the treatment of CVD. This implies that the candidate drugs can target different target proteins involved in the same or different signal pathways, and thereby have potential effects on the whole signal system.
Target Prediction
In search of the candidate targets, the model that efficiently integrates the chemical, genomic and pharmacological information for drug targeting and discovery on a large scale is based on the two powerful methods Random Forest (RF) and Support Vector Machine (SVM). The model is supported by a large pharmacological database of 6511 drugs and 3999 targets extracted from the DrugBank database (available online: http://drugbank.ca/; accessed on 1 June 2011), and shows an impressive performance of prediction for drug-target interaction, with a concordance of 85.83%, a sensitivity of 79.62% and a specificity of 92.76%. the candidate targets were selected according to the criteria that the possibility of interacting with potential candidate targets was higher than 0.6 for the RF model and 0.7 for the SVM model. The obtained candidate targets were finally reserved and were further predicted for their targets.
Target Validation
Molecular docking analysis was carried out using the AutoDock software (available online: http://autodock.scripps.edu/; accessed on 1 February 2012). This approach performs the docking of the small, flexible ligand to a set of grids describing the target protein. During the docking process, the protein was considered as rigid and the molecules as flexible. The crystal structures of the candidate targets were downloaded from the RCSB Protein Data Bank (available online: http://www.pdb.org/; accessed on 1 December 2011), and the proteins without crystal structures were performed based on homology modeling using the Swiss-Model Automated Protein Modelling Server (available online: http://swissmodel.expasy.org/; accessed on 1 February 2012).
TCM is a heritage that is thousands of years old and is still used by millions of people all over the world—even after the development of modern scientific medicine. Chinese herbal combinations generally include one or more plants and even animal products.
The study identified 54 protein targets, which are closely associated with CVD for the three Chinese herbs, of which 29 are common targets (52.7%), which clarifies the mechanism of efficiency of the herbs for the treatment of CVD.
Activation of NFkB
Extracellular stimuli for NFkB activation and NFkB regulated genes
Extracellular stimuli Regulated genes
TNFa Growth factors (G/M-CSF)
Interleukin 1 G/M CSF, M CSF, G CSF
ROS Cell adhesion molecules
UV light ICAM-1, VCAM, E-Selectin, P-selectin
Ischaemia Cytokines
Lipopolysaccharide TNFa, IL-1, IL-2, IL-6, interferon
Bacteria Transcription regulators
Viruses P53, IkB, c-rel, c-myc
Amyloid Antiapoptotic proteins
Glutamate TRAF-1, TRAF-2, c-IAP1, c-IAP2
Pathophysiology
Reactive oxygen species (ROS) are toxic and in conditions of a dysbalance between their overproduction and the diminished activity of various antioxidant enzymes and other molecules induce cellular injury termed oxidative stress. ROS are often related to a number of diseases like atherosclerosis. However, the mechanism is not clear at all. Latest years of research have brought the idea of connection between ROS and NFkB. And indeed, in vitro studies showed a rapid activation of NFkB after exposure of certain cell types to ROS. Today, no specific receptor for ROS has been found, thus, the details of the ROS induced activation of NFkB are missing.
Natural occurring agents which actions are still a matter of debate in the theory and nouvelle small molecular derivates activate or inhibit the transcriptional factor. Synthetic oligo and polypeptide inhibitors of NFkB can penetrate the cell membrane and directly act on the Rel proteins. The most sophisticated approaches towards inhibiting the activation and translocation of NFkB into the nucleus represent gene deliveries, using plasmids or adenoviruses containing genes for various super repressors—modified IkB proteins, or so called NFkB decoys, which interact with activated NFkB and thus, inhibit the interaction between the transcription factor and nuclear DNA enhancers.
A simplified scheme of the activation of NFkB by the degradation of IkB. IkB is phosphorylated by IKK and ubiquinatated by the ubiquitine ligase system (ULS). IkB is further degradated by the 26S proteasome (26S).Activated NFkB can pass the nuclear membrane and interact with kB binding sequences in enhancers of NFkB regulated genes. LPS, lipopolysaccharide; ROS, reactive oxygen species; FasL, Fas ligand; TRAF, TNFa receptor associated factor; NIK, NFkB inducing kinase; MEKK, mitogen activated protein kinase/extracellular signal regulated kinases kinases.
The medicine of this century is a medicine of molecules, the diagnostic procedure and the therapy moves further from the “clinical picture” to the use of achievements in molecular biology and genetics. However, sober scepticism and awareness are indicated. Especially the role of NFkB in multiple signal transducing pathways and the tissue dependent variability of responses to alternations in NFkB pathway may be the reasons for unwanted side effects of the therapy that are after in vitro or in vivo experiments hardly to expect in the clinical use.
Therapeutic Targets
Modern drug discovery is primarily based on the search and subsequent testing of drug candidates acting on a preselected therapeutic target. Progress in genomics, protein structure, proteomics, and disease mechanisms has led to a growing interest in an effort for finding new targets and more effective exploration of existing targets. The number of reported targets of marketed and investigational drugs has significantly increased in the past 8 years. There are 1535 targets collected in the therapeutic target database.
Knowledge of these targets is helpful for molecular dissection of the mechanism of action of drugs and for predicting features that guide new drug design and the
search for new targets. This article summarizes the progress of target exploration and investigates the characteristics of the currently explored targets to analyze their sequence, structure, family representation, pathway association, tissue distribution, and genome location features for finding clues useful for searching for new targets. Possible “rules” to guide the search for druggable proteins and the feasibility of using a statistical learning method for predicting druggable proteins directly from their sequences are discussed.
Current Trends in Exploration of Therapeutic Targets
There are 395 identifiable targets described in 1606 patents. Of these targets, 264 have been found in more than one patent and 50 appear in more than 10 patents. The number of patents associated with a target can be considered to partly correlate with the level of effort and intensity of interest currently being directed to it. Approximately one third of the patents with an identifiable target were approved in the past year. This suggests that the effort for the exploration of these targets is ongoing, and there has been steady progress in the discovery of new investigational agents directed to these targets.
Various degrees of progress have been made toward discovery and testing of agents directed at these targets. However, for some of these targets, many difficulties remain to be resolved before viable drugs can be derived. The appearance of a high number of patents associated with these targets partly reflects the intensity of efforts for finding effective drug candidates against these targets.
There are 62 targets being explored for the design of subtype-specific drugs, which represents 15.7% of the 395 identifiable targets in U.S. patents approved in 2000 through 2004. Compared with the 11 targets of FDA approved subtype-specific drugs during the same period, a significantly larger number of targets are being explored for the design of subtype-specific drugs.
What Constitutes a Therapeutic Target?
The majority of clinical drugs achieve their effect by binding to a cavity and regulating the activity, of its protein target. Specific structural and physicochemical properties, such as the “rule of five” (Lipinski et al., 2001), are required for these drugs to have sufficient levels of efficacy, bioavailability, and safety, which define target sites to which drug-like molecules can bind. In most cases, these sites exist out of functional necessity, and their structural architectures accommodate target-specific drugs that minimally interact with other functionally important but structurally similar sites.
These constraints limit the types of proteins that can be bound by drug-like molecules, leading to the introduction of the concept of druggable proteins (Hopkins and Groom, 2002; Hardy and Peet, 2004). Druggable proteins do not necessarily become therapeutic targets (Hopkins and Groom, 2002); only those that play key roles in diseases can be explored as potential targets.
Prediction of Druggable Proteins by a Statistical Learning Method
Currently, the support vector machine (SVM) method seems to be the most accurate statistical learning method for protein predictions. SVM is based on the structural risk minimization principle from statistical learning theory. Known proteins are divided into druggable and nondruggable classes; each of these proteins is represented by their sequence-derived physicochemical features.
These features are then used by the SVM to construct a hyperplane in a higher dimensional hyperspace that maximally separates druggable proteins and nondruggable ones. By projecting the sequence of a new protein onto this hyperspace, it can be determined whether this protein is druggable from its location with respect to the hyperplane. It is a druggable protein if it is located on the side of druggable class.
References
Böger RH. Asymmetric dimethylarginine, an endogenous inhibitor of nitric oxide synthase, explains the ‘L-arginine paradox’ and acts as a novel cardiovascular risk factor. J Nutr 2004; 134: 2842S–7S.
B Dobutovi, K Smiljani, S Soski, HD Düngen and ER Isenovi. Nitric Oxide and its Role in Cardiovascular Diseases. The Open Nitric Oxide Journal, 2011; 3: 65-71. 1875-0427/11.
Vallance P, Leone A, Calver A, Collier J, Moncada S. Accumulation of an endogenous inhibitor of nitric oxide synthesis in chronic renal failure. Lancet 1992; 339(8793): 572−5.
ER Hedegaard, E Stankevicius, U Simonsen and O Fröbert. Non-endothelial endothelin counteracts hypoxic vasodilation in porcine large coronary arteries. BMC Physiology 2011; 11:8-20. http://www.biomedcentral.com/1472-6793/11/8
S Rajagopalan, D Pfenninger, C Kehrer, A Chakrabarti. Increased Asymmetric Dimethylarginine and Endothelin 1 Levels in Secondary Raynaud’s Phenomenon. Arthritis & Rheumatism 2003; 48(7): 1992–2000. DOI: 10.1002/art.11060
Boger RH. The emerging role of asymmetric dimethylarginine as a novel cardiovascular risk factor. Cardiovasc Res 2003;59:824-833.
Böger RH, Ron ES. L-Arginine Improves Vascular Function by Overcoming the Deleterious Effects of ADMA, a Novel Cardiovascular Risk Factor. Altern Med Rev 2005;10(1):14-23.
Böger RH. Asymmetric dimethylarginine (ADMA) and cardiovascular disease: insights from prospective clinical trials. Vascular Medicine 2005; 10(2): S19-S25. DOI: 10.1191/ 1358863x 05vm602oa.
J Leiper, M Nandi, B Torondel, J Murray-Rust, et al. Disruption of methylarginine metabolism impairs vascular homeostasis.
Murray-Rust, J. et al. Structural insights into the hydrolysis of cellular nitric oxide synthase inhibitors by dimethylarginine dimethylaminohydrolase. Nat. Struct. Biol. 2001; 8:679–683.
D Nilsson, LGustafsson, A Wackenfors, B Gesslein, et al. Up-regulation of endothelin type B receptors in the human internal mammary artery in culture is dependent on protein kinase C and mitogen-activated kinase signaling pathways. MC Cardiovascular Disorders 2008; 8:21-31. doi:10.1186/1471-2261-8-21. http://www.biomedcentral.com/1471-2261/8/21
GL Volti, S Salomone, V Sorrenti, A Mangiameli, et al. Effect of silibinin on endothelial dysfunction and ADMA levels in obese diabetic mice. Cardiovascular Diabetology 2011, 10:62. http://www.cardiab.com/content/10/1/62
Leiper, J., Murray-Rust, J., McDonald, N. & Vallance, P. S-nitrosylation of dimethylarginine dimethylaminohydrolase regulates enzyme activity: further interactions between nitric oxide synthase and DDAH. Proc. Natl. Acad. Sci. USA 2002; 99: 13527–13532.
R Maas. Pharmacotherapies and their influence on asymmetric dimethylargine (ADMA). Vascular Medicine 2005; 10(2): S49-S57. DOI : 10.1191/ 1358863x05vm605oa
Veli-Pekka Valkonen, Tomi-Pekka Tuomainen, R Laaksonen. DDAH gene and cardiovascular risk. Vascular Medicine 2005; 10: S45–48.
AA Elesber, H Solomon, RJ Lennon, V Mathew, et al. Coronary endothelial dysfunction is associated with erectile dysfunction and elevated asymmetric dimethylarginine in patients with early atherosclerosis. European Heart Journal 2006; 27: 824–831. doi:10.1093/eurheartj/ehi749.
S Yasuda, S Miyazaki, M Kanda, Y Goto, et al. Intensive treatment of risk factors in patients with type-2 diabetes mellitus is associated with improvement of endothelial function coupled with a reduction in the levels of plasma asymmetric dimethylarginine an endogenous inhibitor of nitric oxide synthase. European Heart Journal 2006; 27: 1159–1165. doi:10.1093/ eurheartj/ehi876.
F Markos, BA Hennessy, M Fitzpatrick, J O’Sullivan and HM Snow. The effect of tezosentan, a non-selective endothelin receptor antagonist, on shear stress-induced changes in arterial diameter of the anaesthetized dog. Journal of Physiology 2002; 544(3): 913–918. DOI: 10.1113/jphysiol.2002.030478. http://www.jphysiol.org
M Kayrak; A Bacaksiz; MA Vatankulu, SS Ayhan, et al. Association Between Exaggerated Blood Pressure Response to Exercise and Serum Asymmetric Dimethylarginine Levels. Hypertension and Circulatory Control. Circ J 2010; 74: 1135 – 1141.
C Walther, S Gielen, and R Hambrecht. The effect of exercise training on endothelial function in cardiovascular disease in humans. Exerc Sport Sci Rev 2004; 32(4): 129–134 .
D Abraham, S Selvakumar, DM Baker, and JCS Tsui. Nitric Oxide Manipulation: A Therapeutic Target for Peripheral Arterial Disease? Williams, Xu Shi-Wen, Hindawi Publishing Corporation, Cardiology Research and Practice 2012; Article ID 656247, 7 pages doi:10.1155/2012/656247G . Sidney G. Shaw, Ed.
M Stephan-Gueldner, A Inomata. Coronary arterial lesions induced by endothelin antagonists. Toxicology Letters 2000; 112–113: 531–535.
V Ničković, J Nikolić, N Djindjić, М Ilić, et al. Diagnostic significance of dimethylarginine in the development of hepatorenal syndrome in patients with alcoholic liver cirrhosis. Vojnosanit Pregl 2012; 1-6 .UDC: 616.89-008.441.3-06:[616.36-004-07:616.61-008.6-07DOI: 10.2298/ VSP110728009N.
HMA Eid, H Arnesen, EM Hjerkinn, T Lyberg, et al. Effect of diet and omega-3 fatty acid intervention on asymmetric dimethylarginine. Nutrition & Metabolism 2006; 3:4-14. doi:10.1186/1743-7075-3-4.
B Li, X Xu, X Wang, H Yu, X Li, et al. A Systems Biology Approach to Understanding the Mechanisms of Action of Chinese Herbs for Treatment of Cardiovascular Disease. Int. J. Mol. Sci. 2012; 13: 13501-13520; doi:10.3390/ ijms131013501. ISSN 1422-0067. http://www.mdpi.com/journal/ijms
P Celec. Nuclear factor kappa B—molecular biomedicine: the next generation. Biomedicine & Pharmacotherapy 2004; 58:365–371. http://www.elsevier.com/locate/biopha
C. J. ZHENG, L. Y. HAN, C. W. YAP, Z. L. JI, et al. Therapeutic Targets: Progress of Their Exploration and Investigation of Their Characteristics. Pharmacol Rev 2006; 58:259–279. 0031-6997/06/5802-259–279. http://pharmrev.aspetjournals.org/content/suppl/2006/05/26/58.2.259.D
Lev-Ari, A. Stem cells create new heart cells in baby mice, but not in adults, study shows
Lev-Ari, A. Cardiovascular Disease (CVD) and the Role of agent alternatives in endothelial Nitric Oxide Synthase (eNOS) Activation and Nitric Oxide Production
Lev-Ari, A. Bystolic’s generic Nebivolol – positive effect on circulating Endothelial Progenitor Cells endogenous augmentation
Lev-Ari, A. Macrovascular Disease – Therapeutic Potential of cEPCs: Reduction Methods for CV Risk
Lev-Ari, A. Heart patients’ skin cells turned into healthy heart muscle cells
Lev-Ari, A. Resident-cell-based Therapy in Human Ischaemic Heart Disease: Evolution in the PROMISE of Thymosin beta4 for Cardiac Repair
http://pharmaceuticalintelligence.com/2012/04/30/93/
Nitric Oxide and Sepsis, Hemodynamic Collapse, and the Search for Therapeutic Options
Mediterranean Diet is BEST for patients with established Heart Disorders
Endothelin Receptors in Cardiovascular Diseases: The Role of eNOS Stimulation
Inhibition of ET-1, ETA and ETA-ETB, Induction of NO production, stimulation of eNOS and Treatment Regime with PPAR-gamma agonists (TZD): cEPCs
Endogenous Augmentation for Cardiovascular Risk Reduction – A Bibliography
Reveals from ENCODE project will invite high synergistic collaborations to discover specific targets
Related articles
- Nitric Oxide has a ubiquitous role in the regulation of glycolysis -with a concomitant influence on mitochondrial function (pharmaceuticalintelligence.com)
- Clinical Trials Results for Endothelin System: Pathophysiological role in Chronic Heart Failure, Acute Coronary Syndromes and MI – Marker of Disease Severity or Genetic Determination? (pharmaceuticalintelligence.com)
- Latest Study Provides Evidence Confirming the Mechanism of Action of Vasomedical’s EECP Therapy in Treating Cardiovascular Disease (prweb.com)
- Statins’ nonlipid effects on vascular endothelium through eNOS activation (pharmaceuticalintelligence.com)
- Interaction of Nitric Oxide and Prostacyclin in Vascular Endothelium (pharmaceuticalintelligence.com)
- Nitric oxide: role in Cardiovascular health and disease (pharmaceuticalintelligence.com)









