Sleep science
Larry H. Bernstein,MD, FCAP, Curator
LPBI
Perchance to Dream
Mapping the dreaming brain through neuroimaging and studies of brain damage
| March 1, 2016

Prefrontal leucotomies—surgeries to cut a section of white matter in the front of the brain, thus severing the frontal lobe’s connections to other brain regions—were all the rage through the 1950s as treatments for psychoses. The operations drastically altered the mental state of most patients. But along with personality changes, dulled initiative, and reduced imagination came a seemingly innocuous effect of many of these procedures: the patients stopped dreaming.
Mark Solms, a neuropsychologist at the University of Cape Town in South Africa, uncovered the correlation in historical data from around the globe as part of a long-term study to assess the impact, on dreams and dreaming, of damage to different parts of the brain. Between 1985 and 1995, Solms interviewed 332 of his own patients at hospitals in Johannesburg and London who had various types of brain trauma, asking them about their nightly experiences.
Solms identified two brain regions that appeared critical for the experience of dreaming. The first was at the junction of the parietal, temporal, and occipital lobes—a cortical area that supports spatial cognition and mental imagery. The second was the ventromesial quadrant of the frontal lobes, a lump of white matter commonly associated with goal-seeking behavior that links the limbic structures to the frontal cortex. “This lesion site rang a historical bell in my mind—that’s where the prefrontal leucotomy used to be done,” says Solms, adding that the operation controlled the hallucinations and delusions that came with psychosis. “That sort of struck me as, ‘Gosh, that’s what dreaming is.’” Lesions in other areas could intensify or reduce certain aspects of dreams, but damage to either of the regions Solms pinpointed reportedly caused dreaming to cease completely (Psychoanal Q, 64:43-67, 1995).
Advances in neuroimaging have lent more support to Solms’s brain map, and pinned down other areas that researchers now understand play a part in dream development. In 2013, Bill Domhoff, a psychologist from the University of California, Santa Cruz, and colleagues from the University of British Columbia published results that combined neuroimaging scans from separate studies of REM sleep and daydreaming. They discovered that brain regions that light up when there’s a high chance that one is dreaming overlapped with parts of the brain’s default mode network—regions active when the brain is awake but not focused on a specific external task (Front Hum Neurosci, 7:412, 2013). “It very much lines up,” says Domhoff. “It’s just stunning.”
The default mode network allows us to turn our attention inward, and dreaming is the extreme example, explains Jessica Andrews-Hanna, a cognitive scientist at the University of Colorado Boulder. The network takes up a large amount of cortical real estate. Key players are regions on the midline of the brain that support memories and future planning; these brain sections connect to other areas affecting how we process social encounters and imagine other individuals’ thoughts. “When people are sleeping—in particular, when they’re dreaming—the default mode network actually stays very active,” says Andrews-Hanna. With external stimuli largely cut off, the brain operates in a closed loop, and flights of fancy often ensue.
We usually take the bizarre nature of these experiences at face value. “Even in a completely crazy dream, we all think that it’s normal,” says Martin Dresler, a cognitive neuroscientist at Radboud University in the Netherlands. Dresler and many other researchers attribute this blasé acceptance to the deactivation of a brain region called the dorsolateral prefrontal cortex. When we sleep, the dorsolateral prefrontal cortex powers down, and higher executive control—which would normally flag a nonsensical concern, such as running late for a class when you haven’t been in school for a decade, as unimportant—evaporates. “You have this overactive default mode network with no connectivity, with no communication with regions that are important for making sense of the thoughts,” says Andrews-Hanna.
In healthy sleeping subjects, these executive functions can be unlocked in what’s known as lucid dreaming, when the prefrontal cortex reactivates and sleepers gain awareness of and control over their imagined actions. A lucid dreamer can actually “direct” a dream as it unfolds, deciding to fly, for example, or turning a nightmarish monster into a docile pet.
Records of lucid dreaming are limited to REM sleep, the sleep stage where the brain is most active. REM sleep normally induces paralysis to prevent people from acting out their dreams, but the eye muscles are exempt, and this gives skilled lucid dreamers a way to signal their lucidity to researchers.
Dresler’s team is using this phenomenon as a tool to ask specific questions about dreams. Before trained lucid dreamers fall asleep in Dresler’s lab, they agree to flick their eyes from left to right as soon as they realize within a dream that they’re asleep. The dreamed movement causes their actual eyes to move in a similar way under their closed eyelids. Researchers mark this signal as the beginning of a lucid dream, and then track brain patterns associated with specific dreamed actions. Dreaming also occurs in non-REM sleep, but with the brain less active, the eye muscles won’t respond to dream input—so there’s no robust way to tell if lucid dreaming takes place.
When subjects achieved lucidity and consciously dreamed that they performed a predetermined hand movement, Dresler’s research team observed activity in the sensorimotor cortex matching what would occur if the subjects actually moved their hands while awake (Curr Biol, 21:1833-37, 2011). “It’s probably the case that, for most of what we are dreaming about, the very same machinery and the very same brain regions are active compared to wakefulness,” says Dresler. “It’s just that the motor execution is stopped at the spinal level.”
Beyond sleep research, tracking lucid and normal dreaming offers an investigative model to study aspects of psychosis, according to some researchers. “These regions that are activated during lucid dreaming are typically impaired in patients with psychosis,” explains Dresler. “Having insight into your non-normal mental state in dreaming shares neural correlates with having insights into your non-normal state of consciousness in psychosis.” Dresler proposes training patients in early stages of psychosis to dream lucidly, in the hope that it might grant them some therapeutically relevant understanding of their illness.
While executive functions are impaired in many patients suffering from psychosis, their default networks seem to be overactive, says Andrews-Hanna. But how much similarity exists between the brain states of dreaming and psychosis remains controversial. Domhoff emphasizes the unique nature of dreams. “They’re not like schizophrenia, they’re not like meditation, they’re not like any kind of drug trip,” he says. “They’re an enactment of a scenario that is based upon various wishes and concerns.”
Ultimately, says Solms, deciphering dreaming furthers the field’s knowledge of what the brain does, as much as studies conducted during waking hours. “If you’re a clinician, and you understand what the different parts of the brain do in relation to dreaming, then it’s one of the things you can use as a road map for evaluating your patients.”
Dreamed Movement Elicits Activation in the Sensorimotor Cortex
Martin Dresler1, 5, Stefan P. Koch2, 5, Renate Wehrle1, 5, Victor I. Spoormaker1, et al. Curr Biol.8 Nov 2011; 21(21): 1833–1837 doi:10.1016/j.cub.2011.09.029
Since the discovery of the close association between rapid eye movement (REM) sleep and dreaming, much effort has been devoted to link physiological signatures of REM sleep to the contents of associated dreams [1, 2, 3 and 4]. Due to the impossibility of experimentally controlling spontaneous dream activity, however, a direct demonstration of dream contents by neuroimaging methods is lacking. By combining brain imaging with polysomnography and exploiting the state of “lucid dreaming,” we show here that a predefined motor task performed during dreaming elicits neuronal activation in the sensorimotor cortex. In lucid dreams, the subject is aware of the dreaming state and capable of performing predefined actions while all standard polysomnographic criteria of REM sleep are fulfilled [5 and 6]. Using eye signals as temporal markers, neural activity measured by functional magnetic resonance imaging (fMRI) and near-infrared spectroscopy (NIRS) was related to dreamed hand movements during lucid REM sleep. Though preliminary, we provide first evidence that specific contents of REM-associated dreaming can be visualized by neuroimaging.
Highlights
► Eye signals can be used to access dream content with concurrent EEG and neuroimaging
► Dreamed hand movements correspond to activity in the contralateral sensorimotor cortex
Lucid dreaming is a rare but robust state of sleep that can be trained [5]. Phenomenologically, it comprises features of both waking and dreaming [7]: in lucid dreams, the sleeping subject becomes aware of his or her dreaming state, has full access to memory, and is able to volitionally control dreamed actions [6]. Although all standard polysomnographic criteria of rapid eye movement (REM) sleep [8] are maintained and REM sleep muscle atonia prevents overt motor behavior, lucid dreamers are able to communicate their state by predefined volitional eye movements [6], clearly discernable in the electrooculogram (EOG) (Figure 1). Combining the techniques of lucid dreaming, polysomnography, and brain imaging via functional magnetic resonance imaging (fMRI) or near-infrared spectroscopy (NIRS), we demonstrate the possibility to investigate the neural underpinnings of specific dream contents—in this case, dreamed hand clenching. Predecided eye movements served as temporal markers for the onset of hand clenching and for hand switching. Previous studies have shown that muscle atonia prevents the overt execution of dreamed hand movements, which are visible as minor muscle twitches at most [3 and 9].

Figure 1.
Exemplary Lucid REM Sleep as Captured by Polysomnography during Simultaneous fMRI
Note high-frequency electroencephalogram (EEG) and minimal electromyogram (EMG) amplitude due to muscle atonia characteristic of rapid eye movement (REM) sleep (left), with wakefulness for comparison (right). Subjects were instructed to communicate the state of lucidity by quick left-right-left-right (LRLR) eye movements. Filter settings are as follows: EEG, bandpass filter 0.5−70 Hz, with additional notch filter at 50 Hz; electrooculogram (EOG), bandpass filter 0.1–30 Hz; EMG, bandpass filter 16–250 Hz.

Figure 2.
Comparison of Sensorimotor Activation during Wakefulness and Sleep
Functional magnetic resonance imaging (fMRI) blood oxygen level-dependent (BOLD)-response increases were contrasted between left and right hand movements (columns) in the three conditions (rows): executed hand movement during wakefulness (WE) (A), imagined hand movement during wakefulness (WI) (B), and dreamed hand movement during lucid REM sleep (LD) (C). Effects of left (right) hand movements were calculated in a fixed-effects analysis as a contrast “left > right” and “right > left,” respectively. Subpanels depict results in an SPM glass-brain view (sagital and coronal orientation) to demonstrate the regional specificity of the associated cortical activation, along with sensorimotor activation overlaid on an axial slice of the subject’s T1-weighted anatomical scan (position indicated on the glass brain for condition A). Clusters of activation in the glass-brain views are marked using the numbering given in Table S1. Red outlines in the glass-brain views mark the extent of activation found in the WE condition. This region of interest (ROI) was derived from the respective activation map during executed hand movement (A), thresholded at whole-brain corrected pFWE < 0.005, cluster extent >50 voxels, and served as a ROI for analysis of the WI and LD conditions in (B) and (C), respectively. T values are color-coded as indicated. The time course of the peak voxel inside the ROI is depicted (black) along with the predicted hemodynamic response based on the external pacing (A and B) or the predefined LRLR-eye signals during (C). The maximal difference in activation of the peak voxel between conditions is indicated as percentage of BOLD signal fluctuations of the predicted time course (gray).
FMRI results were confirmed by an independent imaging method in a second subject: NIRS data showed a typical hemodynamic response pattern of increased contralateral oxygenation over the sensorimotor region during successful task performance in lucid REM sleep (Figure 3; Figure 4). Notably, during dreaming, the hemodynamic responses were smaller in the sensorimotor cortex but of similar amplitude in the supplementary motor area (SMA) when compared to overt motor performance during wakefulness.

Figure 3.
Near-Infrared Spectroscopy Topography
Concentration changes of oxygenated (Δ[HbO], upper panel) and deoxygenated hemoglobin (Δ[HbR], lower panel) during executed (WE) and imagined (WI) hand clenching in the awake state and dreamed hand clenching (LD). The optical probe array covered an area of ∼7.5 × 12.5 cm2 over the right sensorimotor area. The solid box indicates the ROI over the right sensorimotor cortex with near-infrared spectroscopy (NIRS)-channels surrounding the C4-EEG electrode position. NIRS channels located centrally over midline and more anterior compared to sensorimotor ROI were chosen as ROI for the supplementary motor area (SMA, dotted box).

Figure 4.
Condition-Related NIRS Time Courses
Time courses of HbO (red traces) and HbR (blue traces) from the right sensorimotor ROI (left panel) and the supplementary motor ROI SMA (right panel) for executed (WE) and imagined (WI) hand clenching in the awake state and dreamed hand clenching (LD). The time courses represent averaged time courses from NIRS channels within the respective ROI (Figure 3). For each condition, 0 s denotes the onset of hand clenching indicated by LRLR-signals. Note that the temporal dynamics, i.e., an increase in HbO and a decrease in HbR, are in line with the typical hemodynamic response. Overt movement during wakefulness (dark red/blue traces) showed the strongest hemodynamic response, whereas the motor-task during dreaming leads to smaller changes (light red/blue traces). In the SMA, the hemodynamic response was stronger during the dreamed task when compared to imagery movement during wakefulness
Neurophysiological studies suggest that during REM sleep, the brain functions as a closed loop system, in which activation is triggered in pontine regions while sensory input is gated by enhanced thalamic inhibition and motor output is suppressed by atonia generated at the brain stem level [4 and 12].
Efforts have been made to correlate REMs to gaze direction during dreams—the “scanning hypothesis” [1 and 2]—and indeed similar cortical areas are involved in eye movement generation in wake and REM sleep [17]. In a similar vein, small muscle twitches during REM sleep were presumed to signal a change in the dream content [3]. Dream research methodology mostly relies on the evaluation of subjective reports of very diverse dream contents.
During dreaming, activation was much more localized in small clusters representing either generally weaker activation or focal activation of hand areas only, with signal fluctuations only in the order of 50% as compared to the actually executed task during wakefulness. The SMA is involved in timing, preparation, and monitoring of movements [21], and linked to the retrieval of a learned motor sequence especially in the absence of external cues [22]. Our NIRS data speak for an activation of SMA even during simple movements. This is in line with several PET and fMRI studies reporting SMA activations for simple tasks such as hand clenching, single finger-tapping, and alternated finger-tapping.
While You Were Sleeping
Assessing body position in addition to activity may improve monitoring of sleep-wake periods.
| March 1, 2016
Polysomnography—the combined assessment of brain waves, heart rate, oxygen saturation, muscle activity, and other parameters—is the most precise way to track a person’s sleeping patterns. However, the equipment required for such analyses is expensive, bulky, and disruptive to natural behavior.
Researchers are thus searching for ways to improve the accuracy of wearable devices while maintaining user-friendliness. Maria Angeles Rol of the University of Murcia in Spain and her colleagues have now discovered that by using a device strapped to the patient’s upper arm that measures both arm activity and position (the degree of tilt), they can more precisely detect periods of sleep.
The researchers studied just 13 people in this pilot study, says Barbara Galland of the University of Otago in New Zealand, but adds that nonetheless it “provide[s] an opening for further investigations to demonstrate the value of this novel technique.” (Chronobiol Int, 32:701-10, 2015)
Validation of an innovative method, based on tilt sensing, for the assessment of activity and body position
M. A. Bonmati-Carriona, B. Middletonb, V. L. Revellb, D. J. Skeneb, M. A. Rola* & J. A. Madrid
Chronobiology International: The Journal of Biological and Medical Rhythm Research 2015; 32,(5) :701-710 PDF http://dx.doi.org:/10.3109/07420528.2015.1016613
Surprisingly small sections of brain, and even neuronal and glial networks in a dish, display many electrical indicators of sleep.
| March 1, 2016 http://www.the-scientist.com/?articles.view/articleNo/45394/title/Sleep-s-Kernel
Sleep is usually considered a whole-brain phenomenon in which neuronal regulatory circuits impose sleep on the brain. This paradigm has its origins in the historically important work of Viennese neurologist Constantin von Economo, who found that people who suffered from brain infections that damaged the anterior hypothalamus slept less. The finding was a turning point in sleep research, as it suggested that sleep was a consequence of active processes within the brain. This stood in stark contrast to the ideas of renowned St. Petersburg physiologist Ivan Pavlov, who believed that sleep resulted from the passive withdrawal of sensory input. Although the withdrawal of sensory input remains recognized as playing a role in sleep initiation, there is now much evidence supporting the idea that neuronal and glial activity in the anterior hypothalamus leads to the inhibition of multiple excitatory neuronal networks that project widely throughout the brain.
But we also know from millions of stroke cases that cause brain damage and from experimentally induced brain damage in animal models that, regardless of where a lesion occurs in the brain, including the anterior hypothalamus, all humans or animals that survive the brain damage will continue to sleep. Further, a key question remains inadequately answered: How does the hypothalamus know to initiate sleep? Unless one believes in the separation of mind and brain, then, one must ask: What is telling the hypothalamus to initiate sleep? If an answer is found, it leads to: What is telling the structure that told the hypothalamus? This is what philosophers call an infinite regress, an unacceptable spiral of logic.
For these reasons, 25 years ago the late Ferenc Obál Jr. of A. Szent-Györgyi Medical University in Szeged, Hungary, and I (J.K.) began questioning the prevailing ideas of how sleep is regulated. The field needed answers to fundamental questions. What is the minimum amount of brain tissue required for sleep to manifest? Where is sleep located? What actually sleeps? Without knowing what sleeps or where sleep is, how can one talk with any degree of precision about sleep regulation or sleep function? A new paradigm was needed.
CHARACTERIZING SLEEP: Sleep-like patterns of neural activity are apparent not just at the level of the whole brain, but also in isolated neural circuits. Researchers have even documented sleep-like behavior in cultures of glial and neural cells. By increasing the number of electrophysiological measurements we use to characterize sleep states, the homology between sleep-like states in culture and sleep in intact animals becomes stronger.
See full infographic: WEB | PDF© CATHERINE DELPHIA
There is no direct measure of sleep, and no single measure is always indicative of sleep. Quiescent behavior and muscle relaxation usually occur simultaneously with sleep but are also found in other circumstances, such as during meditation or watching a boring TV show. Sleep is thus defined in the clinic and in experimental animals using a combination of multiple parameters that typically correlate with sleep.
The primary tool for assessing sleep state in mammals and birds is the electroencephalogram (EEG). High-amplitude delta waves (0.5–4 Hz) are a defining characteristic of the deepest stage of non–rapid eye movement (non-REM) sleep. However, similar waves are evident in adolescents who hyperventilate for a few seconds while wide awake. Other measures used to characterize sleep include synchronization of electrical activity between EEG electrodes and the quantification of EEG delta wave amplitudes. Within specific sensory circuits, the cortical electrical responses induced by sensory stimulation (called evoked response potentials, or ERPs) are higher during sleep than during waking. And individual neurons in the cerebral cortex and thalamus display action potential burst-pause patterns of firing during sleep.
Using such measures, researchers have shown that different parts of the mammalian brain can sleep independently of one another. Well-characterized sleep regulatory substances, or somnogens, such as growth hormone releasing hormone (GHRH) and tumor necrosis factor α (TNF-α), can induce supranormal EEG delta waves during non-REM sleep in the specific half of the rat brain where the molecules were injected. Conversely, if endogenous TNF-α or GHRH production is inhibited, spontaneous EEG delta waves during non-REM sleep are lower on the side receiving the inhibitor. A more natural example of sleep lateralization is found in the normal unihemispheric sleep of some marine mammals. (See “Who Sleeps?”)
Much smaller parts of the brain also exhibit sleep-like cycles. As early as 1949, Kristian Kristiansen and Guy Courtois at McGill University and the Montreal Neurological Institute showed that, when neurons carrying input from the thalamus and surrounding cortical tissue are surgically severed, clusters of neurons called cerebral cortical islands will alternate between periods of high-amplitude slow waves that characterize sleep and low-amplitude fast waves typical of waking, independently of surrounding tissue.1 This suggests that sleep is self-organizing within small brain units.
In 1997, Ivan Pigarev of the Russian Academy of Sciences in Moscow and colleagues provided more-concrete evidence that sleep is a property of local networks. Measuring the firing patterns of neurons in monkeys’ visual cortices as the animals fell asleep while performing a visual task, they found that some of the neurons began to stop firing even while performance persisted. Specifically, the researchers found that, within the visual receptive field being engaged, cells on the outer edges of the field stopped firing first. Then, as the animal progressed deeper into a sleep state, cells in more-central areas stopped firing. This characteristic spatial distribution of the firing failures is likely a consequence of network behavior. The researchers thus concluded that sleep is a property of small networks.2
More recently, David Rector at Washington State University and colleagues provided support for the idea of locally occurring sleep-like states. In a series of experiments, they recorded electrical activity from single cortical columns using a small array of 60 electrodes placed over the rat somatosensory cortex. The sensory input from individual facial whiskers maps onto individual cortical columns. As expected, ERPs in the cortical columns induced by twitching a whisker were higher during sleep than during waking. But looking at the activity of individual columns, the researchers observed that they could behave somewhat independently of each other. When a rat slept, most—but not all—of the columns exhibited the sleep-like high-amplitude ERPs; during waking, most—but not all—of the columns were in a wake-like state. Interestingly, the individual cortical columns also exhibited patterns that resembled a sleep rebound response: the longer a column was in the wake-like state, the higher the probability that it would soon transition into a sleep-like state.3
To test how cortical-column state can affect whole-animal behavior, Rector and his team trained rats to lick a sucrose solution upon the stimulation of a single whisker, then characterized the whisker’s cortical-column state. If the column receiving input from the stimulated whisker was in a wake-like state (low-magnitude ERP), the rats did not make mistakes. But if the column was in the sleep-like state (high-magnitude ERP), the animals would fail to lick the sucrose when stimulated and would sometimes lick it even when their whisker was not flicked.4 Even though the animal was awake, if a cortical column receiving stimulation was asleep, it compromised the animal’s performance. These experiments indicate that even very small neuronal networks sleep and that the performance of learned behavior can depend on the state of such networks.
Given that sleep can manifest in relatively small brain regions, perhaps it should not be too surprising that co-cultures of neurons and glia possess many of the electrophysiological sleep phenotypes that are used to define sleep in intact animal brains. During sleep, cortical and thalamic neurons display bursts of action potentials lasting about 500 ms, followed by periods of hyperpolarization lasting about the same length of time. The synchronization of this firing pattern across many neurons is thought to generate the EEG activity characteristic of delta-wave sleep, and undisturbed co-cultures of glia and neurons display periodic bursts of action potentials, suggesting that the culture default state is sleep-like. In contrast, if neuronal and glia networks are stimulated with excitatory neurotransmitters, the culture’s “burstiness”—the fraction of all action potentials found within bursts—is reduced, indicating a transition to a wake-like state. Treatment of co-cultures with excitatory neurotransmitters also converts their gene expression profile from a spontaneous sleep-like pattern to a wake-like pattern.5

SLEEP IN VITRO: Neurons co-cultured with glial cells display patterns of action potentials and slow (delta) waves, suggesting that small neural networks can and do sleep, even outside of the body. In culture, neurons fire in bursts, and slow-wave electrical activity is synchronized while in a default sleep-like state. However, if the culture is stimulated with electricity or excitatory neurotransmitters, delta-wave amplitude and the neurons’ synchrony, or burstiness, are reduced, suggesting that the culture “wakes up.” Conversely, the addition of TNF-α, a sleep-inducing agent, increases burstiness and the amplitudes of delta waves.
See full infographic: WEB | PDF© CATHERINE DELPHIA
Cell cultures also respond to sleep-inducing agents similarly to whole organisms. If a neuronal and glial culture is treated with TNF-α, the synchronization and amplitudes of slow-wave electrical activity increase, indicating a deeper sleep-like state. Moreover, ERPs are of greater magnitude after cultures are treated with TNF-α than during the sleep-like default state, suggesting that the somnogen induces a deeper sleep-like state in vitro as it does in vivo.6
Researchers have even studied the developmental pattern of such sleep phenotypes, using multielectrode arrays to characterize network activity throughout the culture, and the emergence of network properties follows a similar time course as in intact mouse pups. Spontaneous action potentials occur during the first few days in culture, but network emergent properties are not evident until after about 10 days. Then, synchronization of electrical potentials begins to emerge, and the network’s slow waves begin to increase in amplitude. If the cultures are electrically stimulated, slow-wave synchronization and amplitudes are reduced, suggesting the networks wake up. This is followed by rebound-enhanced slow-wave synchronization and amplitudes the next day, suggesting sleep homeostasis is also a characteristic of cultured networks.6
Clearly, even small neural networks can exhibit sleep-like behavior, in a dish or in the brain. But the question remains: What is driving the oscillations between sleep- and wake-like states?
Sleep emerges
In the intact brain, communication among neurons and between neurons and other cells is ever changing. Bursts of action potentials trigger the release of multiple substances and changes in gene expression, both of which alter the efficacy of signal transmission. For instance, neural or glial activity induces the release of ATP into the local extracellular space. Extracellular ATP, in turn, induces changes in the expression of TNF-α and other somnogens known to induce a sleep-like state. Because these effects take place in the immediate vicinity of the cell activity, they target sleep to local areas that were active during prior wakefulness.
In 1993, Obál and I (J.K.) proposed that sleep is initiated within local networks as a function of prior activity.7 The following year, Derk-Jan Dijk and Alex Borbely of the University of Zurich provided support for this idea when they had volunteers hold hand vibrators in one hand during waking to stimulate one side of the somatosensory cortex. In subsequent sleep, the side of the brain that received input from the stimulated hand exhibited greater sleep intensity, determined from amplitudes of EEG slow waves, than the opposite side of the brain. And in 2006, Reto Huber, then at the University of Wisconsin, showed that if an arm is immobilized during waking, amplitudes of EEG slow waves from the side of the brain receiving input from that arm are lower in subsequent sleep.
These experiments indicate that local sleep depth is a function of the activity of the local network during waking—an idea that has been confirmed by multiple human and animal studies. Moreover, local network state oscillations strongly indicate that sleep is initiated within local networks such as cortical columns. But how do the states of a population of small networks translate into whole-animal sleep?
Small local clusters of neurons and glia are loosely connected with each other via electrophysiological and biochemical signaling, allowing for constant communication between local networks. Steven Strogatz of Cornell University showed that dynamically coupled entities, including small neuronal circuits, will synchronize with each other spontaneously without requiring direction by an external actor. Synchronization of loosely coupled entities occurs at multiple levels of complexity in nature from intact animals to molecules—for example, birds flocking, or the transition from water to ice. The patterns generated by bird flocking, or the hardness of ice, are called emergent properties.
We, Obál, and our colleagues proposed that whole-brain sleep is an emergent property resulting from the synchronization of local neuronal network states.7,8,9 This would explain why sleep continues to occur after brain damage: because the remaining local circuits will spontaneously synchronize with each other. This view also allows one to easily envision variations in the depth or degree of sleep and waking because it allows for some parts of the brain to be in sleep-like states while other areas are in wake-like states, just as Rector observed. These independent states of local networks may account for sleep inertia, the minutes-long period upon awakening of poor cognitive performance and fuzzy-mindedness, and may also play a role in the manifestation of dissociated states such as sleepwalking. Most importantly, this paradigm frees sleep regulation from the dualism trap of mind/brain separation: top-down imposition of state is not required for the initiation of local state oscillations or for subsequent whole-organism sleep to ensue.
Our theory is also consistent with the modulation of sleep and wakefulness by sleep regulatory circuits such as those in the hypothalamus. For example, if interleukin-1, a sleep regulatory substance, is applied locally to the surface of the rat cortex, it induces local high-amplitude EEG slow waves indicative of a greater local depth of sleep.10 The responses induced by interleukin-1 in the cortex enhanced neuronal activity in anterior hypothalamic sleep regulatory areas.11 That hypothalamic neuronal activity likely provides information on local sleep- and wake-like states occurring in the cortex to the hypothalamus, where it can modulate the orchestration of the sleep initiated within the smaller brain units.
Finally, our ideas may inform the study of how sleep influences the formation of memories. A fundamental problem a living brain faces is the incorporation of new memories and behaviors while conserving existing ones. We know that cell activity enhances neuronal connectivity and the efficacy of neurotransmission within active circuits, a phenomenon that has been posited to be a mechanism by which memories are formed and solidified. By themselves, however, these use-dependent mechanisms would lead to unchecked growth of connectivity (in response to activity patterns) and positive feedback (since increased connectivity leads to reuse), ultimately resulting in a rigid, non-plastic network.7 Instead, we suggest that biochemical mechanisms—specifically, the use-dependent expression of genes involved in sleep regulation and memory—induce oscillations, representing local wake- and sleep-like states, which serve to stabilize and preserve brain plasiticity.7
For more than a century, researchers have struggled to understand how sleep works and what it does. Perhaps this lack of answers stems from a fundamental misconception about what sleeps. By thinking about sleep in smaller units, such as individual networks in the brain, hopefully the field will start to understand what exactly is going on during this enigmatic—but very common—phenomenon.
James M. Krueger is a regents professor of neuroscience and Sandip Roy is an associate professor of electrical engineering at Washington State University.
References
- K. Kristiansen, G. Courtois, “Rhythmic electrical activity from isolated cerebral cortex,” Electroen Clin Neuro, 1:265-72, 1949.
- I.N. Pigarev et al., “Evidence for asynchronous development of sleep in cortical areas,” Neuroreport, 8:2557-60, 1997.
- D.M. Rector et al., “Local functional state differences between rat cortical columns,” Brain Res, 1047:45-55, 2005.
- J.M. Krueger et al., “Sleep: A synchrony of cell activity-driven small network states,” Eur J Neurosci, 38:2199-09, 2013.
- V. Hinard et al., “Key electrophysiological, molecular, and metabolic signatures of sleep and wakefulness revealed in primary cortical cultures,” J Neurosci, 32:12506-17, 2012.
- K.A. Jewett et al., “Tumor necrosis factor enhances the sleep-like state and electrical stimulation induces a wake-like state in co-cultures of neurons and glia,” Eur J Neurosci, 42:2078-90, 2015.
- J.M. Krueger, F. Obál, “A neuronal group theory of sleep function,” J Sleep Res, 2:63-69, 1993.
- J.M. Krueger et al., “Sleep as a fundamental property of neuronal assemblies,” Nat Rev Neurosci, 9: 910-19, 2008.
- S. Roy et al., “A network model for activity-dependent sleep regulation,” J Theor Biol, 253:462-68, 2008.
- T. Yasuda et al., “Interleukin-1 beta has a role in cerebral cortical state-dependent electro-encephalographic slow-wave activity,” Sleep, 28:177-84, 2005.
- K. Yasuda et al., “Unilateral cortical application of interleukin-1β (IL1β) induces asymmetry in Fos- and IL1β-immunoreactivity: Implications for sleep regulation,” Brain Res, 1131:44-59, 2007.
In Dogged Pursuit of Sleep
Unearthing the root causes of narcolepsy keeps Emmanuel Mignot tackling one of sleep science’s toughest questions.
| March 1, 2016 http://www.the-scientist.com/?articles.view/articleNo/45347/title/In-Dogged-Pursuit-of-Sleep
In November 1986, Emmanuel Mignot arrived at Stanford University’s Center for Sleep Sciences and Medicine for a 16-month stint as a research associate. His goal was to find effective drugs to treat narcolepsy; his study subjects belonged to a colony of canines that suffered from the malady. “[When I got there], the dogs were being maintained, but not much was being done with them other than some chemistry studies on known neurotransmitters,” says Mignot, a professor of psychiatry and behavioral sciences at Stanford University and now director of the center. “As a pharmacologist, I wanted to study potential treatments for narcolepsy and understand the molecular biology to improve treatment in humans.”
The first narcoleptic dog, a French poodle named Monique, was brought to Stanford in 1974 byWilliam Dement, the so-called “father of sleep medicine,” who had founded the center in 1970, the first in the world dedicated to the study of sleep. Dement and other researchers there established a full breeding colony in 1977 when dogs with a genetic form of the neurological disorder were discovered—initially, some puppies from a litter of Dobermans and, later, some Labradors. Narcoleptic dogs and humans both exhibit a combination of symptoms: perpetual sleepiness, cataplexy—muscle paralysis attacks triggered by emotions—and abnormal rapid eye movement (REM) sleep. While the condition in humans and dogs is treatable, there is no cure.
To study which narcolepsy drugs increased wakefulness and decreased cataplexy in the dogs, Mignot and psychiatry professor Seiji Nishino used a food-elicited cataplexy test: administration of the drug followed by release into a room with pieces of food on the floor and careful observation. “The dog would rush into the room and be so happy to eat the treats, and then would have an attack and collapse on the floor.” The researchers counted the number and duration of the attacks after treatment with a drug at various doses. In humans, cataplexy episodes are triggered by a positive emotion such as laughter at a joke or pleasant surprise. “For the dogs, it is food or the joy of playing. That is what is great about dogs as a model for this condition. When you give a treatment to a rat or mouse and they stop having cataplexy, you really don’t know if it is because they don’t feel good or if it is a genuine effect. But the dogs show you emotions like humans. I knew all of these dogs by name. They were my friends. I could see if they were worried or didn’t feel well.”
Mignot worked mostly with the Dobermans and Labs, but there were also dogs donated to the colony that seemed to have a sporadic form of narcolepsy, “There was Vern, a miniature poodle; Wally, a big poodle; Tucker, a mutt; and Beau, my beloved dachshund.” Using the cataplexy test in animals along with in vitro studies of the drugs’ chemical properties, Mignot and Nishino found that antidepressants suppress cataplexy by inhibiting adrenergic reuptake, and that amphetamine-like stimulants promote wakefulness in narcoleptics by increasing the availability of dopamine. “We improved the then-current treatments and started to understand the kinds of chemicals important to regulate narcolepsy symptoms.”
But Mignot wanted to understand the molecular mechanism of narcolepsy, so he turned his focus to the genetic basis of the disorder. A lack of genetics training and no map of the dog genome to guide him did not deter Mignot. He has tirelessly pursued this previously little-studied and, so far, only known neurological disorder that fundamentally perturbs the nature of sleep states.
Here, Mignot talks about pursuing a master’s, PhD, and MD simultaneously, the paper retraction that has been the most difficult episode in his career so far, and his unexpected devotion to a Chihuahua.
Mignot Motivated
Sir Mix-a-Lot. The youngest of six siblings, Mignot had a penchant for collecting fossils and for conducting chemistry experiments in the bathroom of his family’s home in Paris. “I bought chemicals sold by a Chinese shopkeeper on Rue Saint-Dominique to do all kinds of experiments, mixed them, and occasionally made mistakes. There were burn marks and projections on the walls of my bathroom.” In high school, the self-proclaimed “nerd with glasses” became interested in biology, and, after graduation in 1977, went to study for a medical degree at the René Descartes University Faculty of Medicine in Paris.
Collecting degrees. “In the second year of medical school, I got bored from all of the memorization.” He took the entrance exam for the prestigious École Normale Supérieure (ENS), which gives students freedom to pursue their academic interests at other institutions while providing a stipend, housing, and the support of professor mentors. He passed, and entered the ENS in 1979. Mignot worked towards a master’s in biochemistry, and then a PhD in molecular pharmacology while still continuing his medical studies. “Nothing was set up for MD-PhD programs at the time. It was all in parallel, which was crazy. I had an exam every few weeks,” says Mignot. In 1984, he received both his medical degree and, later, a PhD from Pierre and Marie Curie University.
New to narcolepsy. Mignot became interested in the effects of drugs on the brains of psychiatric patients, studying how different compounds affected the metabolism of neurotransmitters in the brains of rats, and pursued a residency in psychiatry to complement his laboratory research. In 1986, he was offered a professorship in pharmacology at the Paris V University School of Medicine. But first, Mignot needed to complete the mandatory military service that he had deferred. “Instead of going to a former French colony to practice medicine, I convinced the French government to send me to Stanford to study modafinil, a wakefulness-promoting drug created by a French pharmaceutical company called Lafon Laboratories for the treatment of narcolepsy. I had never heard about [narcolepsy] during medical school—it must have been a single line in my textbooks. I discovered that Stanford was doing work on sleep and that Dement had started a colony of narcoleptic dogs there. I thought I could study these animals and figure out how modafinil worked.”
So Mignot came to Stanford for 16 months as part of his military service with financial support from Lafon Laboratories. “The company had claimed modafinil worked by a novel mechanism, unrelated to how stimulants work,” says Mignot. But Mignot found that modafinil bound the dopamine transporter, inhibiting the reuptake of the neurotransmitter, boosting wakefulness. “This is a similar mode of action as Ritalin, but the company was claiming otherwise. It took 10 years for my results to be validated, finally, by Nora D. Volkow, now director of the National Institute on Drug Abuse, who showed . . . that indeed the drug displaces the dopamine transporter at doses that increase wakefulness in humans.”
Mignot Moves
Going to the dogs. At Stanford, Mignot immersed himself in his work with the dog colony. “I worked all the time and came home just to sleep. I was definitely not very successful with girls then, because I smelled like dog all the time. I spent all day with the dogs, going to the facility, hugging, playing, and working with them. When we bred them, sometimes the mothers rejected their puppies so we had to come in every few hours, even in the middle of the night, to bottle-feed the puppies. Even after I took a shower, you could still smell the dogs. It was a strange part of my life.”
From pharmacology to genetics. Mignot kept extending his stay at Stanford. “After a few years I realized our pharmacology studies were never going to lead to narcolepsy’s cause. We needed to find the genetic cause in the dog.” In 1988, he resigned a faculty position in Paris—which was being held for him even as he continued to extend his time at Stanford—deciding to search for the mutated gene responsible for narcolepsy in dogs. In 1993, Mignot became the head of the Center for Narcolepsy at Stanford. A connection between an immune gene, the human leukocyte antigen (HLA) allele HLA-DR2, and narcolepsy in humans had already been identified by Yutaka Honda at the University of Tokyo, so Mignot’s lab tried to ascertain whether the same connection was true in the dogs or if the immune gene was simply a genetic linkage marker. These were the days before the dog or human genome had been sequenced, so the work took Mignot’s lab 10 years, and almost 200 narcoleptic Dobermans and Labradors: years of painstaking chromosome walking experiments, DNA fingerprinting, and the construction of a bacterial artificial chromosome library of dog genomic pieces. “What helped us a lot was that we knew the Dobermans and Labs had the same genetic defect because we interbred and got narcoleptic puppies—what’s called a complementation test.” In 1999, Mignot’s team identified the mutated gene as hypocretin receptor 2, whose protein binds hypocretin (also called orexin), a neuropeptide that regulates arousal and wakefulness. Several weeks later, after seeing these findings, Masashi Yanagisawa’s lab independently published a confirmation, showing that hypocretin knockout mice also have narcolepsy.
In parallel narcolepsy studies across ethnic groups, Mignot’s lab found that it was not the initial HLA-DR2allele that predisposed humans to narcolepsy, but another, nearby HLA gene, DQB1*0602.
Humans are not like dogs. “After we found the gene, the research went fast. We decided to look at hypocretin itself and see if it’s abnormal in humans.” Mignot’s lab sequenced the genes for the hypocretin receptor and its ligand in narcoleptic patients, expecting mutations in either to be rare because of the known HLA-narcolepsy linkage and the fact that most cases in humans, unlike in dogs, are not familial. Only one documented case, a child who had narcolepsy onset at six months of age, has been found to harbor a hypocretin gene mutation. “I think you need to knock out both receptor 1 and 2 in humans to get the full narcoleptic phenotype,” says Mignot. “Those with just one mutation may be more prone to tiredness but not full narcolepsy.”
In 2000, Mignot’s and Nishino’s groups reported that hypocretin was not present in narcoleptic patients’ cerebrospinal fluid—a test still used diagnostically today. The same year, independent studies from Mignot’s laboratory and that of Jerome Siegel at the University of California, Los Angeles, found that the lack of hypocretin was not due to gene mutations but to the fact hypocretin cells were missing in the brains of narcoleptic patients. HLA genes were well known to be associated with many autoimmune diseases, and Mignot hypothesized that hypocretin was missing due to an autoimmune attack against hypocretin-secreting neurons. What the abnormality is in those narcolepsy patients with normal hypocretin levels remains a mystery.
Mignot Moves Forward
Still a missing link. “I have been working on this [autoimmunity] hypothesis for 10 years, and we see that this hypothesis is more and more likely, but we cannot find any direct proof. It’s frustrating, but that kind of struggle is the story of my life.” All known autoimmune diseases result in the generation of antibodies in patients, but antibodies against hypocretin or the hypocretin cells have never been detected. So Mignot’s lab tested whether T-cells were the immune component attacking hypocretin. In 2013, his lab published a study identifying the T-cell culprits. But the study was retracted by Mignot himself one year later, when Mignot’s group couldn’t reproduce the results after the scientist who did most of the experiments had left the lab. “It was really painful and the worst time in my career.”
A new lead. “In 2010, a lot of people suddenly started to develop narcolepsy after receiving the Pandemrix vaccine against swine flu. It’s very odd. We still don’t understand why this particular vaccine increased the risk of narcolepsy.” Mignot thinks that a component of the vaccine or the virus itself triggers the immune system to attack hypocretin-producing neurons. “So now I am doing a lot of studies comparing the different vaccines and the wild-type virus to try to understand what could be common to produce this response. I think the vaccine will give us a final clue to isolate the immune T-cells involved in narcolepsy.”
Genetics of sleep. Mignot’s lab is working on a genome-wide association study, which shows that the genetic variants linked to narcolepsy are mostly immune-related, similar to Type 1 diabetes, celiac disease and other autoimmune diseases, further supporting the autoimmune hypothesis. Mignot is also getting a large human study off the ground. “I want to study the genetics of 40,000 people with sleep issues to see if there are genetic traits that cause people to sleep well or not sleep well, to need more sleep or less sleep. This hasn’t been done yet. I think this will help us crack open the mysteries of sleep.”
A new companion. “The dog colony was officially dismantled in 2000 after we found the canine narcolepsy gene. The dogs were adopted and we got Bear, a narcoleptic Schipperke. He passed away over a year ago. I loved that dog and miss him a lot. He was an unusually kind soul. Three months later, a breeder from Vermont called and said he had a narcoleptic Chihuahua. I flew to Vermont and adopted Watson and he’s been with us ever since. I never would have thought to adopt a Chihuahua, but now I can’t think of life without Watson. He is faithful and cuddly. I really think you can bond with any dog.”
The journey continues. “This story of narcolepsy, it’s a difficult story. Finding the gene was very difficult, and finding the autoimmune connection should have been trivial, but it has been an ordeal because there is absolutely no collateral damage. As [Stanford neurologist] Larry Steinman said to me, it’s like a ‘hit and run’—it looks like it was cleaned up and the players disappear. It’s hard, but by learning about this disease, we may discover other diseases where a similar autoimmune destruction happens in the brain but we have never realized it. I wouldn’t be surprised if some forms of depression and schizophrenia have an autoimmune basis in the brain. By experience, the more difficult it is, the most interesting the answer will be.”
Greatest Hits
- Identified the gene for hypocretin receptor 2, which, when mutated, causes an inherited form of narcolepsy in Dobermans and Labradors
- Identified how antidepressant and stimulant drugs work as treatments for narcolepsy
- Identified DQB1*0602 as the main human gene associated with narcolepsy
- By genome-wide association, found immune polymorphisms, such as one in the T-cell receptor alpha, that also predispose people to the disease, further suggesting the disease is autoimmune
- Found that human narcolepsy, unlike canine narcolepsy, is not caused by mutations in the hypocretin receptor 2 gene but is due to an immune-mediated destruction of hypocretin-producing neurons in the brain
DQB1*0602 and DQA1*0102 (DQ1) are better markers than DR2 for narcolepsy in Caucasian and black Americans.
Sleep consumes almost one-third of any human lifetime, yet its biological function remains unknown. Electrophysiological studies have shown that sleep is physiologically heterogeneous. Sleep onset is first characterized by light nonrapid eye movement (NREM) sleep (stage I and II), followed by deep NREM sleep or slow-wave sleep (stage III and IV) and finally rapid eye movement (REM) sleep. This sleep cycle is ∼90 min long and is repeated multiple times during nocturnal sleep. REM sleep, also called paradoxical sleep, is characterized by low-voltage fast electroencephalogram activity, increased brain metabolism, skeletal muscle atonia, rapid eye movements, and dreaming. Total sleep deprivation and/or REM sleep deprivation are both lethal in animals.
NREM and REM sleep are mainly regulated by circadian and homeostatic processes. Recent studies have suggested that across the animal kingdom, circadian rhythms are regulated by similar negative feedback loops involving the rhythmic expression of RNAs encoding proteins that act to shut off the genes encoding them (Hall 1995; Dunlap 1996;Rosbash et al. 1996; Young et al. 1996). From a genetic perspective, much less progress has been made in the noncircadian aspects of sleep regulation. This review demonstrates that a genetic approach to narcolepsy will in time provide a novel insight into the molecular basis of sleep control.
Narcolepsy most often begins in the second decade of life but may be observed at the age of 5 or younger (Honda 1988). The cardinal symptom in narcolepsy is a persistent and disabling excessive daytime sleepiness. Sleep attacks are unpredictable, irresistible, and may lead to continuing activities in a semiconscious manner, a phenomenon referred to as automatic behavior. Naps are usually refreshing, but the restorative effect vanishes quickly.
Sleepiness is not sufficient to diagnose the disorder. Narcoleptic patients also experience symptoms that are secondary to abnormal transitions to REM sleep (Aldrich 1992; Bassetti and Aldrich 1996). The most important of these symptoms is cataplexy, a pathognomonic symptom for the disorder. In cataplexy, humor, laughter, or anger triggers sudden episodes of muscle weakness ranging from sagging of the jaw, slurred speech, buckling of the knees or transient head dropping, to total collapse to the floor (Aldrich 1992; Bassetti and Aldrich 1996). Patients typically remain conscious during the attack, which may last a few seconds or a few minutes. Reflexes are abolished during the attack, as they are during natural REM sleep atonia. Sleep paralysis, another manifestation of REM sleep atonia, is characterized by an inability to move and speak while falling asleep or upon awakening. Episodes last a few seconds to several minutes and can be very frightening. Hypnagogic hallucinations are vivid perceptual dream-like experiences (generally visual) occurring at sleep onset. Sleep paralysis and hypnagogic hallucinations occasionally occur in normal individuals under extreme circumstances of sleep deprivation or after a change in sleep schedule (Aldrich 1992; Bassetti and Aldrich 1996) and thus have little diagnostic value in isolation.
Nocturnal sleep polysomnography is conducted to exclude other possible causes of daytime sleepiness such as sleep apnea or periodic limb movements (Aldrich 1992). The Multiple Sleep Latency Test (MSLT) is also carried out to demonstrate daytime sleepiness objectively. In this test, patients are requested to take four or five naps at 2-hr intervals, during which time to sleep onset (sleep latency) is measured. Short sleep latencies under 5 min are usually observed in narcoleptic patients, together with abnormal REM sleep episodes, referred to as sleep-onset REM periods (SOREMPs). The combination of a history of cataplexy, short sleep latencies, and two or more SOREMPs during MSLT is diagnostic for narcolepsy (Bassetti and Aldrich 1996;Mignot 1996). Note that many naps consist only of NREM sleep suggesting that there is also a broader problem of impaired sleep–wake regulation, with indistinct boundaries between sleep and wakefulness in narcolepsy (Broughton et al. 1986; Bassetti and Aldrich 1996).
The disorder has a large psychosocial impact. Two-thirds of patients have fallen asleep while driving, and 78% suffer from reduced performance at work (Broughton et al. 1981). Depression occurs in up to 23% of cases (Roth 1980). Treatment is purely symptomatic and generally involves amphetamine-like stimulants for excessive daytime sleepiness and antidepressive treatment for cataplexy and other symptoms of abnormal REM sleep (Bassetti and Aldrich 1996; Nishino and Mignot 1997).
Narcolepsy–cataplexy affects 0.02%–0.18% of the general population in various ethnic groups (Mignot 1998). A familial tendency for narcolepsy has long been recognized (Roth 1980). The familial risk of a first-degree relative is 0.9%–2.3% for narcolepsy–cataplexy, which is 10–40 times higher than the prevalence in the general population (Mignot 1998).
In a Finnish twin cohort study consisting of 13,888 monozygotic (MZ) and same-sexed dizygotic (DZ) twin pairs, three narcoleptic individuals were found and each of them was discordant DZ with a negative family history (Hublin et al. 1994). In the literature, 16 MZ pairs with at least one affected twin have been reported and five of these pairs were concordant for narcolepsy (Mignot 1998). Although narcolepsy is likely to have a genetic predisposition, the low rate of concordance in narcoleptic MZ twins indicates that environmental factors play an important role in the development of the disease.
Narcolepsy was shown to be associated with the human leukocyte antigen (HLA) DR2 in the Japanese population (Honda et al. 1984;Juji et al. 1984). DR2 is observed in all Japanese patients versus 33% of Japanese controls (Juji et al. 1984; Matsuki et al. 1988a). A similar association is observed in Caucasians, with >85% versus 22% DR2 positivity (Langdon et al. 1984; Billiard et al. 1986;Rogers et al. 1997). Strikingly however, the DR2association is much lower in African–Americans (65%–67% in narcoleptic patients vs. 27%–38% in controls) (Neely et al. 1987;Matsuki et al. 1992;Rogers et al. 1997). Further studies have shown that HLA DQalleles, located ∼80 kb from the DRregion, are more tightly associated with narcolepsy than HLADR subtypes. More than 90% of narcolepsy–cataplexy patients across all ethnic groups carry a specific allele of HLA DQB1, DQB1*0602 (Matsuki et al. 1992;Mignot et al. 1994); this allele is present in 12%–38% of the general population across many ethnic groups (Matsuki et al. 1992; Mignot et al. 1994; Lin et al. 1997).DQB1*0602 is associated almost exclusively with DR2in Japanese (Lin et al. 1997) and Caucasians (Begovich et al. 1992), whereas it is observed frequently in association with DR2, DR5, or other DRsubtypes in African–Americans (Mignot et al. 1994, 1997a). The increased DR–DQ haplotypic diversity in African–Americans explains the low DR2 association observed in this population.
To further characterize the DQB1 region in narcoleptic subjects, novel polymorphic markers were isolated and characterized (Mignot et al. 1997a). The markers tested included six novel microsatellite markers (DQCAR, DQCARII, G51152, DQRIV, T16CAR, and G411624R). DQA1, a DQ gene whose product is known to pair with DQB1-encoding polypeptides to form the biologically active DQ heterodimer molecule, was also studied. The results obtained are summarized in Figure1. The association with narcolepsy decreases in theT16CAR–DQB2 region (Mignot et al. 1997a) and in the DRB1 region (Mignot et al. 1994, 1997b). The G411624R andT16CAR microsatellites are complex repeats with drastically different sizes, all of which are frequently observed in narcolepsy susceptibility haplotypes, a result suggesting crossovers in the region. In the DRB1 region, association with narcolepsy is still tight with DRB1*1501 (DR2) in Caucasians and Asians but is significantly lower in African–Americans, which suggests crossovers in the region among ethnic groups.

Figure 1.
Schematic summary of the narcolepsy susceptibility region within the HLA complex. Genes and markers are depicted by vertical bars, alleles observed in narcoleptic patients are listed above each marker.DQB2, DQB3, DQB1, DQA1, andDRB1 are HLA genes and pseudogenes. QBP and QAP are the promoter regions ofDQB1 and DQA1, respectively. G411624R, T16CAR, G51152, DQCAR, and DQCARII are microsatellite CA repeats identified in the HLA DQ region (Mignot et al. 1997a).DQRIV is a compound tandem repeat of 4- and 2-bp units located between DQB1 and G51152. TheDQA1*0102allele is subdivided into 01021 and 01022 based on a codon 109 synonymous substitution. Genomic segments in which frequent recombination was detected are indicated by vertical solid lines. Broken lines indicate rare possible ancestral crossovers detected in the area. Crossovers betweenT16CAR and G51152 occur within ethnic groups; crossovers between QAP and DRB1are frequently observed among ethnic groups (Mignot et al. 1997a). Note that the genomic region shared by most narcoleptic patients extends from a region between T16CAR and G51152 to a region between QAP andDRB1. No other genes were found in 86 kb of genomic sequence surrounding the DQB1*0602 gene (Ellis et al. 1997). Additional diversity is also found at the level ofG51152 andDQRIV, this being most likely due to a slippage mechanism rather than crossover (Lin et al. 1997; Mignot et al. 1997a). (+, Δ, *) Frequent alleles found predominantly in Caucasian, Asian, and African–American populations, respectively; (kb) kilobase pairs. Alleles frequently observed in theDQB1*0602/DQA1*0121 haplotype are underlined.DRB1*1501, DRB1*1503, and DRB1*1602 are DR2subtypes.DRB1*1101 and DRB1*12022 are DR5 subtypes.
The DQA1*0102/DQB1*0602 haplotype is common in narcoleptic patients (Mignot et al. 1994). Other haplotypes withDQA1*0102but not DQB1*0602, such as DQA1*0102andDQB1*0604, are frequent in control populations in all ethnic groups and do not predispose to narcolepsy. DQA1*0102 alone is thus not likely to confer susceptibility but may be involved in addition to DQB1*0602 for the development of narcolepsy (Mignot et al. 1994, 1997a).
Microsatellite analysis in the HLA DQ region revealed that only the area surrounding the coding regions of DQB1 andDQA1 is well conserved across all susceptibility haplotypes. Polymorphism can be observed in microsatellite and/or in the promoter regions flanking the DQB1*0602 and DQA1*0102alleles and in the region between these two genes (Mignot et al. 1997a). Mutations by slippage for some loci, and rare ancestral crossovers in a few instances, contribute to this diversity (Mignot et al. 1997a). Sequence analysis of DQ genes from narcoleptic and control individuals has revealed no sequence variation that correlates with the disease (Lock et al. 1988; Uryu et al. 1989; Ellis et al. 1997;Mignot et al. 1997a). No new gene was found in 86 kb of genomic sequence surrounding the HLA DQ gene (Ellis et al. 1997). A study on the dosage effect of DQB1*0602 allele on narcolepsy susceptibility revealed that DQB1*0602 homozygous subjects are at two to four times greater risk than heterozygous subjects for developing narcolepsy (Pelin et al. 1998). Taken together, these results strongly suggest that the DQA1*0102 andDQB1*0602alleles themselves rather than an unknown gene in the region are the actual susceptibility genes for narcolepsy.
……
Narcolepsy is both a significant medical problem and a unique disease model. Research in humans has led to the identification of specific HLA alleles that predispose to the disorder. This has suggested the possibility that narcolepsy may be an autoimmune disorder, a hypothesis that has not been confirmed to date. Cells of the central and peripheral nervous systems and immune systems are known to interact at multiple levels (Morganti-Kossmann et al. 1992; Wilder 1995). For example, peripheral immunity is modulated by the brain via autonomic or neuroendocrinal interactions, whereas the immune system affects the nervous system through the release of cytokines. Cytokines have been shown to modulate sleep directly and have established effects on neurotransmission and neuronal differentiation (Krueger and Karnovsky 1995; Mehler and Kessler 1997). It is therefore possible that neuroimmune interactions that are not autoimmune in nature might be involved in the pathophysiology of narcolepsy.
NREM and REM sleep are mainly regulated by circadian and homeostatic processes. Single gene circadian mutations have been isolated from species as diverse as Arabidopsis(toc1),Neurospora (frq), Drosophila (perand tim), and mouse (Clock) (Hall 1995). Theper and Clock genes isolated inDrosophiliaand mouse, respectively, have been shown to belong to the same family, the PAS domain family (Hall 1995; Rosbash et al. 1996; Young et al. 1996; King et al. 1997). Analysis of frq, tim,andper demonstrate that circadian rhythms of diverse species are regulated by similar negative feedback loops in which gene products negatively regulate their own transcripts (Hall 1995;Dunlap 1996;Rosbash et al. 1996; Young et al. 1996). Putative homologs of theper gene have also been isolated in mammals (Albrecht et al. 1997; Tei et al. 1997). In mouse, RNAs for two perhomologs are expressed rhythmically within the suprachiasmatic nucleus (SCN), a brain region with an established role in generating mammalian circadian rhythms (Shearman et al. 1997;Shigeyoshi et al. 1997; Tei et al. 1997).
Much less progress has been made in the noncircadian aspect of sleep regulation. Sleep can only be recognized and characterized electrophysiologically in mammals and birds, and single gene mutants for this behavior have not been described in the mouse. Canine narcolepsy is the only known single gene mutation affecting sleep state organization as opposed to circadian control of behavior. The molecular cloning of this gene may lead to a better understanding of the molecular basis and biological role of sleep, as has been the case for circadian rhythms following the cloning of frq, per, and Clock.
By Karen Zusi
A web of cell types in one of the brain’s chief wake centers keeps animals up—but also puts them to sleep.
Feature: Desperately Seeking Shut-Eye
By Anna Azvolinsky
New insomnia drugs are coming on the market, but drug-free therapy remains the most durable treatment.
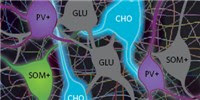
Selectively driving cholinergic fibers optically in the thalamic reticular nucleus promotes sleep
Cholinergic projections from the basal forebrain and brainstem are thought to play important roles in rapid eye movement (REM) sleep and arousal. Using transgenic mice in which channelrhdopsin-2 is selectively expressed in cholinergic neurons, we show that optical stimulation of cholinergic inputs to the thalamic reticular nucleus (TRN) activates local GABAergic neurons to promote sleep and protect non-rapid eye movement (NREM) sleep. It does not affect REM sleep. Instead, direct activation of cholinergic input to the TRN shortens the time to sleep onset and generates spindle oscillations that correlate with NREM sleep. It does so by evoking excitatory postsynaptic currents via α7-containing nicotinic acetylcholine receptors and inducing bursts of action potentials in local GABAergic neurons. These findings stand in sharp contrast to previous reports of cholinergic activity driving arousal. Our results provide new insight into the mechanisms controlling sleep.

Leave a Reply