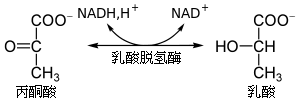
The Colors of Respiration and Electron Transport
Reporter & Curator: Larry H. Bernstein, MD, FCAP
Molecular Biology of the Cell. 4th edition
Electron-Transport Chains and Their Proton Pumps
http://www.ncbi.nlm.nih.gov/books/NBK26904/
Having considered in general terms how a mitochondrion uses electron
transport to create an electrochemical proton gradient, we need to
examine the mechanisms that underlie this membrane-based energy-conversion process. In doing so, we also accomplish a larger purpose.
As emphasized at the beginning of this chapter, very similar chemi-
osmotic mechanisms are used by mitochondria, chloroplasts, archea,
and bacteria. In fact, these mechanisms underlie the function of nearly
all living organisms— including anaerobes that derive all their energy
from electron transfers between two inorganic molecules. It is therefore
rather humbling for scientists to remind themselves that the existence
of chemiosmosis has been recognized for only about 40 years.
We begin with a look at some of the principles that underlie the electron-transport process, with the aim of explaining how it can pump protons
across a membrane.
Although protons resemble other positive ions such as Na+ and K+
in their movement across membranes, in some respects they are unique.
Hydrogen atoms are by far the most abundant type of atom in living
organisms; they are plentiful not only in all carbon-containing
biological molecules, but also in the water molecules that surround
them. The protons in water are highly mobile, flickering through the
hydrogen-bonded network of water molecules by rapidly
dissociating from one water molecule to associate with its neighbor,
as illustrated in Figure 14-20A. Protons are thought to move across a
protein pump embedded in a lipid bilayer in a similar way: they
transfer from one amino acid side chain to another, following a
special channel through the protein.
Protons are also special with respect to electron transport. Whenever
a molecule is reduced by acquiring an electron, the electron (e -) brings
with it a negative charge. In many cases, this charge is rapidly
neutralized by the addition of a proton (H+) from water, so that
the net effect of the reduction is to transfer an entire hydrogen atom,
H+ + e – (Figure 14-20B). Similarly, when a molecule is oxidized,
a hydrogen atom removed from it can be readily dissociated into
its constituent electron and proton—allowing the electron to
be transferred separately to a molecule that accepts electrons,
while the proton is passed to the water. Therefore, in a membrane
in which electrons are being passed along an electron-transport
chain, pumping protons from one side of the membrane to
another can be relatively simple. The electron carrier merely
needs to be arranged in the membrane in a way that causes it to
pick up a proton from one side of the membrane when it accepts
an electron, and to release the proton on the other side of the
membrane as the electron is passed to the next carrier molecule
in the chain (Figure 14-21).
http://www.ncbi.nlm.nih.gov/books/NBK26904/bin/ch14f21.gif
Figure 14-21
How protons can be pumped across membranes. As an electron
passes along an electron-transport chain embedded in a lipid-bilayer
membrane, it can bind and release a proton at each step.
In this diagram, electron carrier B picks up a proton (H+)
from one (more…)
The Redox Potential Is a Measure of Electron Affinities
In biochemical reactions, any electrons removed from one
molecule are always passed to another, so that whenever one
molecule is oxidized, another is reduced. Like any other chemical r
eaction, the tendency of such oxidation-reduction reactions, or
redox reactions, to proceed spontaneously depends on the free-
energy change (ΔG) for the electron transfer, which in turn
depends on the relative affinities of the two molecules for electrons.
Because electron transfers provide most of the energy for living
things, it is worth spending the time to understand them. Many
readers are already familiar with acids and bases, which donate
and accept protons (see Panel 2-2, pp. 112–113). Acids and bases
exist in conjugate acid-base pairs, in which the acid is readily
converted into the base by the loss of a proton. For example,
acetic acid (CH3COOH) is converted into its conjugate base
(CH3COO-) in the reaction:
Image ch14e3.jpg
In exactly the same way, pairs of compounds such as NADH and
NAD+ are called redox pairs, since NADH is converted to NAD+
by the loss of electrons in the reaction:
Image ch14e4.jpg
NADH is a strong electron donor: because its electrons are held
in a high-energy linkage, the free-energy change for passing its
electrons to many other molecules is favorable (see Figure 14-9).
It is difficult to form a high-energy linkage. Therefore its redox
partner, NAD+, is of necessity a weak electron acceptor.
The tendency to transfer electrons from any redox pair can be
measured experimentally. All that is required is the formation
of an electrical circuit linking a 1:1 (equimolar) mixture of the
redox pair to a second redox pair that has been arbitrarily selected
as a reference standard, so the voltage difference can be measured
between them (Panel 14-1, p. 784). This voltage difference is
defined as the redox potential; as defined, electrons move
spontaneously from a redox pair like NADH/NAD+ with a low
redox potential (a low affinity for electrons) to a redox pair like
O2/H2O with a high redox potential (a high affinity for electrons).
Thus, NADH is a good molecule for donating electrons to the
respiratory chain, while O2 is well suited to act as the “sink” for
electrons at the end of the pathway. As explained in Panel 14-1,
the difference in redox potential, ΔE0′, is a direct measure of
the standard free-energy change (ΔG°) for the transfer of an
electron from one molecule to another.
Box Icon
Panel 14-1
Redox Potentials.
Electron Transfers Release Large Amounts of Energy
As just discussed, those pairs of compounds that have the most negative
redox potentials have the weakest affinity for electrons and therefore
contain carriers with the strongest tendency to donate electrons.
Conversely, those pairs that have the most positive redox potentials
have the strongest affinity for electrons and therefore contain carriers
with the strongest tendency to accept electrons. A 1:1 mixture of NADH
and NAD+ has a redox potential of -320 mV, indicating that NADH has
a strong tendency to donate electrons; a 1:1 mixture of H2O and ½O2
has a redox potential of +820 mV, indicating that O2 has a strong
tendency to accept electrons. The difference in redox potential is
1.14 volts (1140 mV), which means that the transfer of each electron
from NADH to O2 under these standard conditions is enormously
favorable, where ΔG° = -26.2 kcal/mole (-52.4 kcal/mole for the two
electrons transferred per NADH molecule; see Panel 14-1). If we
compare this free-energy change with that for the formation of the
phosphoanhydride bonds in ATP (ΔG° = -7.3 kcal/mole, see Figure 2-75), we see that more than enough energy is released by the oxidization
of one NADH molecule to synthesize several molecules of ATP from
ADP and Pi.
Living systems could certainly have evolved enzymes that would
allow NADH to donate electrons directly to O2 to make water in the reaction:
Image ch14e5.jpg
But because of the huge free-energy drop, this reaction would proceed
with almost explosive force and nearly all of the energy would be released
as heat. Cells do perform this reaction, but they make it proceed much
more gradually by passing the high-energy electrons from NADH to
O2 via the many electron carriers in the electron-transport chain.
Since each successive carrier in the chain holds its electrons more
tightly, the highly energetically favorable reaction 2H+ + 2e – + ½O2
→ H2O is made to occur in many small steps. This enables nearly half
of the released energy to be stored, instead of being lost to the
environment as heat.
Spectroscopic Methods Have Been Used to Identify Many Electron
Carriers in the Respiratory Chain
Many of the electron carriers in the respiratory chain absorb visible
light and change color when they are oxidized or reduced. In general,
each has an absorption spectrum and reactivity that are distinct enough
to allow its behavior to be traced spectroscopically, even in crude mixtures.
It was therefore possible to purify these components long before their
exact functions were known. Thus, the cytochromes were discovered
in 1925 as compounds that undergo rapid oxidation and reduction in
living organisms as disparate as bacteria, yeasts, and insects. By observing
cells and tissues with a spectroscope, three types of cytochromes were
identified by their distinctive absorption spectra and designated
cytochromes a, b, and c. This nomenclature has survived, even though
cells are now known to contain several cytochromes of each type and
the classification into types is not functionally important.
The cytochromes constitute a family of colored proteins that are
related by the presence of a bound heme group, whose iron atom
changes from the ferric oxidation state (Fe3+) to the ferrous oxidation
state (Fe2+) whenever it accepts an electron. The heme group consists
of a porphyrin ring with a tightly bound iron atom held by four nitrogen
atoms at the corners of a square (Figure 14-22). A similar porphyrin ring
is responsible for the red color of blood and for the green color of
leaves, being bound to iron in hemoglobin and to magnesium in
chlorophyll, respectively.
http://www.ncbi.nlm.nih.gov/books/NBK26904/bin/ch14f22.jpg
Figure 14-22. The structure of the heme group attached covalently
to cytochrome c.
Figure 14-22
The structure of the heme group attached covalently to cytochrome c.
The porphyrin ring is shown in blue. There are five different
cytochromes in the respiratory chain. Because the hemes in different
cytochromes have slightly different structures and (more…)
Iron-sulfur proteins are a second major family of electron carriers. In these
proteins, either two or four iron atoms are bound to an equal number of
sulfur atoms and to cysteine side chains, forming an iron-sulfur center
on the protein (Figure 14-23). There are more iron-sulfur centers than
cytochromes in the respiratory chain. But their spectroscopic detection
requires electron spin resonance (ESR) spectroscopy, and they are less
completely characterized. Like the cytochromes, these centers carry one
electron at a time.
http://www.ncbi.nlm.nih.gov/books/NBK26904/bin/ch14f23.jpg
Figure 14-23. The structures of two types of iron-sulfur centers.
Figure 14-23
The structures of two types of iron-sulfur centers. (A) A center of the
2Fe2S type. (B) A center of the 4Fe4S type. Although they contain
multiple iron atoms, each iron-sulfur center can carry only one
electron at a time. There are more than seven different (more…)
The simplest of the electron carriers in the respiratory chain—and
the only one that is not part of a protein—is a small hydrophobic
molecule that is freely mobile in the lipid bilayer known as ubiquinone,
or coenzyme Q. A quinone (Q) can pick up or donate either one or
two electrons; upon reduction, it picks up a proton from the medium
along with each electron it carries (Figure 14-24).
http://www.ncbi.nlm.nih.gov/books/NBK26904/bin/ch14f24.jpg
Figure 14-24. Quinone electron carriers.
Figure 14-24
Quinone electron carriers. Ubiquinone in the respiratory chain picks
up one H+ from the aqueous environment for every electron it accepts,
and it can carry either one or two electrons as part of a hydrogen atom
(yellow). When reduced ubiquinone donates (more…)
In addition to six different hemes linked to cytochromes, more than
seven iron-sulfur centers, and ubiquinone, there are also two copper
atoms and a flavin serving as electron carriers tightly bound to respiratory-chain proteins in the pathway from NADH to oxygen. This pathway
involves more than 60 different proteins in all.
As one would expect, the electron carriers have higher and higher
affinities for electrons (greater redox potentials) as one moves along
the respiratory chain. The redox potentials have been fine-tuned
during evolution by the binding of each electron carrier in a particular
protein context, which can alter its normal affinity for electrons. However,
because iron-sulfur centers have a relatively low affinity for electrons,
they predominate in the early part of the respiratory chain; in contrast,
the cytochromes predominate further down the chain, where a higher
affinity for electrons is required.
The order of the individual electron carriers in the chain was
determined by sophisticated spectroscopic measurements (Figure 14-25),
and many of the proteins were initially isolated and characterized as
individual polypeptides. A major advance in understanding the
respiratory chain, however, was the later realization that most of
the proteins are organized into three large enzyme complexes.
http://www.ncbi.nlm.nih.gov/books/NBK26904/bin/ch14f25.gif
Figure 14-25. The general methods used to determine the path of
electrons along an electron-transport chain.
Figure 14-25
The general methods used to determine the path of electrons along
an electron-transport chain. The extent of oxidation of electron
carriers a, b, c, and d is continuously monitored by following their
distinct spectra, which differ in their oxidized and (more…)
The Respiratory Chain Includes Three Large Enzyme Complexes
Embedded in the Inner Membrane
Membrane proteins are difficult to purify as intact complexes
because they are insoluble in aqueous solutions, and some of
the detergents required to solubilize them can destroy normal
protein-protein interactions. In the early 1960s, however, it
was found that relatively mild ionic detergents, such as deoxycholate,
can solubilize selected components of the inner mitochondrial
membrane in their native form. This permitted the identification
and purification of the three major membrane-bound respiratory
enzyme complexes in the pathway from NADH to oxygen (Figure 14-26).
As we shall see in this section, each of these complexes acts as an
electron-transport-driven H+ pump; however, they were
initially characterized in terms of the electron carriers that
they interact with and contain:
http://www.ncbi.nlm.nih.gov/books/NBK26904/bin/ch14f26.gif
Figure 14-26. The path of electrons through the three respiratory
enzyme complexes.
Figure 14-26
The path of electrons through the three respiratory enzyme complexes.
The relative size and shape of each complex are shown. During the
transfer of electrons from NADH to oxygen (red lines), ubiquinone
and cytochrome c serve as mobile carriers that ferry (more…)
The NADH dehydrogenase complex (generally known as complex I)
is the largest of the respiratory enzyme complexes, containing more
than 40 polypeptide chains. It accepts electrons from NADH and
passes them through a flavin and at least seven iron-sulfur centers
to ubiquinone. Ubiquinone then transfers its electrons to a second
respiratory enzyme complex, the cytochrome b-c1 complex.
The cytochrome b-c1 complex contains at least 11 different
polypeptide chains and functions as a dimer. Each monomer
contains three hemes bound to cytochromes and an iron-sulfur
protein. The complex accepts electrons from ubiquinone
and passes them on to cytochrome c, which carries its electron
to the cytochrome oxidase complex.
The cytochrome oxidase complex also functions as a dimer; each
monomer contains 13 different polypeptide chains, including two
cytochromes and two copper atoms. The complex accepts one electron
at a time from cytochrome c and passes them four at a time to oxygen.
The cytochromes, iron-sulfur centers, and copper atoms can carry
only one electron at a time. Yet each NADH donates two electrons,
and each O2 molecule must receive four electrons to produce water.
There are several electron-collecting and electron-dispersing points
along the electron-transport chain where these changes in electron
number are accommodated. The most obvious of these is cytochrome
oxidase.
An Iron-Copper Center in Cytochrome Oxidase Catalyzes Efficient
O2 Reduction
Because oxygen has a high affinity for electrons, it releases a
large amount of free energy when it is reduced to form water.
Thus, the evolution of cellular respiration, in which O2 is
converted to water, enabled organisms to harness much more
energy than can be derived from anaerobic metabolism. This
is presumably why all higher organisms respire. The ability of
biological systems to use O2 in this way, however, requires a
very sophisticated chemistry. We can tolerate O2 in the air we
breathe because it has trouble picking up its first electron; this
fact allows its initial reaction in cells to be controlled closely by
enzymatic catalysis. But once a molecule of O2 has picked up one
electron to form a superoxide radical (O2 -), it becomes dangerously
reactive and rapidly takes up an additional three electrons wherever
it can find them. The cell can use O2 for respiration only because
cytochrome oxidase holds onto oxygen at a special bimetallic
center, where it remains clamped between a heme-linked iron
atom and a copper atom until it has picked up a total of four electrons.
Only then can the two oxygen atoms of the oxygen molecule be
safely released as two molecules of water (Figure 14-27).
Figure 14-27. The reaction of O2 with electrons in cytochrome oxidase.
Figure 14-27
The reaction of O2 with electrons in cytochrome oxidase. As indicated,
the iron atom in heme a serves as an electron queuing point; this
heme feeds four electrons into an O2 molecule held at the bimetallic
center active site, which is formed by the other (more…)
The cytochrome oxidase reaction is estimated to account for 90%
of the total oxygen uptake in most cells. This protein complex is
therefore crucial for all aerobic life. Cyanide and azide are extremely
toxic because they bind tightly to the cell’s cytochrome oxidase
complexes to stop electron transport, thereby greatly reducing
ATP production.
Although the cytochrome oxidase in mammals contains 13
different protein subunits, most of these seem to have a subsidiary
role, helping to regulate either the activity or the assembly of the
three subunits that form the core of the enzyme. The complete
structure of this large enzyme complex has recently been determined
by x-ray crystallography, as illustrated in Figure 14-28. The atomic
resolution structures, combined with mechanistic studies of the effect
of precisely tailored mutations introduced into the enzyme by genetic
engineering of the yeast and bacterial proteins, are revealing the
detailed mechanisms of this finely tuned protein machine.
Figure 14-28. The molecular structure of cytochrome oxidase.
Figure 14-28
The molecular structure of cytochrome oxidase. This protein
is a dimer formed from a monomer with 13 different protein
subunits (monomer mass of 204,000 daltons). The three colored
subunits are encoded by the mitochondrial genome, and they
form the functional (more…)
Electron Transfers Are Mediated by Random Collisions in
the Inner Mitochondrial Membrane
The two components that carry electrons between the three
major enzyme complexes of the respiratory chain—ubiquinone
and cytochrome c—diffuse rapidly in the plane of the inner
mitochondrial membrane. The expected rate of random collisions
between these mobile carriers and the more slowly diffusing
enzyme complexes can account for the observed rates of electron
transfer (each complex donates and receives an electron about
once every 5–20 milliseconds). Thus, there is no need to postulate
a structurally ordered chain of electron-transfer proteins in the
lipid bilayer; indeed, the three enzyme complexes seem to exist as
independent entities in the plane of the inner membrane, being
present in different ratios in different mitochondria.
The ordered transfer of electrons along the respiratory chain
is due entirely to the specificity of the functional interactions
between the components of the chain: each electron carrier is
able to interact only with the carrier adjacent to it in the sequence
shown in Figure 14-26, with no short circuits.
Electrons move between the molecules that carry them in
biological systems not only by moving along covalent bonds
within a molecule, but also by jumping across a gap as large
as 2 nm. The jumps occur by electron “tunneling,” a quantum-
mechanical property that is critical for the processes we are
discussing. Insulation is needed to prevent short circuits that
would otherwise occur when an electron carrier with a low redox
potential collides with a carrier with a high redox potential. This
insulation seems to be provided by carrying an electron deep
enough inside a protein to prevent its tunneling interactions
with an inappropriate partner.
How the changes in redox potential from one electron carrier
to the next are harnessed to pump protons out of the mitochondrial
matrix is the topic we discuss next.
A Large Drop in Redox Potential Across Each of the Three Respiratory
Enzyme Complexes Provides the Energy for H+ Pumping
We have previously discussed how the redox potential reflects
electron affinities (see p. 783). An outline of the redox potentials
measured along the respiratory chain is shown in Figure 14-29.
These potentials drop in three large steps, one across each major
respiratory complex. The change in redox potential between any
two electron carriers is directly proportional to the free energy
released when an electron transfers between them. Each enzyme
complex acts as an energy-conversion device by harnessing some
of this free-energy change to pump H+ across the inner membrane,
thereby creating an electrochemical proton gradient as electrons
pass through that complex. This conversion can be demonstrated
by purifying each respiratory enzyme complex and incorporating
it separately into liposomes: when an appropriate electron donor
and acceptor are added so that electrons can pass through the complex,
H+ is translocated across the liposome membrane.
Figure 14-29. Redox potential changes along the mitochondrial
electron-transport chain.
Figure 14-29
Redox potential changes along the mitochondrial electron-transport
chain. The redox potential (designated E′0) increases as electrons
flow down the respiratory chain to oxygen. The standard free-energy
change, ΔG°, for the transfer (more…)
The Mechanism of H+ Pumping Will Soon Be Understood in
Atomic Detail
Some respiratory enzyme complexes pump one H+ per electron
across the inner mitochondrial membrane, whereas others pump
two. The detailed mechanism by which electron transport is coupled
to H+ pumping is different for the three different enzyme complexes.
In the cytochrome b-c1 complex, the quinones clearly have a role.
As mentioned previously, a quinone picks up a H+ from the aqueous
medium along with each electron it carries and liberates it when it
releases the electron (see Figure 14-24). Since ubiquinone is freely
mobile in the lipid bilayer, it could accept electrons near the inside
surface of the membrane and donate them to the cytochrome b-c1
complex near the outside surface, thereby transferring one H+
across the bilayer for every electron transported. Two protons are
pumped per electron in the cytochrome b-c1 complex, however, and
there is good evidence for a so-called Q-cycle, in which ubiquinone
is recycled through the complex in an ordered way that makes this
two-for-one transfer possible. Exactly how this occurs can now be
worked out at the atomic level, because the complete structure of
the cytochrome b-c1 complex has been determined by x-ray
crystallography (Figure 14-30).
Figure 14-30. The atomic structure of cytochrome b-c 1.
Figure 14-30
The atomic structure of cytochrome b-c 1. This protein is a dimer.
The 240,000-dalton monomer is composed of 11 different protein
molecules in mammals. The three colored proteins form the
functional core of the enzyme: cytochrome b (green), cytochrome (more…)
Allosteric changes in protein conformations driven by electron
transport can also pump H+, just as H+ is pumped when ATP
is hydrolyzed by the ATP synthase running in reverse. For both the
NADH dehydrogenase complex and the cytochrome oxidase complex,
it seems likely that electron transport drives sequential allosteric
changes in protein conformation that cause a portion of the protein
to pump H+ across the mitochondrial inner membrane. A general
mechanism for this type of H+ pumping is presented in Figure 14-31.
Figure 14-31. A general model for H+ pumping.
Figure 14-31
A general model for H+ pumping. This model for H+ pumping
by a transmembrane protein is based on mechanisms that are
thought to be used by both cytochrome oxidase and the light-driven
procaryotic proton pump, bacteriorhodopsin. The protein
is driven through (more…)
H+ Ionophores Uncouple Electron Transport from ATP Synthesis
Since the 1940s, several substances—such as 2,4-dinitrophenol—
have been known to act as uncoupling agents, uncoupling electron
transport from ATP synthesis. The addition of these low-molecular-weight organic compounds to cells stops ATP synthesis by mitochondria
without blocking their uptake of oxygen. In the presence of an
uncoupling agent, electron transport and H+ pumping continue at
a rapid rate, but no H+ gradient is generated. The explanation for
this effect is both simple and elegant: uncoupling agents are lipid-
soluble weak acids that act as H+ carriers (H+ ionophores), and
they provide a pathway for the flow of H+ across the inner mitochondrial
membrane that bypasses the ATP synthase. As a result of this short-
circuiting, the proton-motive force is dissipated completely, and
ATP can no longer be made.
Respiratory Control Normally Restrains Electron Flow
Through the Chain
When an uncoupler such as dinitrophenol is added to cells,
mitochondria increase their oxygen uptake substantially because
of an increased rate of electron transport. This increase reflects
the existence of respiratory control. The control is thought to
act via a direct inhibitory influence of the electrochemical proton
gradient on the rate of electron transport. When the gradient is
collapsed by an uncoupler, electron transport is free to run unchecked
at the maximal rate. As the gradient increases, electron transport
becomes more difficult, and the process slows. Moreover, if an
artificially large electrochemical proton gradient is experimentally
created across the inner membrane, normal electron transport
stops completely, and a reverse electron flow can be detected in
some sections of the respiratory chain. This observation suggests
that respiratory control reflects a simple balance between the
free-energy change for electron-transport-linked proton pumping
and the free-energy change for electron transport—that is, the
magnitude of the electrochemical proton gradient affects both
the rate and the direction of electron transport, just as it affects
the directionality of the ATP synthase (see Figure 14-19).
Respiratory control is just one part of an elaborate interlocking
system of feedback controls that coordinate the rates of glycolysis,
fatty acid breakdown, the citric acid cycle, and electron transport.
The rates of all of these processes are adjusted to the ATP:ADP ratio,
increasing whenever an increased utilization of ATP causes the ratio
to fall. The ATP synthase in the inner mitochondrial membrane,
for example, works faster as the concentrations of its substrates
ADP and Pi increase. As it speeds up, the enzyme lets more H+ flow
into the matrix and thereby dissipates the electrochemical proton
gradient more rapidly. The falling gradient, in turn, enhances the
rate of electron transport.
Similar controls, including feedback inhibition of several key enzymes
by ATP, act to adjust the rates of NADH production to the rate of
NADH utilization by the respiratory chain, and so on. As a result of
these many control mechanisms, the body oxidizes fats and sugars
5–10 times more rapidly during a period of strenuous exercise than
during a period of rest.
Natural Uncouplers Convert the Mitochondria in Brown Fat into
Heat-generating Machines
In some specialized fat cells, mitochondrial respiration is normally
uncoupled from ATP synthesis. In these cells, known as brown fat
cells, most of the energy of oxidation is dissipated as heat rather
than being converted into ATP. The inner membranes of the large
mitochondria in these cells contain a special transport protein that
allows protons to move down their electrochemical gradient, by-
passing ATP synthase. As a result, the cells oxidize their fat stores
at a rapid rate and produce more heat than ATP. Tissues containing
brown fat serve as “heating pads,” helping to revive hibernating animals
and to protect sensitive areas of newborn human babies from the cold.
Bacteria Also Exploit Chemiosmotic Mechanisms to Harness Energy
Bacteria use enormously diverse energy sources. Some, like animal
cells, are aerobic; they synthesize ATP from sugars they oxidize to
CO2 and H2O by glycolysis, the citric acid cycle, and a respiratory
chain in their plasma membrane that is similar to the one in the
inner mitochondrial membrane. Others are strict anaerobes, deriving
their energy either from glycolysis alone (by fermentation) or from an
electron-transport chain that employs a molecule other than oxygen
as the final electron acceptor. The alternative electron acceptor can
be a nitrogen compound (nitrate or nitrite), a sulfur compound
(sulfate or sulfite), or a carbon compound (fumarate or carbonate),
for example. The electrons are transferred to these acceptors by a
series of electron carriers in the plasma membrane that are comparable
to those in mitochondrial respiratory chains.
Despite this diversity, the plasma membrane of the vast majority of
bacteria contains an ATP synthase that is very similar to the one in
mitochondria. In bacteria that use an electron-transport chain to
harvest energy, the electron-transport pumps H+ out of the cell and
thereby establishes a proton-motive force across the plasma membrane
that drives the ATP synthase to make ATP. In other bacteria, the
ATP synthase works in reverse, using the ATP produced by glycolysis
to pump H+ and establish a proton gradient across the plasma
membrane. The ATP used for this process is generated by
fermentation processes (discussed in Chapter 2).
Thus, most bacteria, including the strict anaerobes, maintain a proton
gradient across their plasma membrane. It can be harnessed to drive
a flagellar motor, and it is used to pump Na+ out of the bacterium via
a Na+-H+ antiporter that takes the place of the Na+-K+ pump of
eucaryotic cells. This gradient is also used for the active inward transport
of nutrients, such as most amino acids and many sugars: each nutrient is
dragged into the cell along with one or more H+ through a specific symporter
(Figure 14-32). In animal cells, by contrast, most inward transport across
the plasma membrane is driven by the Na+ gradient that is established by the
Na+-K+ pump.
Figure 14-32. The importance of H+-driven transport in bacteria.
Figure 14-32
The importance of H+-driven transport in bacteria. A proton-motive force
generated across the plasma membrane pumps nutrients into the cell and
expels Na+. (A) In an aerobic bacterium, an electrochemical proton gradient
across the plasma membrane is produced (more…)
Some unusual bacteria have adapted to live in a very alkaline
environment and yet must maintain their cytoplasm at a physiological
pH. For these cells, any attempt to generate an electrochemical H+
gradient would be opposed by a large H+ concentration gradient in
the wrong direction (H+ higher inside than outside). Presumably for
this reason, some of these bacteria substitute Na+ for H+ in all of their
chemiosmotic mechanisms. The respiratory chain pumps Na+ out of
the cell, the transport systems and flagellar motor are driven by an
inward flux of Na+, and a Na+-driven ATP synthase synthesizes
ATP. The existence of such bacteria demonstrates that the principle
of chemiosmosis is more fundamental than the proton-motive force
on which it is normally based.
Summary
The respiratory chain in the inner mitochondrial membrane contains
three respiratory enzyme complexes through which electrons pass on
their way from NADH to O2.
Each of these can be purified, inserted into synthetic lipid vesicles,
and then shown to pump H+ when electrons are transported through it.
In the intact membrane, the mobile electron carriers ubiquinone and
cytochrome c complete the electron-transport chain by shuttling between
the enzyme complexes. The path of electron flow is NADH → NADH
dehydrogenase complex → ubiquinone → cytochrome b-c1 complex →
cytochrome c → cytochrome oxidase complex → molecular oxygen (O2).
The respiratory enzyme complexes couple the energetically favorable
transport of electrons to the pumping of H+ out of the matrix. The
resulting electrochemical proton gradient is harnessed to make ATP
by another transmembrane protein complex, ATP synthase, through
which H+ flows back into the matrix. The ATP synthase is a reversible
coupling device that normally converts a backflow of H+ into ATP
phosphate bond energy by catalyzing the reaction ADP + Pi → ATP,
but it can also work in the opposite direction and hydrolyze ATP to
pump H+ if the electrochemical proton gradient is sufficiently reduced.
Its universal presence in mitochondria, chloroplasts, and procaryotes
testifies to the central importance of chemiosmotic mechanisms in cells.
By agreement with the publisher, this book is accessible by the search
feature, but cannot be browsed.
Copyright © 2002, Bruce Alberts, Alexander Johnson, Julian Lewis,
Martin Raff, Keith Roberts, and Peter Walter; Copyright © 1983, 1989,
1994, Bruce Alberts, Dennis Bray, Julian Lewis, Martin Raff, Keith
Roberts, and James D. Watson .